-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaGenomic Analysis of the Hydrocarbon-Producing, Cellulolytic, Endophytic Fungus
The microbial conversion of solid cellulosic biomass to liquid biofuels may provide a renewable energy source for transportation fuels. Endophytes represent a promising group of organisms, as they are a mostly untapped reservoir of metabolic diversity. They are often able to degrade cellulose, and they can produce an extraordinary diversity of metabolites. The filamentous fungal endophyte Ascocoryne sarcoides was shown to produce potential-biofuel metabolites when grown on a cellulose-based medium; however, the genetic pathways needed for this production are unknown and the lack of genetic tools makes traditional reverse genetics difficult. We present the genomic characterization of A. sarcoides and use transcriptomic and metabolomic data to describe the genes involved in cellulose degradation and to provide hypotheses for the biofuel production pathways. In total, almost 80 biosynthetic clusters were identified, including several previously found only in plants. Additionally, many transcriptionally active regions outside of genes showed condition-specific expression, offering more evidence for the role of long non-coding RNA in gene regulation. This is one of the highest quality fungal genomes and, to our knowledge, the only thoroughly annotated and transcriptionally profiled fungal endophyte genome currently available. The analyses and datasets contribute to the study of cellulose degradation and biofuel production and provide the genomic foundation for the study of a model endophyte system.
Published in the journal: . PLoS Genet 8(3): e32767. doi:10.1371/journal.pgen.1002558
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002558Summary
The microbial conversion of solid cellulosic biomass to liquid biofuels may provide a renewable energy source for transportation fuels. Endophytes represent a promising group of organisms, as they are a mostly untapped reservoir of metabolic diversity. They are often able to degrade cellulose, and they can produce an extraordinary diversity of metabolites. The filamentous fungal endophyte Ascocoryne sarcoides was shown to produce potential-biofuel metabolites when grown on a cellulose-based medium; however, the genetic pathways needed for this production are unknown and the lack of genetic tools makes traditional reverse genetics difficult. We present the genomic characterization of A. sarcoides and use transcriptomic and metabolomic data to describe the genes involved in cellulose degradation and to provide hypotheses for the biofuel production pathways. In total, almost 80 biosynthetic clusters were identified, including several previously found only in plants. Additionally, many transcriptionally active regions outside of genes showed condition-specific expression, offering more evidence for the role of long non-coding RNA in gene regulation. This is one of the highest quality fungal genomes and, to our knowledge, the only thoroughly annotated and transcriptionally profiled fungal endophyte genome currently available. The analyses and datasets contribute to the study of cellulose degradation and biofuel production and provide the genomic foundation for the study of a model endophyte system.
Introduction
Global climate change and decreasing fuel reserves are driving a push towards biologically derived fuels from plant wastes. The optimal biofuel for immediate implementation is one that functions within the context of current infrastructure, in particular with existing engines and distribution systems. This would require chemical similarity to gasoline, which is a mixture of hydrocarbons with an average chain length of eight [1]. Fungi have been recognized as producers of eight carbon (C8) volatiles for nearly 80 years and are a major global carbon recycler [2],[3]; however, despite the interest in these compounds, the genes responsible for their production remain largely undefined.
One such producer of C8 volatiles is the endophyte Ascocoryne sarcoides (NRRL 50072). Originally identified as Gliocladium roseum, this organism was shown to produce a series of molecules of potential interest as biofuels when grown on a cellulose-based medium [4]. The taxonomy was later revised to A. sarcoides and its production profile of Volatile Organic Compounds (VOCs) was amended to remove branched-chain alkanes. However, this follow-up work also confirmed the production of straight-chain alkanes from C6 to C9, as well as branched-chain alcohols varying in length from C3 (2-methyl-1-propanol) to C7 (5-methyl-1-hexanol) (Table S1) [5]–[7]. Understanding and optimizing biological production of such molecules is an area of active research (reviewed in [8]).
Bacteria have been shown to produce alkenes through “head-to-head” condensation of fatty acids; however, products with fewer than 23 carbons, like those from A. sarcoides, are not known to be synthesized by this mechanism [9],[10]. Odd-chain alkanes and alkenes of chain lengths 13–19 have been observed in bacteria as products of the decarbonylation of aldehydes and the decarboxylation of fatty acids, respectively [11],[12]. However, currently there are no known eukaryotic homologs for these enzymes. C8 alcohols and ketones have been identified as the products of linoleic acid breakdown; however, the genes responsible for the downstream reductions that generate C8 alkenes and alkanes are still unknown [13]–[17]. In order to gain a better perspective on these pathways and the cellulolytic machinery used by an endophyte, we coupled genome sequencing and short and long RNA-seq with metabolomic profiling of A. sarcoides.
Generation of metabolic pathway predictions in organisms for which genetic tools have not yet been developed remains a difficult problem. Techniques such as gene expression analyses and metabolomics profiling have the advantage that genetic tractability is not required. In a pioneering study, Askenazi et al, showed that gene expression could be linked to specific metabolite production [18]. The authors profiled the level of lovastatin production in engineered strains of the fungus Aspergillus terreus and showed that strains with similar transcriptional profiles also had similar amounts of lovastatin production [18]. Furthermore, extensive metabolic network analyses have demonstrated the ability to link the transcription of individual genes to metabolites [19],[20]. Metabolite-transcriptional coupling has since been validated extensively for the monitoring of different stress responses [21]–[23].
We used RNA-seq based gene expression measurements to accurately map gene structures and to generate candidate gene lists for novel metabolic pathways. In particular, we used gene expression and the co-occurrence of a compound across multiple experimental perturbations to generate candidate genes and pathways for the production of C8 volatiles and several other alkanes and alkenes that currently have no known eukaryotic pathway. In addition, we extensively mapped and annotated the A. sarcoides cellulose breakdown machinery using RNA-seq expression analysis after growth on different carbon substrates. Together with the high quality genome assembly and annotation, these data provide the most complete genomic characterization of any fungal endophyte to date. The analyses and datasets contribute to the development of biofuels from microbial metabolites and the related study of cellulose degradation and may be a reservoir of information for studying the plant-endophyte relationship.
Results
Genome assembly and annotation
The A. sarcoides NRRL 50072 genome was sequenced resulting in approximately 38-fold coverage of the estimated 34 Mb genome [24]. Reads were assembled into 16 scaffolds incorporating 99.5% of the total genomic base pairs. The genome size and overall GC content (45%) is within the average range for other Leotiomycetes fungi [25]. We predicted 10,831 genes resulting in 100% recovery of annotated Core Eukaryotic Genes Mapping Approach (CEGMA) genes which is a benchmark for a high quality genome assembly (Text S1) [26]. Roughly 70% of the gene models had at least one match to one of the 42 available fully sequenced fungal genomes. Approximately 22% of the gene models are seemingly species-specific and did not match to anything currently in GenBank [27]; the remaining 8% were homologous to genes outside of the fungal kingdom. Eighty-seven percent of the gene models were validated with long-read transcriptome profiling (Text S1) and 75% of the potential exon-exon junctions were confirmed (see Figure 1A and 1B). Although a subset of the unvalidated gene models and exon junctions may be spurious, the majority are most likely true genes that are silent under these specific conditions [28],[29].
Fig. 1. Validating gene models and novel TARs. 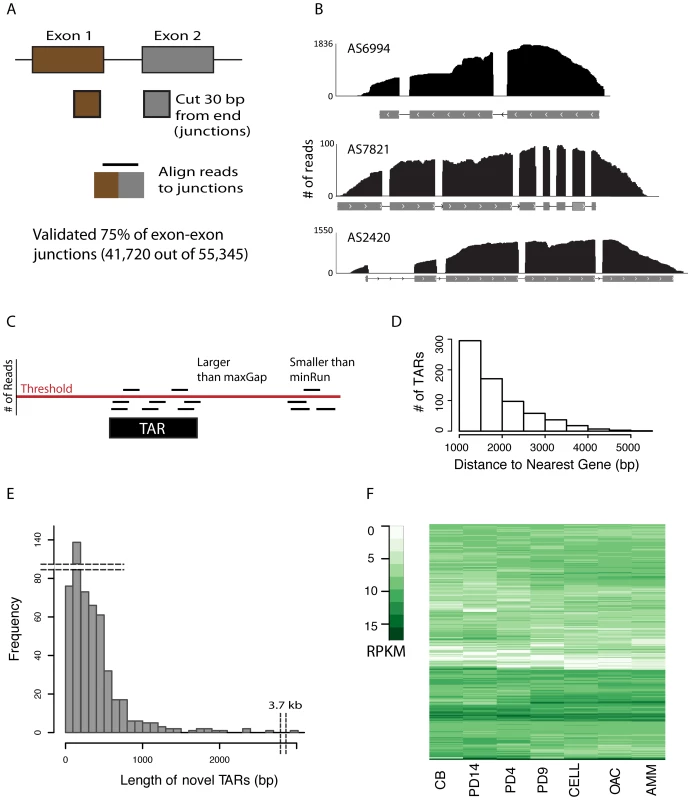
(A) Schematic showing splice junction library generation. (B) For each of the three gene models shown, the x-axis is the genomic coordinates and the gray boxes represent individual exons, with arrows indicating strand. Reads having any overlap with the genic region are represented by black lines, the height of which correspond to the number of reads covering a particular base pair. Note that a read can align outside the exonic region, but that this was not observed at intron boundaries, although it did occur in the UTRs. (C) Schematic illustrating de novo assembly of reads into transcriptionally active regions (TARs). Three parameters are shown: threshold, min run, and max gap. Threshold sets the number of reads required for the region to be considered in the assembly. minRun sets the number of base pairs in the contiguous region required, and maxGap sets the number of discontiguous base pairs permitted to still be considered part of the assembly. Only the black box has sufficient base pairs above the threshold with the permitted contiguous length to be considered a TAR. (D) The minimum distance between each TAR and its nearest neighboring gene was computed. The number of TARs at least 1 kb away from any gene are shown (novel TARs). (E) Histogram of the length of novel TARs. Note the break in both the x and y-axis to indicate the outliers for TAR length and frequency. (F) Columns represent the culture growth conditions, rows individual novel TARs, and elements are color coded according to their RPKM value from white (no expression) to dark green (high expression). RNA-seq analysis and novel TAR identification
We subjected A. sarcoides to seven different growth conditions to assay diversity in both transcription and compound production (Table S2). Volatile metabolite production was analyzed by gas chromatography mass spectrometry (GC/MS) for six of these seven conditions (no GC/MS dataset was obtainable on the day 9 potato dextrose harvest; Table S1 and S2). We monitored A. sarcoides cultures for production of volatiles and selected this subset of six conditions for RNA-seq analysis, which provided differential compound production profiles. Under these six conditions, A. sarcoides produced 48 identifiable volatile metabolites including 18 alcohols and 7 alkanes/alkenes including heptane, octane, and nonane. All volatile metabolites were scored with a binary scale to indicate their presence or absence in each culture headspace. We chose this digitized scoring because different analyses required variation in culture and headspace volumes and our method of detection of VOCs (Solid Phase Micro Extraction (SPME), see Materials and Methods) is sensitive to such variation [30]. The large number of functionally diverse metabolites in the headspaces also precluded the use of external or internal standards to determine the absolute amount detected for each compound across all conditions.
Coupled transcriptional profiles for the six conditions obtained via RNA-seq resulted in more than 200 million reads alignable to the reference genome or exon junctions (Table S3) and greater than 99% similarity between the two technical replicates (Figures S1 and S2). Six diverse sampling conditions were chosen for the RNA-seq analysis in lieu of replicates in order to more thoroughly explore the transcriptional landscape of A. sarcoides and more completely map gene structure throughout the genome. The genome and transcriptome data can be accessed at http://asco.gersteinlab.org.
In addition to the 10,831 gene models predicted, we identified a number of RNA-seq reads which map outside of the gene models. A subset of these reads formed well-defined regions on the reference genome. 602 of these regions are at least 1 kb away from any annotated genes and are designated as transcriptionally active regions (TARs) (Figure 1C, Figures S3 and for sensitivity analysis and examples). These TARs were seemingly devoid of open reading frames and in some cases were quite long (up to 3.7 kb in length). Forty percent of these TARs illustrated condition-specificity (standard deviation greater than 1; see Figure 1D–1F) as has previously been observed in S. cerevisiae and H. sapiens [31]. The importance of these polyadenylated non-coding RNAs in regulating gene expression has only recently been discovered [31],[32] and their exact role remains an active area of research.
Annotation and expression of cellulose degradation machinery
Given the emphasis on cellulose breakdown and utilization for the development of alternative fuels, we were interested in exploring and annotating the cellulolytic capabilities of A. sarcoides. We analyzed the transcription profiles of A. sarcoides for growth on three different carbon sources: cellulose (CELL), cellobiose (CB), and potato dextrose (PD4). While cellulose and cellobiose share the same β(1–4) linkage between monomer units, potato dextrose contains predominantly glucose-monomer. Differential gene expression between the potato dextrose and the two more complex substrates (CELL and CB) provides information on the pathways and mechanisms of cellulose breakdown; whereas, differences between the CELL and CB provides information on the genes necessary to utilize a soluble versus an insoluble polymer. Such differences are particularly useful as they can inform methods aimed at increasing cellulose breakdown efficiency. We first examined the differential expression across these three conditions (Table S15) [29],[33]. There were 1,435 genes that were expressed under all three conditions (Figure 2A). A smaller number, 142 genes, were only expressed during growth on cellulose or cellobiose, including the endo - and exo-cellulases, as expected based on their role in cellulose utilization. 398 and 380 genes were exclusively expressed on cellobiose and cellulose, respectively, reflecting the significant differences in the machinery necessary to utilize a soluble disaccharide versus an insoluble polymer and in the resulting downstream changes in the cellular state. We focused on the subset of genes with homologs in the CAZY database, a manually curated repository for carbohydrate metabolism (see Text S1) [34]. In total, 52% (89 of 169) of glycosyl-hydrolase homologs (GH), 45% (25 of 56) of glycosyl-transferases (GT), 50% (3 of 6) of carbohydrate-binding module genes (CBM), 41% (9 of 22) of carbohydrate esterases (CE), and 0% (0 of 1) of polylyase (PL) were differentially expressed across the three conditions (Figure 2B; Table S4).
Fig. 2. Analysis of cellulose-related expression. 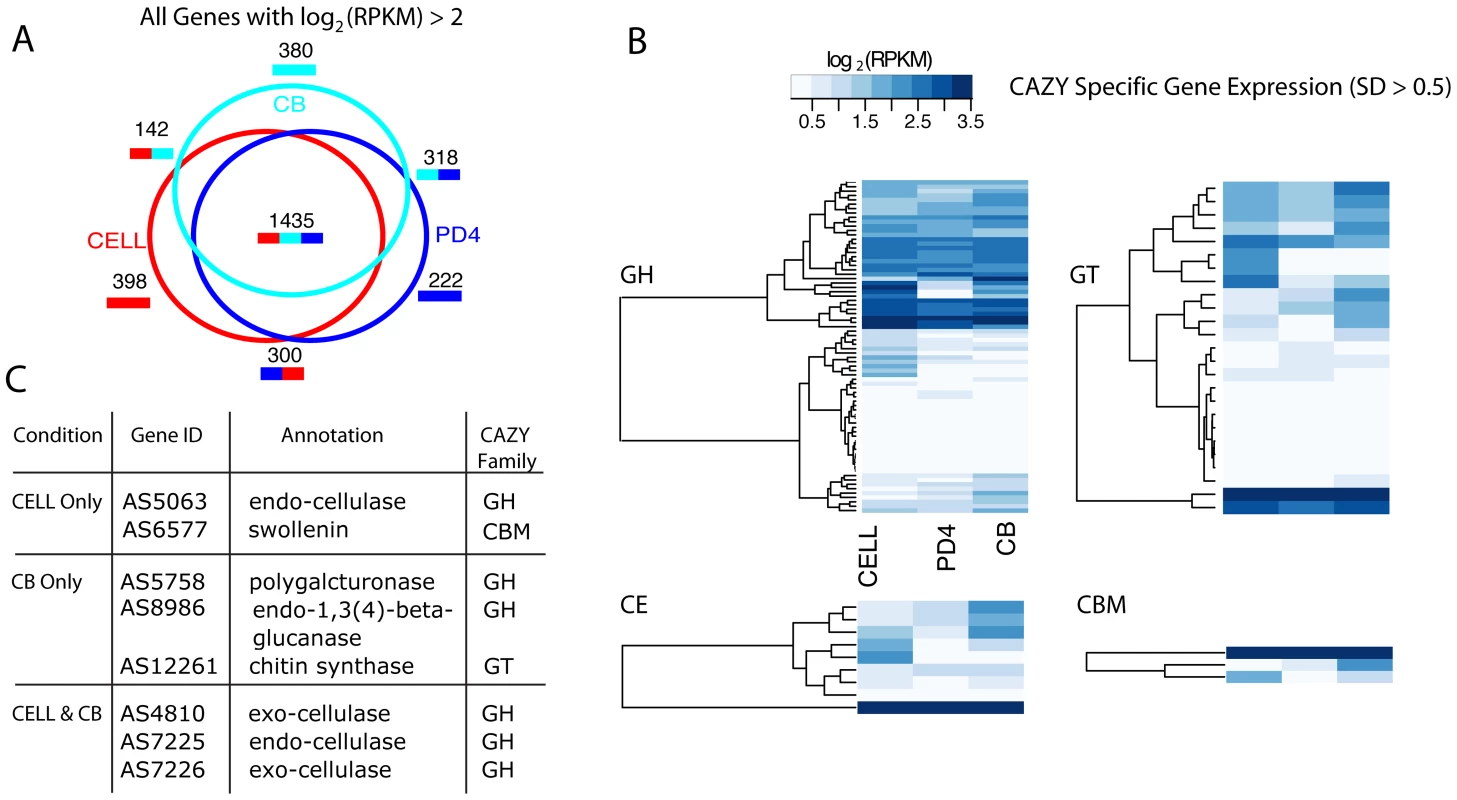
A. sarcoides transcription was profiled when grown on potato-dextrose media for 4 days (PD4), cellulose (CELL) and cellobiose (CB). (A) The total number of genes with quantile normalized log2(RPKM) greater than 2 was computed for each condition. The venn diagram shows the overlap of these genes across the three conditions. (B) Genes were partitioned according to their homology to the four main CAZY families: Glycoside Hydrolase (GH), Glucosyl Transferase (GT), Carbohydrate Esterase (CE), Carbohydrate Binding Modules (CBM). The homologs were then filtered to include only those genes which showed a standard deviation across the three conditions greater than 0.5. Each family was separately clustered (hierarchical, Euclidean distance, single linkage). The colorbar represents the quantile normalized log2 (RPKM) value from white (low expression) to dark blue (high expression). Note: CBM can co-occur with all families. Only those genes that had exclusively a CBM domain were clustered in the CBM matrix to avoid duplication. (C) A table of the most highly expressed genes includes genes not directly involved in degradation, such as swollenin and chitin synthase (see Results for more details). The most highly expressed gene in the cellulose condition was AS6577, which is homologous to the gene encoding the protein swollenin. Swollenin was first identified in the cellulolytic model organism, Trichoderma reesei. Heterologous expression in yeast and Aspergillus niger showed that swollenin mediates disruption of plant cell walls without releasing monomeric sugars [35]. Supplementation of a cellulase mixture with swollenin increased saccharification rates suggesting this protein may play an important role in efficient cellulose breakdown [36]. While A. sarcoides growth on a lignin-containing medium was not analyzed, we identified the full pathways for 5-carbon sugar utilization e.g. arabinose and xylose, sugars which comprise 10–25% of carbohydrates resulting from hemi-cellulolysis [37]. We further validated the presence of these pathways by demonstrating A. sarcoides growth on media with either xylose or arabinose as the sole carbon source (Materials and Methods).
Identification of genes for biosynthesis of secondary metabolites
The genes responsible for both cellulose degradation and the production of secondary metabolites are non-randomly distributed in a number of sequenced genomes, such that they are clustered into regions of higher than average gene density [38],[39]. Therefore, we searched for clusters in A. sarcoides as a strategy to identify genes involved in these processes. We generated a simulated set of scaffolds where the number of genes was kept constant but the placement was randomized to identify regions of the genome with higher than expected gene density. We identified 77 clusters ranging in length from 10–72 kb (p<.05, Text S1). Twenty-six clusters contained genes or domains known to be involved in secondary metabolism, particularly oxidoreductases and permeases. We noted five gene-clusters that were involved in the production of secondary metabolites usually restricted to plants, including two clusters containing genes homologous to those involved in the synthesis of patatin (Table S5). Patatin is a plant storage glycoprotein implicated in plant-fungal communication [40]. Expression of this protein in Arabidopsis negatively affects resistance to Botrytis cinerea and Pseudomonas syringae, but it increases resistance to the cucumber mosaic virus [40]. Interestingly, all genes in this cluster were transcriptionally silent under the conditions we tested. Given their known functional role in mediating plant-fungal interactions, it is possible they are strictly regulated by interactions with the plant host.
The classes of genes most frequently involved in secondary metabolite production are Polyketide synthases (PKS) and Non-ribosomal peptide synthetases (NRPS). We identified 19 PKS and NRPS clusters through fungal-specific Hidden Markov Models of beta ketoacyl synthase (KS) and acyltransferase (AT) domains and an additional 8 gene clusters and 11 gene models composed solely of enoyl reductase and/or dehydratase accessory domains (Text S1, Tables S6 and S7). The identified PKS genes ranged in size from a few kb to the 13 kb and 13-exon hybrid PKS/NRPS AS8071, which is by far the largest predicted gene model in A. sarcoides. Examination of the 3 kb region upstream and downstream of each PKS element also revealed a number of major facilitator superfamily transporters and permeases which may confer resistance to both PKS-derived and exogenous toxins [41]. However, comprehensive searches of previously identified PKS clusters [42], laeA element identification to delineate possible cluster boundaries [43], and use of domain to structure software [44] failed to yield any predictions for possible biosynthetic products. Intriguingly, one PKS, AS1082 was first found to contain a beta ketoacyl synthase domain, but subsequent searches revealed that it contained two distinct KS domains and an acyl carrier protein domain. However, no acyl transferase domain, which typically functions in substrate loading, was identified. While separately encoded acyl transferase enzymes that act in trans have been found in bacteria, only trans-acting enoyl reductase domains have yet been characterized in fungi [45].
Correlating VOC production with gene expression to elucidate biosynthetic pathways
A more direct method to investigate the A. sarcoides genes responsible for production of the novel metabolites is the use of association analysis. As mentioned above, the concordance of gene expression and metabolite production can be used to guide prediction of genes involved in metabolic pathways [18]. A complication in the application of these methods for novel metabolic pathways, as opposed to those generated either via PKS or as part of conserved metabolism, is that we know neither the genes that are involved nor the pathway structure (i.e. the reactant-product pairings that result in the downstream compound). For example, we do not know the genes responsible for the production of octane, nor do we definitively know the starting compound or what intermediates may have been subsequently generated. Thus, we need a series of analyses that simultaneously infer the potential genes and the pathway trajectory as defined by the chemical elements (Figure 3A–3D; Figures S5, S6, S7).
Fig. 3. Coupled transcriptomics and metabolomics to generate pathway predictions. 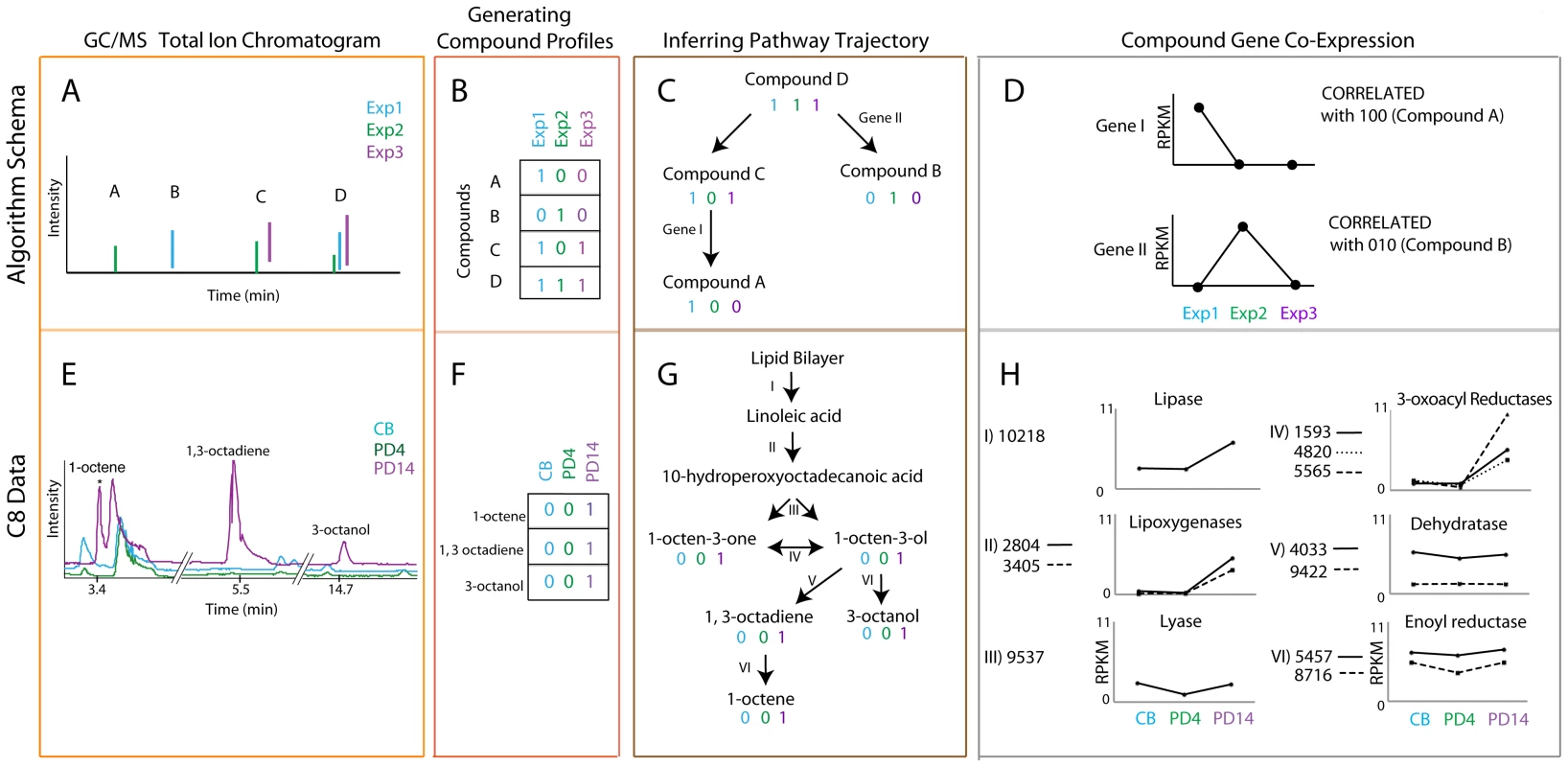
The top panels (A–D) represent the algorithm schema and the bottom panels (E–H) represent the corresponding steps with data for an example pathway, C8 production. Cyan, green, and purple are used to denote different experimental conditions (1, 2, and 3 and CB, PD4, and PD14 for the schematic and the C8 pathway data, respectively). GC/MS total ion chromatograms (orange box, A & E) are used to generate compound co-occurrence profiles (red box, B & F). These compound co-occurrence profiles are used to group and order the compounds based on patterns of correlation and anti-correlation to build a possible biosynthetic pathway (brown box C & G). Genes for which the expression profile matches the compound profile are considered correlated and therefore likely candidates for the biosynthetic pathway of interest (gray box D & H). Retrosynthesis is then used to match correlated genes with a reaction in the pathway, represented by roman numerals denoted on pathway arrows (brown box, C&G). It was previously shown that by examining the “correlation” and “anti-correlation” of sets of genes across a wide spread of phylogenetic space, the importance, ordering, operons, and additional members of the pathway can be discerned [46]–[50]. Furthermore, genes belonging to the same pathway or complex often show both coordinated regulation and conservation [50]. By substituting the phylogenetic profiles from these previous studies with our compound profiles generated from compound presence or absence across all conditions, the resulting character matrix can be used to determine the relatedness of these compounds (Figure 3B and 3F, Figures S7 and S8). On the basis of these relationships, compounds can be then grouped into pathways. To apply this correlation analysis, each metabolite produced by A. sarcoides under each of the six growth conditions was assigned a “1” if it was detected in the particular condition and “0” if it was not detected, as depicted in the schema in Figure 3A–3B and 3E–3F). To further inform the metabolite analysis, we also used a recent meta-analysis that profiled the production of 10 Ascocoryne isolates under varying growth conditions resulting in 20 different GC/MS profiles [5]. Compounds that consistently co-occurred across the genus are more likely to be in the same pathway and were given more weight than those showing inconsistent behavior (Figure S7). We then grouped sets of compounds that co-occurred into single or related sets of pathways (Figure 3C, compounds A and C) and those that rarely or never co-occurred into different pathways (Figure 3C, compound B). To identify possible metabolite-gene linkages, we then computed the correlation between the compound profile and expression of each gene under the different conditions. Correlations between compounds and expression were used instead of strictly quantitative changes in gene expression because this more effectively integrated the expression analysis with the binary compound production data. To ensure the correlations were significant, we computed a p-value for the compound co-expression scores (See Text S1, Tables S8, S9, S10, S11, S12, S13). For a set of compounds with the same compound profile, there may be many genes with correlated expression, not only those involved in the compound production. Therefore, retrosynthesis was used to disambiguate which of the significantly correlated genes were most likely involved in the production of those compounds (Figure S8).
As one example of this method identifying candidate genes, we identified 60 genes with homology to putative alcohol dehydrogenases (EC 1.1.1.1), which have a wide range of specificities and annotation quality. However, only three of the identified alcohol dehydrogenases were significantly co-expressed with any compound production profile. In particular, AS5307 was co-expressed with the compound profile that had a predominance of branched medium chain alcohols, including 3-methyl butanol, 3-methyl-3-buten-1-ol, and 2-methyl-1-propanol. We predict that these three dehydrogenases, from amongst the 60, play a key role in the production of the observed medium-chain alcohol metabolites.
Co-expression has been used to assign functions to genes with known homologs as well as to genes without primary sequence or domain level annotations [51]. All genes co-expressed with a particular compound profile were examined as shown in Figure 4, where each line represents a single gene. A subset of the genes was homologous to well-known secondary metabolite pathway elements, but some had no known function (Figures S9, S10, S11). In the latter cases, gene co-expression was used to infer additional pathway elements as well as associated regulators and transporters. Below, we provide an example set of predictions for a C8 product pathway. The full set of predicted pathway schemas and potential enzymes are provided in the supplement. An R package containing the code and documentation for RNA-seq processing and the association analysis is provided in Text S2.
Fig. 4. Compound gene co-expression profiles. 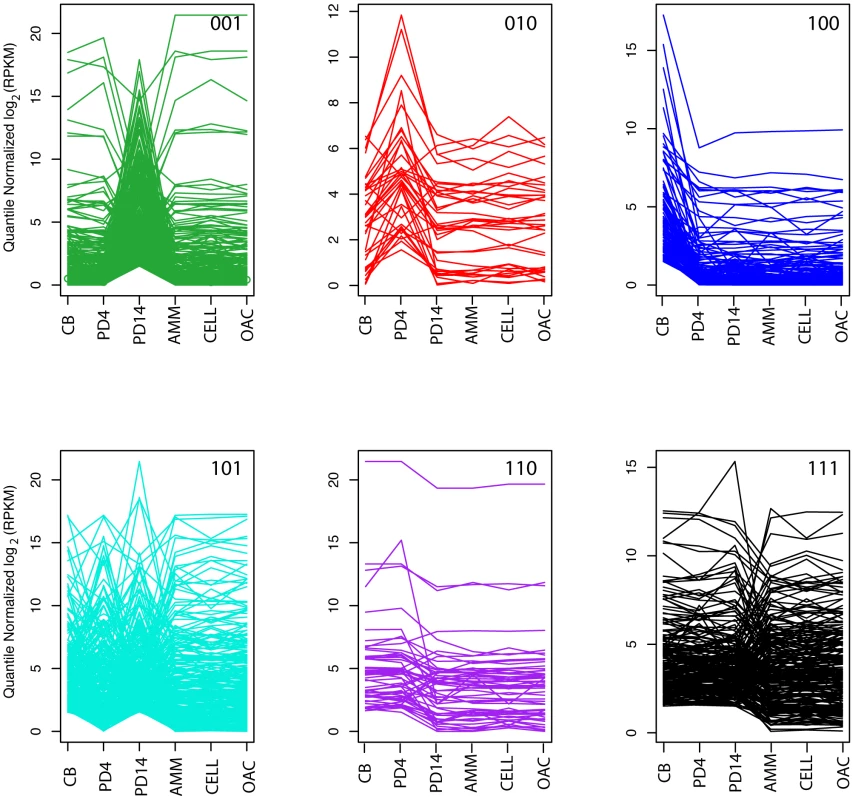
Each plot shows the quantile-normalized log2 (RPKM) for each set of genes of co-expressed with a particular compound profile (green 001, red 010, blue 100, cyan 101, purple 110, and black 111) across all 6 conditions (CB, PD4, PD14, AMM, CELL, and OAC). The first three conditions (CB, PD4, and PD14) represent the conditions where the compounds analyzed in this study were detected. The remaining conditions serve as the nulls (see Text S1for details). Within the plots, each line corresponds to a single gene. Pathway predictions for biosynthesis of C8 metabolites
Given the average chain length is about eight for hydrocarbons in gasoline, the production of molecules with similar lengths represents an obvious starting point for next generation biofuels that will be compatible with pre-existing infrastructure [52],[53]. We identified candidate pathway elements for the production of reduced C8 volatiles in A. sarcoides and assigned correlated genes to each step of the reconstructed C8 pathway (Figure 3). As an example, lipoxygenases (EC 1.13.12.12) are known to be involved in the formation of C8 alcohols and ketones in fungi via the catabolism of linoleic acid [3],[54]. There are five lipoxygenases in the A. sarcoides genome, and two of these are correlated with C8 production (AS2804 and AS3405, Figure 3H, II). The most strongly correlated lipoxygenase, AS2804 is homologous to the Aspergillus gene ppoC (Figure 3H, II) (p<.01). Recently, Brodhun et al showed that expression of ppoC is sufficient to catalyze the breakdown of linoleic acid into a wide range of compounds including: 1-octen-3-ol, 2-octen-1-ol, 2-octenal, and 3-octanone in a crude E. coli lysate [17]. All of these compounds were observed as products of A. sarcoides with the exception of 2-octenal (Table S1). The original hypotheses for the production of these C8 volatiles from linoleic acid involved two enzymes, a lipoxygenase to form a peroxidated intermediate, and a lyase (EC 4.1.2.-) to catalyze its breakdown into smaller, volatile products. However, an active lyase has yet to be successfully purified in fungi [14]–[16],[55], and recent work argues against the need for this activity [17]. We identified one lyase, AS9537; however, its expression did not correlate with the production of C8 volatiles (Figure 3G, III), arguing against the dual-enzyme hypothesis for C8 production and supporting the more central role for the lipooxygenase (AS2804).
In addition to the oxygenated C8 volatiles observed by Brodhun et al. from Aspergillus, A. sarcoides produces the reduced compounds 1,3-octadiene; 1,3-trans-5-cis-octatriene; 1,5-octadien-3-ol; 1-octene; and 3-octanol suggesting that downstream processing of linoleic acid breakdown products has occurred. One potential route to these compounds is shown in Figure 3G, whereby 1-octen-3-one is further reduced to 1-octen-3-ol by FabG (EC 1.1.1.100), a 3-oxoacyl-[acyl-carrier protein] reductase (Figure 3G, IV). In total, A. sarcoides has 10 genes with strong homology to FabG (Table S14), however, only the three co-expressed with C8 production are shown in Figure 3H, IV (AS1593, AS4820, and AS5565, with AS5565 exhibiting the largest expression change). The nearest sequenced relatives of A. sarcoides, Botryotinia fuckeliana, and Sclerotinia sclerotiorum, have only two and three FabG genes, respectively (Figures S13 and S14). Since the reduced C8 compounds have not previously been found outside the Ascocoryne genus and the expression of some FabG genes do correlate with these compound production profiles, it is possible that at least some of these additional FabG genes may participate in the reduction of eight carbon volatile compounds. In addition to the FabG homologs, 317 oxidoreductases particularly aldo-keto reductases, were identified in A. sarcoides. Of these 11 were correlated with C8 production (Table S14). Oxidreductases are able to reduce various functional groups, such as ketones and alcohols, and are expected to participate in the biosynthesis of the C8 reduced products and other volatiles (Figure 3G, 3H; IV and VI). In addition, of all sequenced fungal genomes, only A. fumigatus (626) and T. reesei (494) have a commensurate number. Both B. fuckeliana and S. sclerotiorum have less than 200 oxidoreductases, which is approximately the median number for sequenced fungi. The above average number of oxidoreductases found within A. sarcoides suggests a large reducing capability and extensive secondary metabolism potential.
Discussion
The unknown pathway for the production of potential biofuel compounds in A. sarcoides is part of a more general trend. Microorganisms produce an extraordinary diversity of natural products that have the potential to be used in numerous applications from medicines to biofuels to commodity chemicals [37],[56],[57]. However, identifying the genes responsible for their production remains a major hurdle for organisms that are not genetically tractable. Despite promising developments in pathway prediction algorithms, a substantial gap remains between metabolic capabilities and genetic characterization [58]–[61]. As an example, Metacyc, a repository of metabolic pathways, contains 8,869 compounds linked to 1,908 known pathways, but this represents less than 1% of the compounds estimated to be produced by micro-organisms [62],[63]. An integrated omics approach could provide a relatively simple means of exploring the biosynthetic potential of uncharacterized non-model organisms.
By examining changes in the A. sarcoides transcriptome across a diverse array of conditions, we were able to explore a wide fraction of genes and refine gene and exon boundaries to improve the genome annotation quality. Additionally, with co-expression patterns we generated hypotheses for the genes involved in undefined metabolic pathways and regulatory mechanisms. Through TAR building we identified a number of long, highly expressed regions seemingly devoid of open reading frames that may have a regulatory role. The recovery of 100% of all CEGMA [26] genes suggests a high quality genome assembly, and the number of scaffolds is on par with the number of expected chromosomes in Ascomycete fungi [64]. We used an expanded version of association analysis to generate hypotheses for products from unknown pathways. Such methods are flexible enough to integrate coupled transcriptome and metabolomics data and will take on increasing importance as the throughput of both transcriptome and metabolomics continues to increase. The means to leverage these datasets will be key to our understanding of novel metabolite production particularly for genetically intractable organisms. From its plant mediators to its oxidoreductases and its cellulases, A. sarcoides's gene complement represents several avenues for further research and its diverse array of enzymatic capabilities will contribute to the study of cellulose degradation and secondary metabolite production.
Materials and Methods
Genomic DNA isolation
Isolate NRRL 50072 was obtained under a material transfer agreement from Montana State University (GA Strobel, Bozeman, MT). Genomic DNA was isolated using the Plant DNeasy MaxiPrep kit (Qiagen) according to the manufacturer's instructions with the following modifications: mycelia were grown in potato dextrose broth for approximately 3 weeks at 25°C, shaking at 150 rpm and were harvested via filtration. The filtrate (1 g) was homogenized by mortar and pestle under liquid nitrogen before the addition of 80 µL RNase (100 mg/mL), 80 µL proteinase K (10 mg/mL) and lysis buffer P1 (Qiagen). Homogenized material was heated for 10 min at 65°C and then processed through the remainder of the Qiagen protocol.
Sample preparation of RNA for Illumina RNA-seq
Please see Table S2 for detailed growth and inoculation conditions for CB, PD4, and PD14 as distinguished by the short code referred to in both the text and figure legends. For the remaining 3 conditions (OAC, AMM, and CELL), media were prepared and inoculated with 50 mg filtered culture (1× PD) as reported in Griffin et al., 2010 [5]. Carbon starvation (OAC) was prepared as a minimal medium base with sodium acetate (50 mM) as the sole carbon source. Nitrogen starvation (AMM) was prepared as a minimal medium base with no ammonium chloride and with 83.3 mM glucose. Cellulose substrate (CELL) was prepared as a minimal medium base with cellulose (15 g/L) as the sole carbon source. All were titrated to a pH to 6.0 with NaOH. Vials were incubated for 2 days at 23°C before GC/MS analysis and RNA extraction. For each of these conditions, seven vials were inoculated, with three subjected to GC/MS analysis while the remaining four vials were concurrently used for RNA harvesting. Total RNA was isolated using the Ambion RiboPure Kit (California, USA), and then poly-A purified and prepared for sequencing as in Nagalakshmi et al., 2008 [28].
Sample preparation of RNA for long-read transcriptome (454)
Sample PD9 was selected for RNA preparation and long-read transcriptomics, which was used to confirm gene models. RNA was extracted from a 9-day old 1 L PDB culture grown at 23°C, 150 rpm (Table S2). Extraction performed as in Nagalakshmi et al., 2008 [28].
Metabolomics profiling
All conditions were as specified under the RNA-seq preparation. GC/MS was carried out in parallel with cultures harvested for RNA seq with the exception of PD9, which was not profiled. Control samples for each media condition were prepared for use in GC/MS analysis with the same methods as described in the RNA seq conditions section above, but without the addition of inoculums. Analysis of culture headspaces was performed on a gas-chromatograph coupled to a time-of-flight mass spectrometer (GCT Premier, Waters). Automated culture sampling was mediated by a CTC CombiPAL Autosampler (Leap Technologies) and all cultures were sampled with a 50/30 µm divinylbenzene/carboxen/polydimethylsiloxane StableFlex Fiber (Supelco). GC injection and column parameters, GC temperature program and MS data acquisition parameters were as described previously [5]. Parameters for SPME headspace sampling were as follows. OAC, CELL, and AMM vial cultures were analyzed via automated sampling with a pre-extraction SPME fiber conditioning (7 min, 250°C), 35 min headspace extraction at 30°C, and a splitless GC injection (30 sec, 240°C, 0.75 mm ID injection liner). Manual headspace sampling of CB, PD4, and PD14 flask cultures used the following sampling parameters: pre-extraction SPME fiber conditioning (12 min, 250°C), 30 min headspace extraction (room temperature, approximately 20–25°C), and splitless GC injection (30 seconds, 240°C, 0.75 mm ID injection liner). Data were analyzed with the MassLynx Software Suite™ (Waters). Chromatographic peaks were identified with a combination of spectral search comparisons with the Wiley Registry™ of Mass Spectral Data, 8th Edition, elemental composition analysis and the comparison of retention times and spectra with pure standards for compounds where noted (Sigma-Aldrich). Compounds identified during the analysis of control media samples, including contaminants resulting from the SPME fiber and Wax capillary column, as well as media derived compounds, were excluded from the final compound report for each condition. See Table S1 for the full compound profiles.
C5 and C6 growth assays
Growth assays were performed in 96 well plates in 200 µL media containing trace metals as in Griffin et al., 2010 [65], 0.67 g/L Yeast Nitrogen Base (Difco) and supplemented with 100 mM of either glucose, xylose, arabinose, mannose, cellobiose, or sodium acetate, titrated to a pH of 6 with KOH. Wells containing no added carbon source served as the control. The cultures were inoculated by adding 5 µL of 5×107 spores/mL in Phosphate Buffered Saline (Gibco), and the cultures were grown for 5 days at 23°C. Growth was determined by visual inspection.
Genome assembly and annotation
Initial assembly with single end shot gun titanium reads with Roche's GS DeNove Assembler (Newbler) resulted in 137 scaffolds with an N50 of 2.8 Mb [24]. Following addition of a paired end 3 kb-insert sequencing run, these were assembled into 16 scaffolds encompassing 99.5% of the total sequenced base pairs. Called genes were first aligned to the GenBank non-redundant database using blastx (v2.2.24) [27],[66]. A hit was defined as a match when overlap with the length of the query protein was greater than 60% and E-value<1e-10. We extracted the subset of genes found in the CAZY database, a repository for manually curated carbohydrate machinery, and performed a similar procedure [67].
Domains were identified using the hmmsearch function from HMMer [68] with both a set of fungal-specific protein domains [69] and the entire PFAM database [70]. A domain was considered a match if the E-value was greater than 1 and the length of the match was at least 15. Pathway predictions and enzyme classification was completed through KEGG/KAAS [71],[72]. GO predictions were made by first mapping the set of A. sarcoides genes to their corresponding Aspergillus nidulans homolog [73],[74]. Please see Text S1for a full description of the gene cluster and PKS/NRPS identification strategies.
RNA-seq analysis
In the case of the Illumina runs, mapping was done via building bowtie indices for splice junction libraries, and the genome respectively using default parameters (tolerated up to 2 mismatches and screened for quality scores) [75]. Splice junction libraries were generated as described in Habeggar et al., with 30 bp exon ends [76]. The bowtie reads were converted to mapped read format (MRF) and mrfQuantifier was used to compute a variation of reads per kilobase of exon per million mapped sequence reads (RPKM) for each gene using RSEQtools [76]. Briefly, we computed RPKM as the number of nucleotides that map per kilobase of exon model per million mapped nucleotides for each gene rather than the read count. It is computed by summing the total number of reads that cover each base pair of an annotation feature of interest (in this case of exons) and normalizing by the total length of the feature. For the conditions denoted by CELL, OAC, and AMM, technical replicates were performed yielding one lane per replicate. In the case of PD4, there were two technical replicates performed 2 months apart. A comparison of the RPKM of the genes between lane replicates showed greater than 99% agreement (Figure S2), although the correlation was slightly less between the two AMM replicates than between any other conditions.
The 454 long reads (average size 410 bp) were mapped against the gene models and the genome using BLAT with default parameters [77]. In all cases, only reads that unambiguously mapped to a single location were used for the downstream analysis. For each gene, we calculated the RPKM score as described above. To estimate depth of coverage, the percentage of genes that were detectable using subsamples of reads was computed where detectable was defined as having at least 1, 2, 5, or 10 reads, respectively, overlapping the gene (Figure S12).
Identification of transcriptionally active regions (TARs)
A database of transcriptionally active regions (TARs) was constructed from those RNA-seq long reads that map uniquely to the genome via BLAT [77]. The TAR database was built by employing the minrun/maxgap segmenting module [76]. Gene coverage values were calculated for a range of minrun/maxgap parameters to assess their impact on observed gene coverage. Included in the coverage analysis were TAR file sets with maximum read gaps between zero and five and minimum read run from 30 to 40 (See Text S1for a full description; Figures S3 and S4).
Supporting Information
Zdroje
1. SarpalASKapurGSMukherjeeSTiwariAK 2001 PONA analyses of cracked gasoline by 1H NMR spectroscopy. Part II. Fuel 80 521 528 doi:10.1016/S0016-2361(00)00123-X
2. MurahashiS 1938 \Über die riechstoffe des matsutake (Armillaria Matsutake Ito et Imai Agaricaceae). Sci Pap Inst Phys Chem Res(Tokyo) 34 155 172
3. CombetEHendersonJEastwoodDCBurtonKS 2006 Eight-carbon volatiles in mushrooms and fungi: properties, analysis, and biosynthesis. Mycoscience 47 317 326 doi:10.1007/s10267-006-0318-4
4. StrobelGAKnightonBKluckKRenYLivinghouseT 2008 The production of myco-diesel hydrocarbons and their derivatives by the endophytic fungus Gliocladium roseum (NRRL 50072). Microbiology 154 3319 3328 doi:10.1099/mic.0.2008/022186-0
5. GriffinMASpakowiczDJGianoulisTAStrobelSA 2010 Volatile organic compound production by organisms in the Ascocoryne genus and a reevaluation of myco-diesel production by NRRL 50072. Microbiology mic.0.041327-0 doi:10.1099/mic.0.041327-0
6. StrobelGTomsheckAGearyBSpakowiczDStrobelS 2010 Endophyte Strain NRRL - 50072 producing volatile organics is a species of Ascocoryne. Mycology: An International Journal on Fungal Biology 1 187 doi:10.1080/21501203.2010.510122
7. StrobelGAKnightonBKluckKRenYLivinghouseT 2010 The production of myco-diesel hydrocarbons and their derivatives by the endophytic fungus Gliocladium roseum (NRRL 50072). Microbiology 156 3830 3833 doi:10.1099/mic.0.30824-0
8. FortmanJLChhabraSMukhopadhyayAChouHLeeTS 2008 Biofuel alternatives to ethanol: pumping the microbial well. Trends Biotechnol 26 375 381 doi:10.1016/j.tibtech.2008.03.008
9. BellerHRGohE-BKeaslingJD 2010 Genes Involved in Long-Chain Alkene Biosynthesis in Micrococcus luteus. Appl Environ Microbiol 76 1212 1223 doi:10.1128/AEM.02312-09
10. SukovichDJSeffernickJLRichmanJEHuntKAGralnickJA 2010 Structure, function, and insights into the biosynthesis of a head-to-head hydrocarbon in Shewanella oneidensis strain MR-1. Appl Environ Microbiol 76 3842 3849 doi:10.1128/AEM.00433-10
11. SchirmerARudeMALiXPopovaEdel CardayreSB 2010 Microbial Biosynthesis of Alkanes. Science 329 559 562 doi:10.1126/science.1187936
12. RudeMABaronTSBrubakerSAlibhaiMDel CardayreSB 2011 Terminal Olefin (1-Alkene) Biosynthesis by a Novel P450 Fatty Acid Decarboxylase from Jeotgalicoccus Species. Appl Environ Microbiol 77 1718 1727 doi:10.1128/AEM.02580-10
13. TresslRBahriDEngelKH 1981 Lipid oxidation in fruits and vegetables. p. Available: 10.1021/bk-1981-0170.ch016
14. TresslRBahriDEngelKH 1982 Formation of eight-carbon and ten-carbon components in mushrooms (Agaricus campestris). Journal of Agricultural and Food Chemistry 30 89 93
15. WurzenbergerMGroschW 1984 The formation of 1-octen-3-ol from the 10-hydroperoxide isomer of linoleic acid by a hydroperoxide lyase in mushrooms (Psalliota bispora). Biochimica et Biophysica Acta (BBA)-Lipids and Lipid Metabolism 794 25 30
16. WurzenbergerMGroschW 1984 Stereochemistry of the cleavage of the 10-hydroperoxide isomer of linoleic acid to 1-octen-3-ol by a hydroperoxide lyase from mushrooms (psalliota bispora). Biochimica et Biophysica Acta (BBA) - Lipids and Lipid Metabolism 795 163 165 doi:10.1016/0005-2760(84)90117-6
17. BrodhunFSchneiderSGöbelCHornungEFeussnerI 2010 PpoC from Aspergillus nidulans is a fusion protein with only one active haem. Biochem J 425 553 565 doi:10.1042/BJ20091096
18. AskenaziMDriggersEMHoltzmanDANormanTCIversonS 2003 Integrating transcriptional and metabolite profiles to direct the engineering of lovastatin-producing fungal strains. Nat Biotech 21 150 156 doi:10.1038/nbt781
19. BradleyPHBrauerMJRabinowitzJDTroyanskayaOG 2009 Coordinated concentration changes of transcripts and metabolites in Saccharomyces cerevisiae. PLoS Comput Biol 5 e1000270 doi:10.1371/journal.pcbi.1000270
20. RedestigHCostaIG 2011 Detection and interpretation of metabolite–transcript coresponses using combined profiling data. Bioinformatics 27 i357 i365 doi:10.1093/bioinformatics/btr231
21. HiraiMYYanoMGoodenoweDBKanayaSKimuraT 2004 Integration of transcriptomics and metabolomics for understanding of global responses to nutritional stresses in Arabidopsis thaliana. Proceedings of the National Academy of Sciences of the United States of America 101 10205 10210 doi:10.1073/pnas.0403218101
22. HancockTTakigawaIMamitsukaH 2010 Mining metabolic pathways through gene expression. Bioinformatics 26 2128 2135 doi:10.1093/bioinformatics/btq344
23. SaitoNOhashiYSogaTTomitaM 2010 Unveiling cellular biochemical reactions via metabolomics-driven approaches. Curr Opin Microbiol 13 358 362 doi:10.1016/j.mib.2010.04.006
24. MarguliesMEgholmMAltmanWEAttiyaSBaderJS 2005 Genome sequencing in microfabricated high-density picolitre reactors. Nature 437 376 380 doi:10.1038/nature03959
25. FitzpatrickDLogueMStajichJButlerG 2006 A fungal phylogeny based on 42 complete genomes derived from supertree and combined gene analysis. BMC Evolutionary Biology 6 99 doi:10.1186/1471-2148-6-99
26. ParraGBradnamKKorfI 2007 CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23 1061 1067 doi:10.1093/bioinformatics/btm071
27. BensonDAKarsch-MizrachiILipmanDJOstellJWheelerDL 2007 GenBank. Nucleic Acids Research 36 D25 D30 doi:10.1093/nar/gkm929
28. NagalakshmiUWangZWaernKShouCRahaD 2008 The Transcriptional Landscape of the Yeast Genome Defined by RNA Sequencing. Science 320 1344 1349 doi:10.1126/science.1158441
29. MortazaviAWilliamsBAMcCueKSchaefferLWoldB 2008 Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Meth 5 621 628 doi:10.1038/nmeth.1226
30. Supelco Bulletin 869A n.d. Solid Phase Microextraction: Solventless Sample Preparation for Monitoring Flavor Compounds by Capillary Gas Chromatography. Available: www.sigmaaldrich.com/etc/medialib/docs/Supelco//4524.pdf
31. OzsolakFKapranovPFoissacSKimSWFishilevichE 2010 Comprehensive Polyadenylation Site Maps in Yeast and Human Reveal Pervasive Alternative Polyadenylation. Cell 143 1018 1029 doi:10.1016/j.cell.2010.11.020
32. BumgarnerSLDowellRDGrisafiPGiffordDKFinkGR 2009 Toggle involving cis-interfering noncoding RNAs controls variegated gene expression in yeast. Proceedings of the National Academy of Sciences 106 18321 18326 doi:10.1073/pnas.0909641106
33. GersteinMBLuZJVan NostrandELChengCArshinoffBI 2010 Integrative Analysis of the Caenorhabditis elegans Genome by the modENCODE Project. Science 330 1775 1787 doi:10.1126/science.1196914
34. CantarelBLCoutinhoPMRancurelCBernardTLombardV 2009 The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res 37 D233 238 doi:10.1093/nar/gkn663
35. SaloheimoMPaloheimoMHakolaSPereJSwansonB 2002 Swollenin, a Trichoderma reesei protein with sequence similarity to the plant expansins, exhibits disruption activity on cellulosic materials. European Journal of Biochemistry 269 4202 4211 doi:10.1046/j.1432-1033.2002.03095.x
36. ChenX-aiIshidaNTodakaNNakamuraRMaruyamaJ-ichi 2010 Promotion of Efficient Saccharification of Crystalline Cellulose by Aspergillus fumigatus Swo1. Appl Environ Microbiol 76 2556 2561 doi:10.1128/AEM.02499-09
37. FischerCRKlein-MarcuschamerDStephanopoulosG 2008 Selection and optimization of microbial hosts for biofuels production. Metabolic Engineering 10 295 304 doi:10.1016/j.ymben.2008.06.009
38. JamesEGSarahECChristina CuomoLJJenniferRWSerafim BatzoglouSI 2005 Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438 1105 1115
39. MartinezDBerkaRMHenrissatBSaloheimoMArvasM 2008 Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nature biotechnology 26 553 560
40. CameraSLBalaguéCGöbelCGeoffroyPLegrandM 2009 The Arabidopsis Patatin-Like Protein 2 (PLP2) Plays an Essential Role in Cell Death Execution and Differentially Affects Biosynthesis of Oxylipins and Resistance to Pathogens. MPMI 22 469 481 doi:10.1094/MPMI-22-4-0469
41. Del SorboGSchoonbeekHDe WaardMA 2000 Fungal transporters involved in efflux of natural toxic compounds and fungicides. Fungal Genet Biol 30 1 15
42. YadavGGokhaleRSMohantyD 2009 Towards Prediction of Metabolic Products of Polyketide Synthases: An In Silico Analysis. PLoS Comput Biol 5 e1000351 doi:10.1371/journal.pcbi.1000351
43. BouhiredSWeberMKempf-SontagAKellerNPHoffmeisterD 2007 Accurate prediction of the Aspergillus nidulans terrequinone gene cluster boundaries using the transcriptional regulator LaeA. Fungal Genetics and Biology 44 1134 1145 doi:10.1016/j.fgb.2006.12.010
44. YadavGGokhaleRSMohantyD 2003 SEARCHPKS: a program for detection and analysis of polyketide synthase domains. Nucleic Acids Research 31 3654 3658 doi:10.1093/nar/gkg607
45. PielJ 2010 Biosynthesis of polyketides by trans-AT polyketide synthases. Nat Prod Rep 27 996 1047 doi:10.1039/b816430b
46. KooninEVWolfYI 2008 Genomics of bacteria and archaea: the emerging dynamic view of the prokaryotic world. Nucleic Acids Research 36 6688 6719 doi:10.1093/nar/gkn668
47. PellegriniMMarcotteEMThompsonMJEisenbergDYeatesTO 1999 Assigning protein functions by comparative genome analysis: Protein phylogenetic profiles. Proc Natl Acad Sci U S A 96 4285 4288
48. KorbelJOJensenLJvon MeringCBorkP 2004 Analysis of genomic context: prediction of functional associations from conserved bidirectionally transcribed gene pairs. Nat Biotech 22 911 917 doi:10.1038/nbt988
49. MarcotteEMPellegriniMNgH-LRiceDWYeatesTO 1999 Detecting Protein Function and Protein-Protein Interactions from Genome Sequences. Science 285 751 753 doi:10.1126/science.285.5428.751
50. OverbeekRFonsteinMD'SouzaMPuschGDMaltsevN 1999 The use of gene clusters to infer functional coupling. Proceedings of the National Academy of Sciences 96 2896 2901 doi:10.1073/pnas.96.6.2896
51. KumarCGEvertsRELoorJJLewinHA 2010 Functional annotation of novel lineage-specific genes using co-expression and promoter analysis. BMC Genomics 11 161 doi:10.1186/1471-2164-11-161
52. StephanopoulosG 2007 Challenges in Engineering Microbes for Biofuels Production. Science 315 801 804 doi:10.1126/science.1139612
53. LeeSKChouHHamTSLeeTSKeaslingJD 2008 Metabolic engineering of microorganisms for biofuels production: from bugs to synthetic biology to fuels. Current Opinion in Biotechnology 19 556 563 doi:10.1016/j.copbio.2008.10.014
54. AndreouABrodhunFFeussnerI 2009 Biosynthesis of oxylipins in non-mammals. Progress in Lipid Research 48 148 170 doi:10.1016/j.plipres.2009.02.002
55. GrechkinANHambergM 2004 The “heterolytic hydroperoxide lyase” is an isomerase producing a short-lived fatty acid hemiacetal. Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids 1636 47 58 doi:10.1016/j.bbalip.2003.12.003
56. BerdyJ 2005 Bioactive Microbial Metabolites. J Antibiot 58 1 26
57. DoddsDRGrossRA 2007 Chemicals from Biomass. Science 318 1250 1251 doi:10.1126/science.1146356
58. CroesDCoucheFWodakSJvan HeldenJ 2005 Metabolic PathFinding: inferring relevant pathways in biochemical networks. Nucleic Acids Research 33 W326 W330 doi:10.1093/nar/gki437
59. GaoJEllisLBMWackettLP 2010 The University of Minnesota Biocatalysis/Biodegradation Database: improving public access. Nucleic Acids Res 38 D488 D491 doi:10.1093/nar/gkp771
60. MoriyaYShigemizuDHattoriMTokimatsuTKoteraM 2010 PathPred: an enzyme-catalyzed metabolic pathway prediction server. Nucleic Acids Res 38 W138 143 doi:10.1093/nar/gkq318
61. RahmanSAAdvaniPSchunkRSchraderRSchomburgD 2005 Metabolic pathway analysis web service (Pathway Hunter Tool at CUBIC). Bioinformatics 21 1189 1193 doi:10.1093/bioinformatics/bti116
62. CaspiRFoersterHFulcherCAKaipaPKrummenackerM 2008 The MetaCyc Database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases. Nucleic Acids Res 36 D623 D631 doi:10.1093/nar/gkm900
63. WinkM 1988 Plant breeding: importance of plant secondary metabolites for protection against pathogens and herbivores. Theoret Appl Genetics 75 225 233 doi:10.1007/BF00303957
64. WielochW 2006 Chromosome visualisation in filamentous fungi. J Microbiol Methods 67 1 8 doi:10.1016/j.mimet.2006.05.022
65. GriffinMASpakowiczDJGianoulisTAStrobelSA 2010 Volatile organic compound production by organisms in the genus Ascocoryne and a re-evaluation of myco-diesel production by NRRL 50072. Microbiology 156 3814
66. AltschulSFGishWMillerWMyersEWLipmanDJ 1990 Basic local alignment search tool. J Mol Biol 215 403 410 doi:10.1016/S0022-2836(05)80360-2
67. CantarelBLCoutinhoPMRancurelCBernardTLombardV 2009 The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res 37 D233 238 doi:10.1093/nar/gkn663
68. EddySR 2009 A new generation of homology search tools based on probabilistic inference. Genome Inform 23 205 211
69. AlamIHubbardSOliverSRattrayM 2007 A kingdom-specific protein domain HMM library for improved annotation of fungal genomes. BMC Genomics 8 97 doi:10.1186/1471-2164-8-97
70. FinnRDTateJMistryJCoggillPCSammutSJ 2007 The Pfam protein families database. Nucleic Acids Research 36 D281 D288 doi:10.1093/nar/gkm960
71. KanehisaMArakiMGotoSHattoriMHirakawaM 2008 KEGG for linking genomes to life and the environment. Nucleic Acids Res 36 D480 484 doi:10.1093/nar/gkm882
72. MoriyaYItohMOkudaSYoshizawaACKanehisaM 2007 KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35 W182 185 doi:10.1093/nar/gkm321
73. AshburnerMBallCABlakeJABotsteinDButlerH 2000 Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25 25 29 doi:10.1038/75556
74. ArnaudMBChibucosMCCostanzoMCCrabtreeJInglisDO 2010 The Aspergillus Genome Database, a curated comparative genomics resource for gene, protein and sequence information for the Aspergillus research community. Nucleic Acids Res 38 D420 D427 doi:10.1093/nar/gkp751
75. LangmeadBTrapnellCPopMSalzbergSL 2009 Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10 R25 doi:10.1186/gb-2009-10-3-r25
76. HabeggerLSbonerAGianoulisTARozowskyJAgarwalA 2010 RSEQtools: A modular framework to analyze RNA-Seq data using compact, anonymized data summaries. Bioinformatics. Available: http://bioinformatics.oxfordjournals.org/content/early/2010/12/05/bioinformatics.btq643.abstract
77. KentWJ 2002 BLAT–the BLAST-like alignment tool. Genome Res 12 656 664 doi:10.1101/gr.229202
Štítky
Genetika Reprodukční medicína
Článek Physiological Notch Signaling Maintains Bone Homeostasis via RBPjk and Hey Upstream of NFATc1Článek Intronic -Regulatory Modules Mediate Tissue-Specific and Microbial Control of / TranscriptionČlánek Probing the Informational and Regulatory Plasticity of a Transcription Factor DNA–Binding DomainČlánek Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin ComplexesČlánek An Alu Element–Associated Hypermethylation Variant of the Gene Is Associated with Childhood ObesityČlánek Three Essential Ribonucleases—RNase Y, J1, and III—Control the Abundance of a Majority of mRNAsČlánek Genomic Tools for Evolution and Conservation in the Chimpanzee: Is a Genetically Distinct Population
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2012 Číslo 3
-
Všechny články tohoto čísla
- Comprehensive Research Synopsis and Systematic Meta-Analyses in Parkinson's Disease Genetics: The PDGene Database
- Genomic Analysis of the Hydrocarbon-Producing, Cellulolytic, Endophytic Fungus
- Networks of Neuronal Genes Affected by Common and Rare Variants in Autism Spectrum Disorders
- Akirin Links Twist-Regulated Transcription with the Brahma Chromatin Remodeling Complex during Embryogenesis
- Too Much Cleavage of Cyclin E Promotes Breast Tumorigenesis
- Imprinted Genes … and the Number Is?
- Genetic Architecture of Highly Complex Chemical Resistance Traits across Four Yeast Strains
- Exploring the Complexity of the HIV-1 Fitness Landscape
- MNS1 Is Essential for Spermiogenesis and Motile Ciliary Functions in Mice
- A Fundamental Regulatory Mechanism Operating through OmpR and DNA Topology Controls Expression of Pathogenicity Islands SPI-1 and SPI-2
- Evidence for Positive Selection on a Number of MicroRNA Regulatory Interactions during Recent Human Evolution
- Variation in Modifies Risk of Neonatal Intestinal Obstruction in Cystic Fibrosis
- PIF4–Mediated Activation of Expression Integrates Temperature into the Auxin Pathway in Regulating Hypocotyl Growth
- Critical Evaluation of Imprinted Gene Expression by RNA–Seq: A New Perspective
- A Meta-Analysis and Genome-Wide Association Study of Platelet Count and Mean Platelet Volume in African Americans
- Mouse Genetics Suggests Cell-Context Dependency for Myc-Regulated Metabolic Enzymes during Tumorigenesis
- Transcriptional Control in Cardiac Progenitors: Tbx1 Interacts with the BAF Chromatin Remodeling Complex and Regulates
- Synthetic Lethality of Cohesins with PARPs and Replication Fork Mediators
- APOBEC3G-Induced Hypermutation of Human Immunodeficiency Virus Type-1 Is Typically a Discrete “All or Nothing” Phenomenon
- Interpreting Meta-Analyses of Genome-Wide Association Studies
- Error-Prone ZW Pairing and No Evidence for Meiotic Sex Chromosome Inactivation in the Chicken Germ Line
- -Dependent Chemosensory Functions Contribute to Courtship Behavior in
- Diverse Forms of Splicing Are Part of an Evolving Autoregulatory Circuit
- Phenotypic Plasticity of the Drosophila Transcriptome
- Physiological Notch Signaling Maintains Bone Homeostasis via RBPjk and Hey Upstream of NFATc1
- Precocious Metamorphosis in the Juvenile Hormone–Deficient Mutant of the Silkworm,
- Igf1r Signaling Is Indispensable for Preimplantation Development and Is Activated via a Novel Function of E-cadherin
- Accurate Prediction of Inducible Transcription Factor Binding Intensities In Vivo
- Mitochondrial Oxidative Stress Alters a Pathway in Strongly Resembling That of Bile Acid Biosynthesis and Secretion in Vertebrates
- Mammalian Neurogenesis Requires Treacle-Plk1 for Precise Control of Spindle Orientation, Mitotic Progression, and Maintenance of Neural Progenitor Cells
- Tcf7 Is an Important Regulator of the Switch of Self-Renewal and Differentiation in a Multipotential Hematopoietic Cell Line
- REST–Mediated Recruitment of Polycomb Repressor Complexes in Mammalian Cells
- Intronic -Regulatory Modules Mediate Tissue-Specific and Microbial Control of / Transcription
- Age-Dependent Brain Gene Expression and Copy Number Anomalies in Autism Suggest Distinct Pathological Processes at Young Versus Mature Ages
- A Genome-Wide Association Study Identifies Variants Underlying the Shade Avoidance Response
- -by- Regulatory Divergence Causes the Asymmetric Lethal Effects of an Ancestral Hybrid Incompatibility Gene
- Genome-Wide Association and Functional Follow-Up Reveals New Loci for Kidney Function
- A Natural System of Chromosome Transfer in
- Cell Size and the Initiation of DNA Replication in Bacteria
- Probing the Informational and Regulatory Plasticity of a Transcription Factor DNA–Binding Domain
- Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin Complexes
- Temporal Transcriptional Profiling of Somatic and Germ Cells Reveals Biased Lineage Priming of Sexual Fate in the Fetal Mouse Gonad
- Rapid Analysis of Genome Rearrangements by Multiplex Ligation–Dependent Probe Amplification
- Metabolic Profiling of a Mapping Population Exposes New Insights in the Regulation of Seed Metabolism and Seed, Fruit, and Plant Relations
- The Atypical Calpains: Evolutionary Analyses and Roles in Cellular Degeneration
- The Silkworm Coming of Age—Early
- Development of a Panel of Genome-Wide Ancestry Informative Markers to Study Admixture Throughout the Americas
- Balanced Codon Usage Optimizes Eukaryotic Translational Efficiency
- The Min System and Nucleoid Occlusion Are Not Required for Identifying the Division Site in but Ensure Its Efficient Utilization
- Neurobeachin, a Regulator of Synaptic Protein Targeting, Is Associated with Body Fat Mass and Feeding Behavior in Mice and Body-Mass Index in Humans
- Statistical Analysis of Readthrough Levels for Nonsense Mutations in Mammalian Cells Reveals a Major Determinant of Response to Gentamicin
- Gene Reactivation by 5-Aza-2′-Deoxycytidine–Induced Demethylation Requires SRCAP–Mediated H2A.Z Insertion to Establish Nucleosome Depleted Regions
- The miR-35-41 Family of MicroRNAs Regulates RNAi Sensitivity in
- Genetic Basis of Hidden Phenotypic Variation Revealed by Increased Translational Readthrough in Yeast
- An Alu Element–Associated Hypermethylation Variant of the Gene Is Associated with Childhood Obesity
- Modelling Human Regulatory Variation in Mouse: Finding the Function in Genome-Wide Association Studies and Whole-Genome Sequencing
- Novel Loci for Adiponectin Levels and Their Influence on Type 2 Diabetes and Metabolic Traits: A Multi-Ethnic Meta-Analysis of 45,891 Individuals
- Polycomb-Like 3 Promotes Polycomb Repressive Complex 2 Binding to CpG Islands and Embryonic Stem Cell Self-Renewal
- Insulin/IGF-1 and Hypoxia Signaling Act in Concert to Regulate Iron Homeostasis in
- EMF1 and PRC2 Cooperate to Repress Key Regulators of Arabidopsis Development
- Three Essential Ribonucleases—RNase Y, J1, and III—Control the Abundance of a Majority of mRNAs
- Contrasted Patterns of Molecular Evolution in Dominant and Recessive Self-Incompatibility Haplotypes in
- A Machine Learning Approach for Identifying Novel Cell Type–Specific Transcriptional Regulators of Myogenesis
- Genomic Tools for Evolution and Conservation in the Chimpanzee: Is a Genetically Distinct Population
- Nos2 Inactivation Promotes the Development of Medulloblastoma in Mice by Deregulation of Gap43–Dependent Granule Cell Precursor Migration
- Intracranial Aneurysm Risk Locus 5q23.2 Is Associated with Elevated Systolic Blood Pressure
- Heritability and Genetic Correlations Explained by Common SNPs for Metabolic Syndrome Traits
- A Genome-Wide Association Study of Nephrolithiasis in the Japanese Population Identifies Novel Susceptible Loci at 5q35.3, 7p14.3, and 13q14.1
- DNA Damage in Nijmegen Breakage Syndrome Cells Leads to PARP Hyperactivation and Increased Oxidative Stress
- DNA Resection at Chromosome Breaks Promotes Genome Stability by Constraining Non-Allelic Homologous Recombination
- Genetic Analysis of Floral Symmetry in Van Gogh's Sunflowers Reveals Independent Recruitment of Genes in the Asteraceae
- A Splice Site Variant in the Bovine Gene Compromises Growth and Regulation of the Inflammatory Response
- Promoter Nucleosome Organization Shapes the Evolution of Gene Expression
- The Nucleoside Diphosphate Kinase Gene Acts as Quantitative Trait Locus Promoting Non-Mendelian Inheritance
- The Ciliogenic Transcription Factor RFX3 Regulates Early Midline Distribution of Guidepost Neurons Required for Corpus Callosum Development
- Phosphorylation of the RNA–Binding Protein HOW by MAPK/ERK Enhances Its Dimerization and Activity
- A Genome-Wide Scan of Ashkenazi Jewish Crohn's Disease Suggests Novel Susceptibility Loci
- Parkinson's Disease–Associated Kinase PINK1 Regulates Miro Protein Level and Axonal Transport of Mitochondria
- LMW-E/CDK2 Deregulates Acinar Morphogenesis, Induces Tumorigenesis, and Associates with the Activated b-Raf-ERK1/2-mTOR Pathway in Breast Cancer Patients
- Mapping the Hsp90 Genetic Interaction Network in Reveals Environmental Contingency and Rewired Circuitry
- Autoregulation of the Noncoding RNA Gene
- The Human Pancreatic Islet Transcriptome: Expression of Candidate Genes for Type 1 Diabetes and the Impact of Pro-Inflammatory Cytokines
- Spo0A∼P Imposes a Temporal Gate for the Bimodal Expression of Competence in
- Antagonistic Regulation of Apoptosis and Differentiation by the Cut Transcription Factor Represents a Tumor-Suppressing Mechanism in
- A Downstream CpG Island Controls Transcript Initiation and Elongation and the Methylation State of the Imprinted Macro ncRNA Promoter
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- PIF4–Mediated Activation of Expression Integrates Temperature into the Auxin Pathway in Regulating Hypocotyl Growth
- Metabolic Profiling of a Mapping Population Exposes New Insights in the Regulation of Seed Metabolism and Seed, Fruit, and Plant Relations
- A Splice Site Variant in the Bovine Gene Compromises Growth and Regulation of the Inflammatory Response
- Comprehensive Research Synopsis and Systematic Meta-Analyses in Parkinson's Disease Genetics: The PDGene Database
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



