-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaTelomerase-Null Survivor Screening Identifies Novel Telomere Recombination Regulators
Telomeres are protein–DNA structures found at the ends of linear chromosomes and are crucial for genome integrity. Telomeric DNA length is primarily maintained by the enzyme telomerase. Cells lacking telomerase will undergo senescence when telomeres become critically short. In Saccharomyces cerevisiae, a very small percentage of cells lacking telomerase can remain viable by lengthening telomeres via two distinct homologous recombination pathways. These “survivor” cells are classified as either Type I or Type II, with each class of survivor possessing distinct telomeric DNA structures and genetic requirements. To elucidate the regulatory pathways contributing to survivor generation, we knocked out the telomerase RNA gene TLC1 in 280 telomere-length-maintenance (TLM) gene mutants and examined telomere structures in post-senescent survivors. We uncovered new functional roles for 10 genes that affect the emerging ratio of Type I versus Type II survivors and 22 genes that are required for Type II survivor generation. We further verified that Pif1 helicase was required for Type I recombination and that the INO80 chromatin remodeling complex greatly affected the emerging frequency of Type I survivors. Finally, we found the Rad6-mediated ubiquitination pathway and the KEOPS complex were required for Type II recombination. Our data provide an independent line of evidence supporting the idea that these genes play important roles in telomere dynamics.
Published in the journal: . PLoS Genet 9(1): e32767. doi:10.1371/journal.pgen.1003208
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1003208Summary
Telomeres are protein–DNA structures found at the ends of linear chromosomes and are crucial for genome integrity. Telomeric DNA length is primarily maintained by the enzyme telomerase. Cells lacking telomerase will undergo senescence when telomeres become critically short. In Saccharomyces cerevisiae, a very small percentage of cells lacking telomerase can remain viable by lengthening telomeres via two distinct homologous recombination pathways. These “survivor” cells are classified as either Type I or Type II, with each class of survivor possessing distinct telomeric DNA structures and genetic requirements. To elucidate the regulatory pathways contributing to survivor generation, we knocked out the telomerase RNA gene TLC1 in 280 telomere-length-maintenance (TLM) gene mutants and examined telomere structures in post-senescent survivors. We uncovered new functional roles for 10 genes that affect the emerging ratio of Type I versus Type II survivors and 22 genes that are required for Type II survivor generation. We further verified that Pif1 helicase was required for Type I recombination and that the INO80 chromatin remodeling complex greatly affected the emerging frequency of Type I survivors. Finally, we found the Rad6-mediated ubiquitination pathway and the KEOPS complex were required for Type II recombination. Our data provide an independent line of evidence supporting the idea that these genes play important roles in telomere dynamics.
Introduction
Telomeres are special DNA-protein structures found at the ends of eukaryotic chromosomes. Telomeres are crucial for genome integrity because they prevent chromosome ends from degradation or fusing with each other [1]. In budding yeast Saccharomyces cerevisiae, telomeric DNA consists of ∼350 base pairs (bp) of TG1–3/C1–3 A repeats with a terminal single-stranded TG1–3 tract called a G-overhang [2]. Telomeric DNA can be maintained by either telomerase-mediated elongation or homologous recombination [3]–[5]. Telomerase is a highly specialized reverse transcriptase that adds telomeric DNA sequences to the 3′ G-overhang using its intrinsic RNA template [3]. In Saccharomyces cerevisiae, the core components of telomerase are the catalytic subunit Est2 and its RNA template subunit TLC1 [6], [7]. In wild-type yeast cells, the telomerase pathway supercedes the recombination pathway as the predominant mechanism of telomeric DNA elongation [8], [9]. In telomerase-null cells, telomeric DNA is maintained via a recombination pathway termed “alternative lengthening of telomeres” (ALT) [10]. Approximately 85% of immortalized human tumor cells use telomerase to maintain telomeres while 15% apply the ALT mechanism to maintain telomeres [11].
In telomerase-null S. cerevisiae mutants, most cells undergo senescence after about 50–100 divisions when telomeres shorten to less than approximately 100 bp [7], [12], [13]. Surprisingly, a select few of these senescing cells are able to bypass the short telomere survival crisis through lengthening their telomeres via a Rad52-dependent recombination pathway [14]. These cells are called post-senescence survivors or “survivors” for short [14]. Survivors are categorized into two types: Type I and Type II, which possess different telomeric DNA structures and are defined by their dependence on Rad51 or Rad50 respectively [15]. Type I survivors exhibit highly amplified subtelomeric Y' elements and short terminal telomeric TG tracts. The formation of Type I survivors depends on the canonical homologous recombination proteins Rad51, Rad54, Rad55 and Rad57 [14]. On the other hand, Type II survivors have long heterogeneous terminal telomeric TG tracts generated by recombination, and their formation depends on the Mre11-Rad50-Xrs2 (MRX) complex and Rad59 [14]. Type II survivors resemble the ALT cells observed in mammals [5]. In S. cerevisiae, about 90% of survivors generated on solid medium are categorized as Type I, while 10% are Type II. Nevertheless, Type II survivors grow at faster rates than Type I survivors, eventually overtaking their counterparts in liquid-grown cultures [14].
In addition to the proteins in the Rad52 epistasis group, which are well-defined in the canonical survivor formation pathways, other genes involved in survivor formation have sporadically been identified. For example, SGS1, MEC1/TEL1, MDT1, DEF1, CLB2 and SUA5 are required for the generation of Type II survivors, while RIF1 and RIF2 have strong influences toward Type I survivor emerging frequency [16]–[22]. Notably, some of the genes mentioned above appear to contribute to both survivor generation and telomere length regulation. Deletion of RIF1 or RIF2 causes telomere lengthening, while deletion of MRE11, RAD50, XRS2, TEL1, DEF1 or SUA5 results in telomere shortening [16], [23]–[25]. These observations suggest that genes involved in telomere recombination pathways and telomere length regulation are in some way linked. So far, there have been 251 telomere length maintenance (TLM) genes identified by genome-wide screens [23], [26] and other studies [16], [27]–[34]. Furthermore, 29 additional genes previously miss-classified as essential genes in the Saccharomyces genome deletion project have now officially been implicated as TLM genes [24]. In this study we deleted the TLC1 gene encoding the RNA template subunit of telomerase in each of these 280 TLM mutants. We then examined the survivor types that arose and in doing so we were able to identify novel regulators that contribute to telomere recombination. The genes we characterized as telomere recombination regulators may also affect general DNA recombination at other genomic loci.
Results
Screening of TLM gene deletion library on solid medium identifies genes affecting the emerging ratio of Type I versus Type II survivors
To search for genes affecting survivor formation, we knocked out the RNA component of telomerase TLC1 in 280 haploid TLM mutants reported to have longer or shorter telomeres than the wild-type strain [23], [24], [26] (Table S1). Knocking out TLC1 in most TLM mutants is typically achieved by transformation of an integrating plasmid but for some strains with extremely short telomeres or severe growth defects, recovering a TLC1 deletion clone using this approach was not possible. For such cases, we mated tlc1Δ mutant (BY4741 background) with tlmΔ mutants (BY4742 background) to generate heterozygous diploid strains, and then performed tetrad dissection to obtain haploid mutants lacking both TLC1 and TLM genes (Table S1).
After a telomerase-null tlmΔ mutant library was established, each mutant was passaged repeatedly on solid plates to screen for genes that might affect Type I survivor formation. Most of the mutant cells underwent senescence but a small percentage of cells were able to overcome crisis and became survivors [5]. Genomic DNA was extracted from each survival isolate, digested with the XhoI restriction enzyme, and analyzed by Southern blot with a TG probe to determine if the cells were Type I or Type II survivors (Figure 1A) (see Materials and Methods). In the first round of screening for genes affecting Type I survivor formation, we passaged two independent senescing colonies from each mutant on solid plates to obtain survivors. Because the emerging frequency of Type I survivors (∼90%) is much higher than that of Type II survivors (∼10%), most double mutants passaged on a solid plate, like the tlc1Δ single mutant, turned out to be Type I survivors [5]. However, if both of the two colonies picked from a single mutant strain had telomere structures consistent with that of Type II survivors, it was concluded that the gene missing in this Type II strain might contribute to Type I survivor generation and should be analyzed further. For each tlc1Δ tlmΔ mutant selected in this first round of rough screening, eight single colonies were passaged on solid plates in the second round of screening until survivors arose. When more than four colonies became Type II survivors, this TLM gene was subjected to a third round of screening in which fifty colonies of the tlc1Δ tlmΔ mutant were passaged again on solid plates. From these fifty colonies at least forty colonies typically generated survivors that could be examined. The emerging frequency of Type II survivors in each strain was then calculated (Table 1). Using this screening approach we identified eleven mutants in which the emerging frequencies of Type II survivors was elevated significantly (Table 1). Among these eleven genes, RIF1 and RIF2 deletion in telomerase-null tlc1Δ mutant generated Type II frequencies of 52.2% and 85.7% respectively (Table 1, the column of “Deleting TLC1 in tlm mutants”), percentages which are consistent with a prior study performed by Teng et al. [22]. The other nine genes that affected survivor formation have never before been reported to have such a function. The Type II emerging frequencies in these nine mutants ranged from 45.7% to 93.6% (Table 1, the column of “Deleting TLC1 in tlm mutants”) and were significantly elevated compared to that of the tlc1Δ cells, which had a Type II emerging frequency of 4% (Figure 1B). In contrast with the eleven genes that affected Type I survivor generation, the PIF1, helicase gene [35], [36], appeared to be essential for Type I survivor generation (discussed later).
Fig. 1. Identification of genes affecting the emerging ratio of Type I versus Type II survivors. 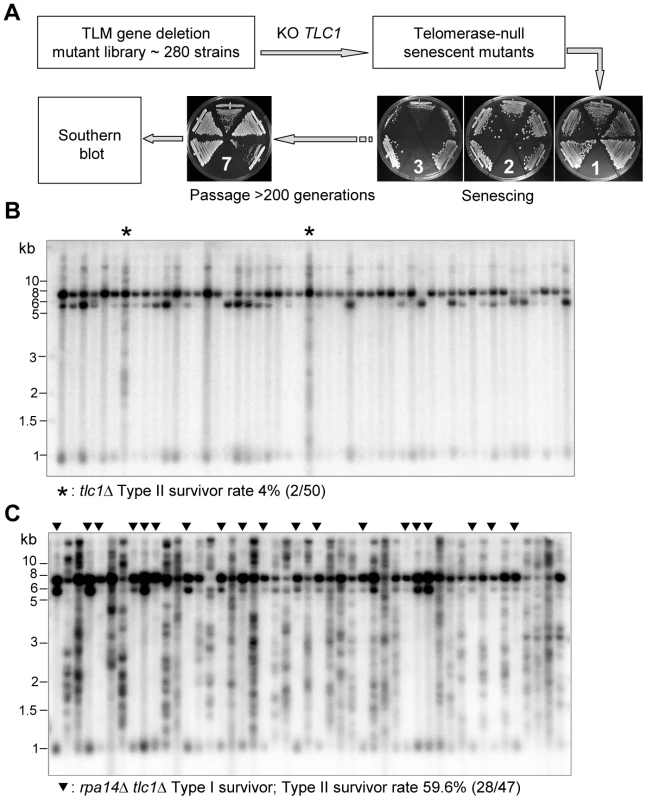
(A) Schematic illustration of the screening procedures for genes that affect the emerging ratio of Type I vs Type II survivors (refer to details in main text). The tlc1Δ single mutants and the rpa14Δ tlc1Δ double mutants were generated through tetrad dissection from heterozygous diploids with one copy of RPA14 and TLC1 deleted. Fifty independent colonies of each mutant were randomly selected and passaged on plates, and the telomere structures of survivors were examined by Southern blot using a TG probe. (B and C) The Southern blot analysis of survivor types in tlc1Δ strain (control) (B) and the rpa14Δ tlc1Δ mutant (C). The asterisks (*) in (B) indicate Type II survivors. The triangles (▾) in (C) indicate Type I survivors. Tab. 1. List of S. cerevisiae TLM genes affecting Type I versus II survivor ratio in tlc1Δ cells. 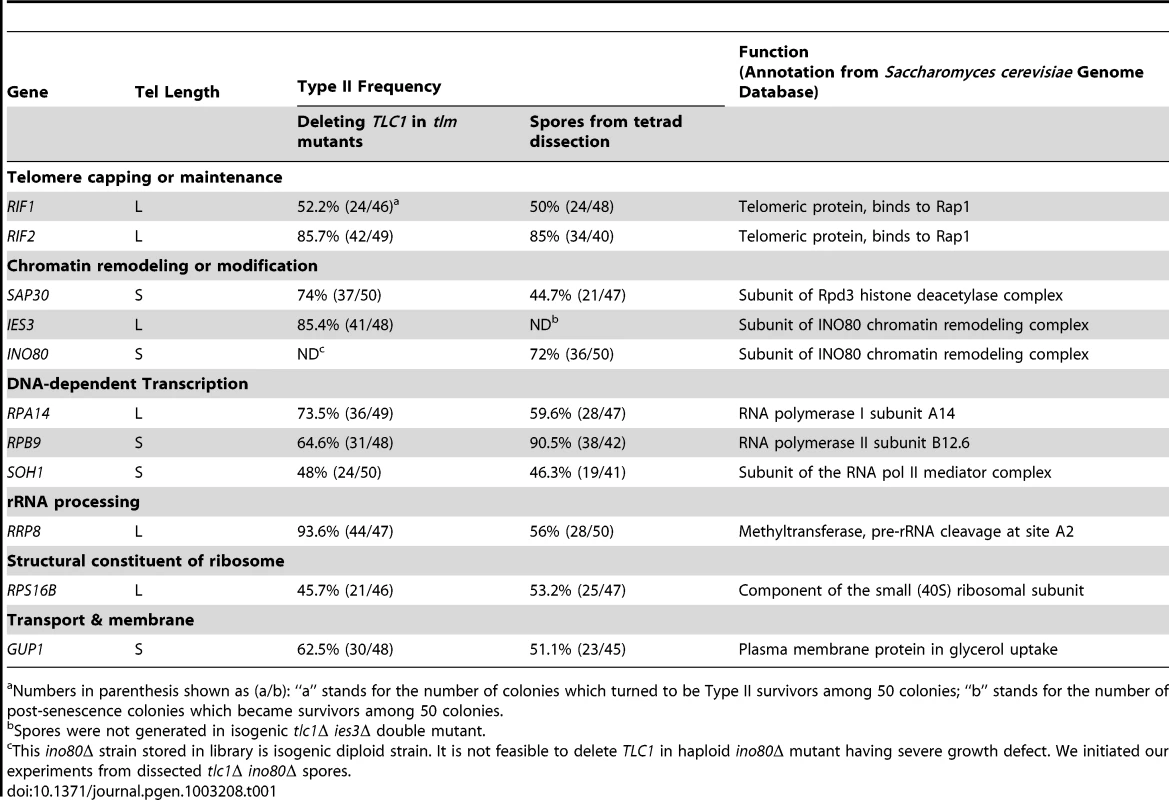
Numbers in parenthesis shown as (a/b): “a” stands for the number of colonies which turned to be Type II survivors among 50 colonies; “b” stands for the number of post-senescence colonies which became survivors among 50 colonies. Very recently, Chang et al. showed that the long telomeres in rif1Δ tlc1Δ and rif2Δ tlc1Δ mutants were preferentially extended by a recombination pathway and senescent cells with long telomeres were more efficient at bypassing senescence via the Type II survivor pathway [37]. These led Chang et al. to propose that rif1Δ tlc1Δ and rif2Δ tlc1Δ mutants affect the ratio of survivor types by altering telomere length at the point of senescence [37]. In order to examine the idea that telomere length affects the type of survivor generated, we generated eleven TLC1/tlc1Δ TLM/tlmΔ diploid strains and performed tetrad dissections to obtain tlc1Δ single and tlc1Δ tlmΔ double mutants (Table 1, the column of “Spore from tetrad dissection”). Because the ino80Δ tlc1Δ double mutant used in the previous experiments was obtained from tetrad dissection, it was not included in this experiment. Fifty senescing clones of the other ten mutant strains, including tlc1Δ single mutants from each diploid mutant, were streaked on plates until survivors arose. Telomere structures of the survivors generated on plates were examined by Southern blot (Figure S1). A representative Southern blot result of rpa14Δ tlc1Δ mutant is shown in Figure 1C. The results of these experiments are summarized below and are listed in the column of “Spore from tetrad dissection” in Table 1. The frequency of Type II survivor formation in the sap30Δ tlc1Δ, rpa14Δtlc1Δ, rrp8Δ tlc1Δ and gup1Δ tlc1Δ double mutants was decreased when compared to that of the corresponding double mutant that had not been through sporogenesis. The frequency of Type II survivor formation in the rpb9Δ tlc1Δ or rps16bΔ tlc1Δ double mutants was increased when compared to that of the corresponding double mutant that had not been through sporogenesis. The frequency of Type II survivor formation in rif1Δ tlc1Δ, rif2Δ tlc1Δ and soh1Δ tlc1Δ double mutants did not change significantly. Recovery of the ies3Δ tlc1Δ double mutant from sporogenesis was not successful. We also examined telomere length around the time of survivor formation and found that similar to the rif1Δ tlc1Δ and rif2Δ tlc1Δ mutants, the critical telomere length in gup1Δ tlc1Δ and ino80Δ tlc1Δ mutants was about 50 bp longer than those in tlc1Δ single mutants from the same crosses (Figure S2). However, in soh1Δ tlc1Δ and rpb9Δ tlc1Δ mutants, the critical telomere lengths were about 30 bp shorter than those in tlc1Δ mutants from the same crosses (Figure S2). Additionally, in the rps16bΔ tlc1Δ, sap30Δ tlc1Δ and rrp8Δ tlc1Δ mutants, the critical telomere lengths were slightly longer (<30 bp) than those in tlc1Δ mutants from the same crosses (Figure S2). In the rpa14Δ tlc1Δ mutant, the critical telomere length was similar to that in tlc1Δ mutant from the same cross (Figure S2). Our data support the idea put forth by Cheng et al. that telomere length affects survivor formation [37]. Our data also show the frequency of Type II emergence in the nine mutants we identified ranged from 44.7% to 90.5%, which was much higher than the Type II emerging frequencies of less than 10% that were usually observed in tlc1Δ cells (Table 1 and Figure S1) [5].
INO80 chromatin remodeling complex affects the emerging frequency of Type I survivor generation
The INO80 complex is one of the ATP-dependent chromatin remodeling complexes that can move or evict nucleosomes, thereby changing chromatin structure and affecting the accessibility of DNA to other factors [38]. The yeast INO80 complex contains multiple subunits, including five essential and ten (Ino80, Ies1, Ies2, Ies3, Ies4, Ies5, Ies6, Taf14, Arp8 and Nhp10) non-essential subunits [38]. A recent study has shown that Ies3 interacts with the telomerase component Est1 [34]. In est1Δ cells, deleting IES3 or ARP8 caused a delay of survivor generation in liquid culturing [34], suggesting that the INO80 complex affects telomere recombination. In our survivor screening we noted that two subunits in the INO80 complex, Ino80 and Ies3, significantly affected the generation of Type I survivors (Table 1). When passaged on solid medium, the ino80Δ tlc1Δ and ies3Δ tlc1Δ mutants produced Type II survivors at frequencies of 70% and 85.4% respectively (Figure 2A and 2B), which were significantly elevated in comparison with the 8.3% we observed in tlc1Δ cells (Figure 2B and Figure S3A). These results suggested that the INO80 complex may be required for efficient Type I survivor formation. To examine this possibility further we examined the impact of depleting each of the other four non-essential subunits of the INO80 complex on the efficiency of Type I survivor formation in tlc1Δ cells. The Southern blot results revealed that the deletion of each of the non-essential INO80 subunits IES1, IES4, IES5 and NHP10 led to the generation of more Type II than Type I survivors (Figure S3). The frequency of Type II emergence in each of these mutants in tlc1Δ cells was above 60% (Figure 2B), which was much higher than that of the tlc1Δ single mutant. These results indicate that the INO80 complex greatly influences the emerging ratio of Type I vs Type II survivors.
Fig. 2. The effect of the Ino80 complex on survivor formation. 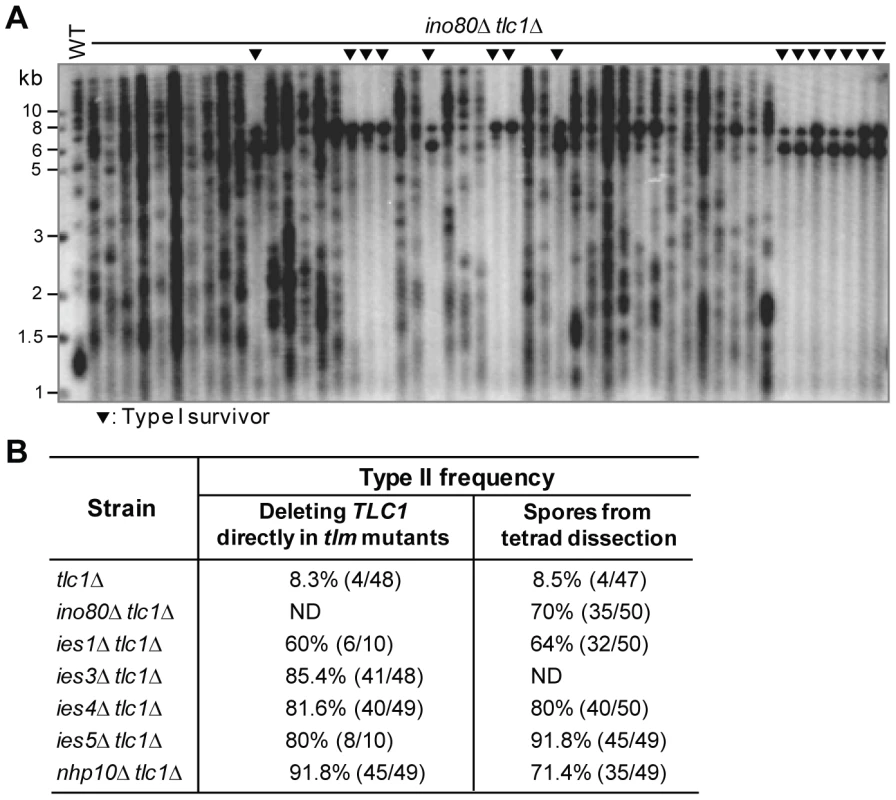
(A) Fifty independent survivor colonies of the ino80Δ tlc1Δ mutant, which was generated from INO80/ino80Δ TLC1/tlc1Δ diploid mutant, were randomly picked and their genomic DNA was isolated for Southern blot assay using a TG1–3 probe. The black triangles indicate Type I survivors. (B) Chart of Type II survivor frequencies in the mutants of Ino80 complex subunits. ND: not done (see Table 1). Pif1 is required for Type I recombination
PIF1 is a non-essential gene which encodes a 5′ to 3′ DNA and DNA/RNA helicase in S. cerevisiae [36], [39]. Previous studies have demonstrated that Pif1 can be translated from different start sites and has two forms which are localized to either the mitochondria or the nucleus [35], [40]. In the mitochondria Pif1 affects recombination of mitochondrial DNA (mtDNA) and plays an important role in maintaining mtDNA stability [41]–[43]. In the nucleus, Pif1 inhibits telomere lengthening by removing telomerase from telomeric DNA [35], [44] and participates in Okazaki fragment maturation [45], [46] and ribosomal DNA replication [47]. Additionally, Pif1 is able to unwind G-quadruplex structures in vitro [48], and likely acts on these structures in vivo as well [48], [49].
In our primary screening the pif1Δ tlc1Δ double mutant had difficulties generating survivors on solid medium, and as a result most clones died out during sequential streaks. The pif1Δ tlc1Δ clones that overcame senescence on solid medium showed a Type II survivor pattern (Figure 3A and 3B), suggesting that Pif1 promotes Type I survivor formation. To further validate the role of Pif1 in Type I survivor generation, we streaked fifty independent pif1Δ tlc1Δ colonies on plates. We noted that forty post-senescence colonies (80%) died during the sequential streaks, indicating that deletion of PIF1 in telomerase-null strains inhibits the creation of post-senescence survivors. The other 10 colonies also underwent senescence, but were able to generate survivors at the 7th streaking. Cells at this stage were harvested, and their telomeres were examined by Southern blot assay (Figure 3C). Only two colonies (4%), which grew at a normal rate, gave rise to type II survivors (Figure 3C and 3D), indicating that type II survivors can indeed form in the absence of Pif1. Interestingly, eight colonies (16%) of extremely slow growing survivors showed distinct patterns of telomeric DNA without either long heterogeneous TG tracts or substantial Y' amplification (Figure 3C and 3D), suggesting that a new type of survivor emerged in pif1Δ tlc1Δ post-senescence cells. In these cells the terminal TG tracts seemed to be even shorter than that in Type I survivors but were unexpectedly maintained during subsequent passages. This abnormality of telomeric DNA was also observed by Dewar et al. [50]. Nevertheless our results suggested that Pif1 is required for Type I survivor formation. To confirm this further, since RAD50 and RAD51 are respectively required for Type II and Type I survivor formation we checked whether survivors could form in either a rad50Δ pif1Δ tlc1Δ or a rad51Δ pif1Δ tlc1Δ triple mutant. The isogenic rad50Δ pif1Δ tlc1Δ or rad51Δ pif1Δ tlc1Δ spores were dissected and serially passaged in liquid culture. As expected, two spores of the rad50Δ pif1Δ tlc1Δ triple mutant underwent senescence gradually and virtually died out at the 9th or 11th passage (Figure 3E). A Southern blot analysis showed that tlc1Δ and pif1Δ tlc1Δ mutants displayed Type II survivor telomere structures after eleven passages, whereas rad50Δ pif1Δ tlc1Δ mutant did not (Figure 3F). These results further support our claim that Pif1 is required for Type I survivor generation. For the rad51Δ pif1Δ tlc1Δ triple mutant, three spores behaved differently in liquid culture. One spore could generate survivors, while the other two spores could not (Figure 3G), suggesting that Pif1 might also affect Type II survivor generation.
Fig. 3. Type I recombination requires Pif1 helicase. 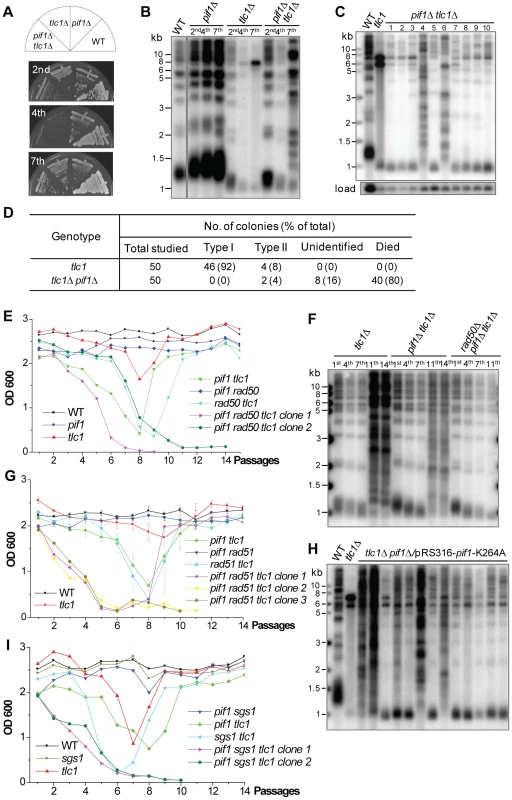
(A) The heterozygous diploid mutant, in which one copy of TLC1 and PIF1 were deleted, was sporulated and dissected, and then individual spores from tetrads were restreaked seven times to allow survivors to form. (B) The genomic DNA of independent colonies from each mutant assayed in (A) was subjected to Southern blot analysis using a TG probe after the 2nd, 4th and 7th streaking as indicated. (C) Fifty independent tlc1Δ pif1Δ senescing colonies were picked and restreaked to generate survivors and the genomic DNA of ten living post-senescence colonies at the 7th streaking was subjected to Southern blot analysis. The blot membrane was re-probed with a CDC15 probe as a loading control [50]. (D) Statistical results of the survivors generated from fifty independent colonies in tlc1Δ and pif1Δ tlc1Δ mutants. (E) The heterozygous diploid triple mutant of PIF1/pif1Δ RAD50/rad50Δ TLC1/tlc1Δ was dissected and the isogenic spores were subjected to cell viability assay in liquid culture. The results of two spores for pif1Δ tlc1Δ rad50Δ mutant are shown. (F) Genomic DNA of the tlc1Δ, pif1Δ tlc1Δ and pif1Δ rad50Δ tlc1Δ strains assayed in (E) was subjected to a Southern blot analysis after the 1st, 4th, 7th, 11th and 14th passages. (G) The heterozygous diploid triple mutant of PIF1/pif1Δ RAD51/rad51Δ TLC1/tlc1Δ was dissected and the isogenic spores were subjected to a liquid culture cell viability assay. (H) Fifty independent tlc1Δ pif1Δ/pRS316-pif1-K264A senescing colonies were serially restreaked and DNA of thirteen surviving post-senescence colonies after the 7th streaking was subjected to the Southern blot assay shown. (I) The heterozygous diploid triple mutant of PIF1/pif1Δ SGS1/sgs1Δ TLC1/tlc1Δ was dissected and the isogenic spores were subjected to cell viability assay in liquid culture. The results of two spores for pif1Δ tlc1Δ sgs1Δ mutant are shown. To investigate whether Pif1's helicase activity is required for Type I survivor formation, we constructed the pRS316-pif1-K264A plasmid and transformed it into pif1Δ cells, as the lysine residue of 264 in the ATP-binding domain of Pif1 is essential for Pif1's helicase activity [35]. Fifty senescing colonies of pif1Δ tlc1Δ/pRS316-pif1-K264A strain were randomly selected and passaged to allow survivors to generate. Thirty-seven post-senescence colonies died during the sequential streaks, while thirteen colonies generated survivors. A Southern blot analysis revealed that the telomere structures of these post-senescence survivors were very similar to those of the pif1Δ tlc1Δ mutant survivors (Figure 3H). We therefore concluded that Pif1's helicase activity plays a key role in telomeric DNA recombination.
Helicases are nucleic acid-dependent ATP-ases that are capable of unwinding DNA or RNA duplex substrates and play important roles in almost every cellular process including DNA replication and repair, transcription, translation, RNA processing and so on [51], [52]. In S. cerevisiae, there are 132 open-reading-frames that encode helicase or helicase-like proteins [35]. Thirteen of them have been shown to have DNA helicase activity. We knocked out TLC1 in each of these thirteen DNA helicase gene mutants (Figure S4) and carried out survivor screenings to investigate if these genes affect Type I or Type II survivor generation. In contrast with PIF1, the other twelve DNA helicase genes and TLC1 double deletion mutants generated Type I survivors on solid medium, indicating that they are not essential for Type I survivor formation (Figure S4A). In liquid medium, sgs1Δ tlc1Δ cells generated Type I survivors, while the other twelve DNA helicase genes and TLC1 double deletion mutants generated Type II survivors after passaging 12 times (about 200 population doublings) (Figure S4B). This result is consistent with a previous report which shows Sgs1 helicase is required for Type II survivor formation [53]. We obtained the pif1Δ sgs1Δ tlc1Δ triple mutant dissected from the heterozygous PIF1/pif1Δ SGS1/sgs1Δ TLC1/tlc1Δ diploid mutant. The pif1Δ sgs1Δ tlc1Δ mutant was cultured in liquid medium and no survivors were recovered (Figure 3I). It was therefore concluded that Pif1 and Sgs1 may define the Type I and Type II survivor formation pathways respectively.
Screening of TLM gene deletion library in liquid medium identifies genes affecting Type II survivor formation
In order to screen for genes that might affect Type II survivor formation, we grew the 280 telomerase-null tlmΔ mutants serially in liquid medium to generate survivors (Figure 4A) [5]. If Type II survivors arise, they eventually out-compete their Type I counterparts in liquid culture because of their aforementioned growth advantage [5]. There were, however, some strains that lacked the genes required for Type II survivor formation, and thus generated only Type I survivors. The viability of these senescing mutants was recorded during passages and survivor cells were harvested at the end of serial culturing. The genomic DNA of the liquid-cultured cells was isolated and subjected to Southern blot with a telomeric TG1–3 probe. Twenty-four tlc1Δ tlmΔ double mutants formed Type I survivors, suggesting these twenty-four genes were required for Type II survivor formation (Figure 4B and Table 2). To further confirm the Type I phenotypes of these mutants, we used a Y' probe and performed Southern blot hybridization to examine the DNA structure. The results clearly showed significant amplification of Y'-elements, a characteristic typical of Type I survivors (Figure 4C).
Fig. 4. Southern blot analyses of Type I survivors generated in tlmΔ tlc1Δ mutants in liquid culture. 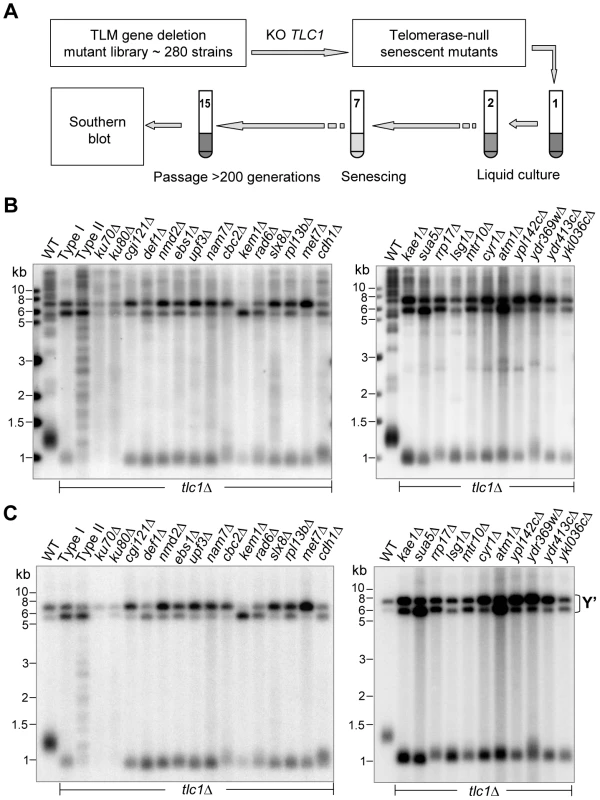
(A) Schematic illustration of the screening procedures for genes that affect Type II survivor formation (refer to details in main text). (B and C) Southern blot analyses of survivor types in the tlc1Δ strain (Type I and Type II serves as controls) and twenty-six tlmΔ tlc1Δ double mutants using a TG probe (B) and a Y' probe (C). Tab. 2. List of <i>S. cerevisiae</i> TLM genes required for Type II survivor formation. 
Among these twenty-four genes, twenty-two had never before been identified for their involvement in Type II survivor formation (Table 2). The two genes identified in our screening that have been previously reported to maintain such a function include SUA5 and DEF1 [16], [18]. It is important to note that survivors generated in tlc1Δ yku70Δ or tlc1Δ yku80Δ cells exhibited distinctive telomeric DNA patterns that differed from classical Type I and Type II survivor structures (Figure 4B and 4C, left panels) [54], [55]. Moreover, both tlc1Δ yku70Δ and tlc1Δ yku80Δ cells exhibited more rapid senescence and became survivors as soon as the telomeric DNA from germinating spores could be examined, observations which are consistent with earlier reports [55], [56]. The results of the yku70 and yku80 mutants were presented in this section with the other mutants which displayed Type I survivors because survivor generation in tlc1Δ yku70Δ and tlc1Δ yku80Δ mutants is more dependent upon RAD51 than RAD50 [54].
Type II survivor formation involves the Rad6-Bre1 pathway
As mentioned above, we identified twenty-two genes not previously known to be required for Type II survivor formation (Table 2). RAD6 remains of particular interest as previous studies have shown that RAD6 plays important roles in recombinational repair [57]. Rad6 is an E2 ubiquitin-conjugating enzyme and it interacts with three E3 ubiquitin ligases (Bre1, Rad18 and Ubr1) known to be involved in different DNA repair pathways [58], [59]. Rad6 and Bre1 are responsible for H2B-K123 ubiquitination, which is required for H3-K4 methylation [60]. Rad6 and Rad18 are involved in post-replication repair via their role in ubiquitination of PCNA [61]. Rad6 and Ubr1 have been linked to DNA repair through their function in degradation of cohesin [62]. Our Southern blot analysis showed that rad6Δ tlc1Δ double mutant cells in liquid culture generated only Type I survivors (Figure 5A), suggesting that Rad6 is required for Type II survivor formation. To validate this result, we knocked out RAD51, which is required for Type I survivor formation, in the rad6Δ tlc1Δ cells. All four clones of the rad6Δ rad51Δ tlc1Δ mutant underwent senescence and were unable to generate survivors (Figure 5B), confirming that Rad6 is required for Type II survivor formation.
Fig. 5. The effect of the Rad6 on survivor formation. 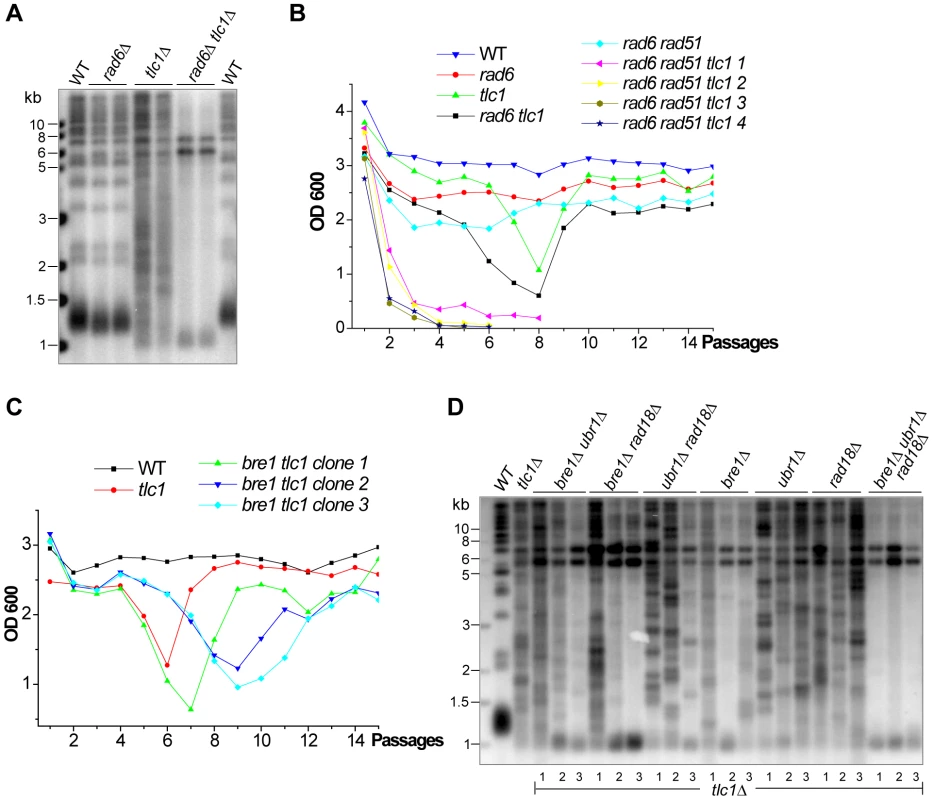
(A) The isogenic strains of tlc1Δ, rad6Δ and rad6Δ tlc1Δ were serially passaged in liquid medium to generate survivors and then genomic DNA of survivors was subjected to Southern blot analysis. (B) The heterozygous diploid RAD6/rad6Δ RAD51/rad51Δ TLC1/tlc1Δ strain was sporulated and tetrads were dissected, and the spores with the indicated genotypes (including four clones of rad6Δ rad51Δ tlc1Δ were subjected to a cell viability assay. (C) The heterozygous diploid TLC1/tlc1Δ RAD18/rad18Δ BRE1/bre1Δ UBR1/ubr1Δ mutant was sporulated and tetrads were dissected, and the spores with the indicated genotypes were subjected to a cell viability assay. Cell viabilities of three clones of the bre1Δ tlc1Δ mutant are shown in this panel, while those of the ubr1Δ tlc1Δ, rad18Δ tlc1Δ, bre1Δ ubr1Δ tlc1Δ, bre1Δ rad18Δ tlc1Δ, ubr1Δ rad18Δ tlc1Δ, and rad18Δ bre1Δ ubr1Δ tlc1Δ mutants are shown in Figure S5. (D) Genomic DNA of the isogenic strains (indicated on top of the panel) was isolated and subjected to Southern blot assay. Three colonies (labeled at the bottom of the panel) of each strain were examined. In order to determine the downstream pathways utilized by Rad6 during Type II survivor generation we constructed the heterozygous diploid strain of TLC1/tlc1Δ RAD18/rad18Δ BRE1/bre1Δ UBR1/ubr1Δ. The isogenic haploid tlc1Δ strains of single-, double-, triple - and quadruple-mutants were derived by sporulation. Three independent colonies of tlc1Δ bre1Δ, tlc1Δ ubr1Δ, tlc1Δ rad18Δ, tlc1Δ bre1Δ ubr1Δ, tlc1Δ bre1Δ rad18Δ, tlc1Δ ubr1Δ rad18Δ, and tlc1Δ rad18Δ bre1Δ ubr1Δ were passaged in liquid medium to allow survivor formation. The analysis of strain viability is shown in Figure 5C and Figure S5. An aliquot of each liquid-grown survivor was harvested on the second day of recovery, and its telomeric DNA was examined by Southern blot (Figure 5D). The tlc1Δ ubr1Δ, tlc1Δ rad18Δ and tlc1Δ ubr1Δ rad18Δ survivors displayed no obvious amplification of Y'-subtelomeric elements whereas the tlc1Δ bre1Δ, tlc1Δ bre1Δ ubr1Δ and Δtlc1Δ bre1Δ rad18Δ survivors that lacked the BRE1 gene displayed significant Y'-element amplification (Figure 5D). These data suggest Bre1 plays an even more positive regulatory role in Type II survivor generation than Ubr1 and Rad18. Interestingly, the tlc1Δ rad18Δ bre1Δ ubr1Δ mutant cells only allowed the development of Type I survivors. These results indicate that Rad6 functions through its downstream pathways and most importantly Bre1 to promote Type II survivor formation.
The KEOPS complex is required for Type II recombination
In addition to RAD6, CGI121 and KAE1 were also identified during our liquid-culture screen as contributing to Type II survivor formation (Table 2). Cgi121 and Kae1 belong to the KEOPS complex, which is evolutionarily conserved from archaea to mammals [63]. In S. cerevisiae, the KEOPS complex consists of five subunits (Cgi121, Bud32, Kae1, Gon7 and Pcc1) and plays multiple roles in transcription, tRNA modification (t6A), chromosome segregation and telomere uncapping-elongation [29], [64]–[66]. The deletion mutants of BUD32 and GON7 were in our original TLM library but in our initial screening the severe growth defects of the bud32Δ and gon7Δ haploid strain made it impossible for us to knock-out TLC1. PCC1 was not in the 280 TLM gene list, and therefore was not covered in our initial screening. In order to determine whether Bud32, Gon7 and Pcc1 were also involved in telomere recombination, we constructed the heterozygous diploid mutants in which one copy of TLC1 and BUD32, GON7 or PCC1 were deleted. The double mutants of bud32Δ tlc1Δ, gon7Δ tlc1Δ and pcc1Δ tlc1Δ were obtained from tetrad dissection and were serially passaged in liquid medium. All the survivors displayed Type I patterns of Y' amplification (Figure 6A), indicating that Type II recombination could not take place in the absence of Bud32, Gon7 or Pcc1.
Fig. 6. Each subunit of the KEOPS complex is essential for Type II recombination. 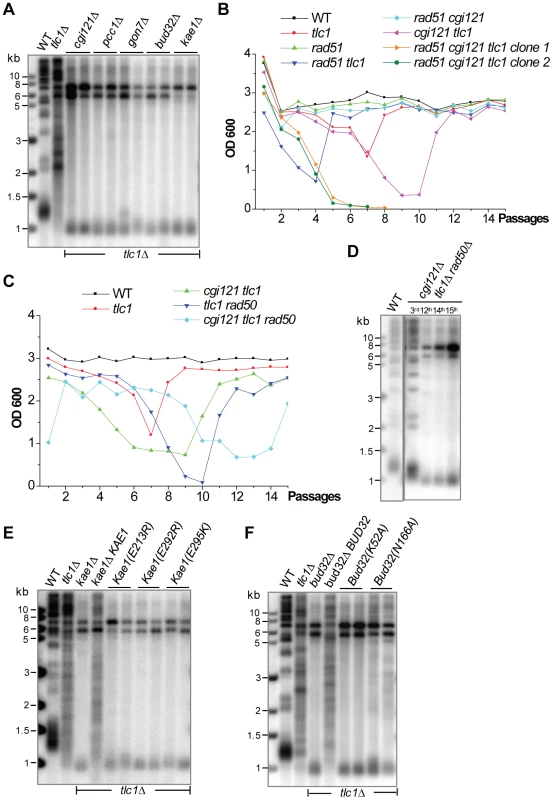
(A) Southern blot of the genomic DNA of the survivors generated by serial liquid culturing of cgi121Δ tlc1Δ, pcc1Δ tlc1Δ, gon7Δ tlc1Δ, bud32Δ tlc1Δ and kae1Δ tlc1Δ mutants. (B) Liquid culture cell viability analyses of sibling spores generated in the diploid CGI121/cgi121Δ TLC1Δtlc1Δ RAD51/rad51Δ strain. The cgi121Δ tlc1Δ rad51Δ triple mutant died out at the 7th passage. (C) Cell viability analysis of sibling spores generated in the diploid CGI121/cgi121Δ TLC1Δtlc1Δ RAD50/rad50Δ strain. Two spores were analyzed in parallel and the results were similar. (D) Southern blot analysis of telomere DNA in survivors generated by serial liquid culturing of the cgi121Δ tlc1Δ rad50Δ triple mutant. (E) and (F) Southern blot analyses of telomere DNA in survivors generated by serial liquid culturing of the tlc1Δ kae1(E213R), tlc1Δ bud32(E292R), tlc1Δ bud32(E295K) mutants (in E), tlc1Δ bud32(K52A) and tlc1Δ bud32(N166A) mutants (in F). To further confirm the critical role the KEOPS complex plays in telomere recombination, we tested whether survivor formation in cgi121Δ tlc1Δ cells would be affected in the absence of RAD51 or RAD50. Unfortunately, we could not examine the genetic interaction between RAD51 or RAD50 and the other four KEOPS subunits in telomere recombination because the bud32Δ, kae1Δ, gon7Δ and pcc1Δ mutants all exhibited severe growth defects. Therefore we focused on CGI121 by generating a heterozygous diploid strain in which one copy of TLC1, CGI121 and RAD51 (or RAD50) was deleted. The isogenic strains of single, double and triple mutants were derived from tetrad dissection and serially cultured in liquid medium. The cgi121Δ tlc1Δ rad51Δ triple mutant died out rapidly, while other tlc1Δ mutants were able to recover robust growth when survivors arose (Figure 6B). Consistently, the cgi121Δ tlc1Δ rad50Δ triple mutant was able to bypass the senescence crisis by generating Type I survivors (Figure 6C and 6D). These results support the conclusion that CGI121 and likely the entire KEOPS complex is required for Type II recombination.
Previous studies have shown that Kae1 has ATP-binding activity and Bud32 acts as a protein kinase and the activities of both of these gene products appear to be essential for all the roles played by the KEOPS complex [29], [63], [66], [67]. Based on the previous biochemical and structural analyses of Kae1 and Bud32 [63], we constructed kae1(E213R) tlc1Δ, bud32(K52A) tlc1Δ and bud32(N166A) tlc1Δ double mutant strains in which the Kae1 ATP-binding site and the Bud32 kinase catalytic sites were mutated. These mutants were unable to generate Type II survivors, but rather exclusively developed Type I survivors when cultured in liquid medium (Figure 6E and 6F). Likewise, the kae1(E292R) and kae1(E295K) mutants, which no longer maintain an interation between Kae1 and Bud32, also displayed a defect in Type II survivor generation (Figure 6E). These data indicate that both the Kae1-Bud32 interaction and their biochemical activities were indispensable for Type II telomere recombination. We therefore concluded that the whole KEOPS complex was necessary for Type II recombination.
Some TLM genes involved in telomere recombination also affect DNA recombination in general
Previously, several labs performed genome-wide screens searching for genes that affect DNA repair and/or recombination and dozens of genes were documented [68]–[71]. In this study we have identified ten genes which affect Type I telomere recombination and twenty-two genes which affect Type II telomere recombination. Fifteen of these genes have already been reported to have potential roles in general DNA repair and/or recombination (Table S2). In order to determine whether the other seventeen genes also play roles in general DNA repair/recombination, we performed three assays used previously [72], [73] to examine relative levels of inter-chromosomal homologous recombination (Figure 7A) and intra-chromosomal homologous recombination in haploid (Figure 7B) and diploid strains (Figure 7C). Each assay detected genomic gene conversion events through the recovery of an intact LEU2 marker by the integration of two seperated fragments (Figure 7A–7C, upper panels). Ten of the seventeen mutants we tested exhibited an extremely slow growth phenotype and were not viable for testing using the general recombination assays. The remaining seven mutants (nam7Δ, ebs1Δ, upf3Δ, nmd2Δ, rps16bΔ, soh1Δ and cgi121Δ) showed decreased activities in inter - or intra-chromosomal homologous recombination (Figure 7A–7C). Therefore it is likely that these seven genes participate in telomere recombination as well as recombination at other genomic loci.
Fig. 7. General homologous recombination activities in nam7Δ, ebs1Δ, upf3Δ, nmd2Δ, rps16bΔ, soh1Δ, and cgi121Δ mutants, and break-induced-replication efficiencies in ies1Δ, ies3Δ, pif1Δ, rad6Δ, and cgi121Δ mutants. 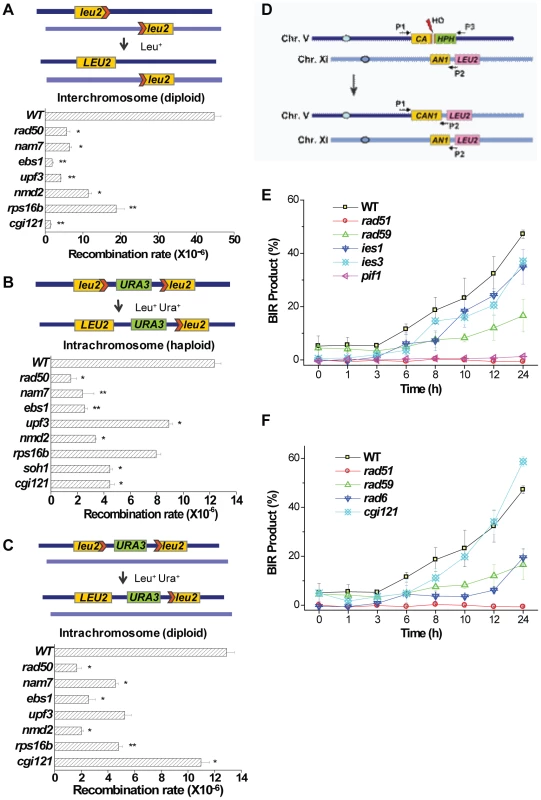
(A) Inter-chromosomal recombination assay in the indicated homozygous diploid deletion mutants. (B) Intra-chromosomal recombination assay in the indicated haploid mutants. (C) Intra-chromosomal recombination assay in the indicated homozygous diploid deletion mutants. The upper panels are the schematic illustrations of each recombination event. The lower panels show the measured recombination rates in the indicated mutants. These assays were performed as reported previously [72], [73] and the statistical significance was indicated as follows: **P-value<0.02 and *P-value<0.05. The rad50Δ mutant was included as a positive control. We failed in constructing soh1Δ homozygous mutants in (A) and (C) because the mating efficiency of this mutant was extremely low. (D) Schematic illustration of the system to detect break-induced-replication (BIR) efficiencies as reported in Lydeard et al [74]. After galactose induction, HO endonuclease causes a break in the indicated site, BIR repair process generates an intact CAN1 marker which can be detected by PCR procedures using primers P1 and P2. (E and F) BIR efficiencies were measured in pif1Δ, ies1Δ and ies3Δ mutants (E), and rad6Δ and cgi121Δ mutants (F). Semi-quantitative PCR was used to measure BIR efficiency as shown in Figure S6. The INO80 complex, Pif1, and Rad6 affect telomere recombination through break-induced-replication mechanism
Break-induced-replication (BIR) only requires one free DNA end to take place and it has been proposed to be the principal mechanism for telomere recombination and survivor generation [14]. To examine whether the INO80 complex, Pif1, Rad6 and the KEOPS complex participate in telomere recombination via Rad51-dependent BIR process we used a system developed by Lydeard et al. to measure the BIR efficiencies in ies1Δ, ies3Δ, pif1Δ, rad6Δ and cgi121Δ mutants [74] (Figure 7D). The rad51Δ and rad59Δ mutant strains served as positive controls [14]. Our results showed that similar to the rad51Δ mutant, the pif1Δ mutant displayed little BIR efficiency (Figure 7E). In ies1Δ, ies3Δ and rad6Δ mutants, the BIR efficiencies were greatly decreased as also seen in the rad59Δ mutant (Figure 7E and 7F). In contrast, the BIR efficiency in the cgi121Δ mutant was comparable to that of the wild-type strain (Figure 7F). Taken together, these data indicate that Pif1 is required for Rad51-dependent break-induced-replication, and the INO80 complex and Rad6, but not the KEOPS complex, contribute to this BIR process.
Discussion
Unlike most other chromosomal loci, eukaryotic telomeres have unique structures attributed to their repetitive DNA sequence and binding proteins [75]. Linear chromosome ends can be recognized as DNA double-stranded breaks and are thus often subjected to repair by non-homologous-end-joining and homologous recombination. It is possible that telomerase-null senescing cells are able to escape the fate of death as telomeres undergo lengthening and repair via homologous recombination. The distinct DNA makeup of Type I and Type II recombinational telomeres allowed us to carry out a genetic screening to identify genes that affect telomere recombination in telomerase-null cells.
Our candidate approach for screening telomere recombination genes had a few shortcomings. In our screening we only covered the 280 known TLM genes, which make up only 5.6% of the ∼5,000 non-essential genes in S. cerevisiae. It would be ideal to cover all non-essential genes in our screen. However, such a study would be too massive to undertake since the screening procedures included knocking out TLC1 in every strain, two to three-weeks passaging cells until they reach senescence and Southern blot experiments for multiple survivors in each mutant (see Figure 1A and Figure 4A). The candidate approach we chose therefore had a strong bias. As a result, we might have missed potential genes that do not affect telomere length, but play important roles in telomere recombination. Another challenge to our screening approach came from the nature of different growth rates of the various mutants. Although we used heterozygous diploid mutants to generate spores of tlc1Δ tlmΔ double mutants (Table S1), for quite a few mutants we were not able to distinguish between a defect in a survivor pathway and synthetic lethality (Table S1). The third issue that we were not able to resolve was to distinguish between hypo-Type I recombination and hyper-Type II recombination. The decrease of Type I survivor frequency seen in the mutants, such as rpa14Δ tlc1Δ (Figure 1C) could be caused by either inhibition of Type I recombination or promotion of Type II recombination. In some Type II survivors, the amplified Y'-elements were detected in Southern blot assays (Figure 1C, Figure S1), suggesting that the increase of Type II survivor frequency in these mutants was a result of enhanced Type II recombination rather than inhibited Type I recombination. This model is supported by the observation that in the nine mutants shown in Figure 1C and Figure S1, the emerging events of Type I survivors were significantly reduced, but were not entirely blocked. The fourth issue that we had not taken into consideration during our primary screening was the effect of the initial telomere length of each mutant on the recombination pathways. It was recently proposed that longer telomeres, like those observed in rif1Δ and rif2Δ mutants could influence the type of recombination pathway used at the telomere [37]. Additionally, it was shown that the mre11-A470T tlc1Δ mutant promotes telomere recombination and bypass senescence efficiently because the Type I recombination occurs before growth limitation [76]. Therefore, it would have been more appropriate to perform all of our screening steps starting with TLC1/tlc1Δ TLM/tlmΔ diploids to obtain tlc1Δ single and tlc1Δ tlmΔ double mutants following tetrad dissection. The fifth issue with our screening approach was that we assumed the TLM genes only affect Type I or Type II recombination. Surprisingly, the telomere structure in the yku and pif1 mutants might actually be different from that of a typical Type I or Type II survivor (Figure 3 and Figure 4). Therefore, genes that influence pathway(s) of telomere recombination other than that of Type I or Type II might have been overlooked. The sixth issue with our screen was that we only identified ten novel genes affecting Type I survivor formation (Table 1). This number might be underrepresent the true total because our primary screening was carried out with a relatively stringent criteria and as such we may have overlooked some genes that have minor influences on the frequency Type I survivor emergence.
Although our screening approach had some imperfections, we successfully identified thirty-two TLM genes that influence telomere recombination when overcoming senescence. Ten of these TLM genes affected the emerging frequency of Type I survivors while twenty-two were required for Type II survivor generation. A large portion of 280 TLM genes have not previously been characterized for their roles in telomere function other than the length of the telomeres in these deletion strains was altered. The positive results of our screen provide more direct evidence supporting the idea that some of these uncharacterized TLM genes do affect telomeres [23], [24], [26]. Additionally, telomere recombination is a means by which cells repair defective telomeres and thus the genes involved in telomeric DNA recombination may also play a role in general DNA recombination/repair. Indeed, the TLM genes that affected either Type I or Type II recombination were also required for general DNA recombination (Figure 7A–7C). The annotated functions of the thirty-two genes that we identified point to several pathways that might contribute to telomere maintenance (Table 1 and Table 2). Some of the genes are known for functions like “rRNA processing,” “structural constituent of ribosome,” and “transport and membrane.” These gene products seem unlikely to play a direct role in telomere recombination. In contrast, the Pif1 helicase and the KEOPS complex are involved in “telomere capping and maintenance” [29], [35] and INO80 complex and Rad6 are associated with “chromatin remodeling and modification.” These genes are likely to play direct roles in telomere recombination.
The senescing pif1Δ tlc1Δ cells did not produce Type I survivors on solid medium (Figure 3C) and the rad50Δ pif1Δ tlc1Δ triple mutant was not able to generate survivors in liquid medium (Figure 3E). These results indicated that Pif1 was required for Type I survivor generation. Interestingly, not all the rad51Δ pif1Δ tlc1Δ triple mutants were able to generate Type II survivors in liquid medium (Figure 3G). Therefore, we favor a model where Pif1 helicase is required for amplification of Y'-elements to form Type I survivors and promotes TG1–3 recombination to form Type II survivors (Figure 3). Previous studies have shown that Pif1 takes part in mitochondrial DNA recombination [41]–[43], however, our data are the first to indicate that Pif1 is also involved in telomeric DNA recombination (Figure 3). In the survivors of pif1Δ tlc1Δ mutants one group exhibited a severely delayed growth phenotype and had a unique telomere structure that differed from the characteristics of either Type I or Type II (Figure 3C). We speculate that these types of survivors require RAD50 to maintain telomeres since no survivors were recovered in the pif1Δ tlc1Δ rad50Δ triple mutant (Figure 3E). In the future it will be interesting to examine how the short telomeres are maintained in these survivors.
Chromatin remodeling complexes have been shown by others to play roles in DNA repair processes via homologous recombination [77]. However, a causal link between chromatin structure alteration and recombination has not yet been well established. We found that in the absence of active chromatin remodeling by the INO80 complex, telomere Type I recombination was unable to efficiently take place (Figure 2A and 2B), suggesting that the alteration of chromatin structure is a pre-requisite to the Type I recombination process at telomeres. Our results could provide an explanation for the previous observation that ies3Δ est1Δ cells generated survivors later than the est1Δ single mutant [34], as ies3Δ est1Δ cells likely have a lower efficiency of Type I survivor generation than est1Δ cells. SAP30, which encodes a subunit of histone deacetylase Rpd3 complex, was also identified in our screening. Deletion of SAP30 dramatically reduced the emerging rate of the Type I survivors (Table 1), suggesting that the Rpd3 histone deacetylase complex may also inhibit Type I recombination. The SWR1 complex is another chromatin remodeling complex that belongs to the INO80 family of remodeling enzymes. SWR1 and INO80 complexes share four common subunits: Rvb1, Rvb2, Arp4 and Act1 [38]. It will be intriguing to determine if other chromatin remodeling enzymes like SWR1 or histone modification enzymes play roles in telomere recombination.
The KEOPS complex gains its name from “Kinase, Endopeptidase and Other Proteins of small Size” [29], and is comprised of five small proteins (Bud32, Kae1, Pcc1, Gon7 and Cgi121) which form a stable complex in vitro and in vivo [29], [63], [65], [67]. Bud32 has kinase activity while Kae1 maintains endopeptidase activity [29]. The KEOPS complex or its subunit(s) are involved in several biological processes, to which each KEOPS subunit seems to contribute unequally. Pcc1, Gon7, Kae1 and Bud32, for example, are recruited to several genomic loci and affect gene transcription [65]. Kae1 contributes to faithful chromosome segregation [64] while Bud32, Cgi121, Gon7 regulate cell polarity in bud-site selection [78]. Additionally, Bud32, Kae1 and Pcc1 are essential for a universal tRNA modification called threonyl carbamoyl adenosine (t6A), for which Cgi121 is dispensable [66]. Moreover, all the subunits of the KEOPS complex appear to play roles in telomere uncapping and telomere length regulation [29]. Our screen elucidated a novel function of the KEOPS complex in telomere recombination, as deficiency of any subunit of the KEOPS complex led to the failure in generating Type II survivors in the tlc1Δ mutant (Figure 5A). The molecular mechanism by which the KEOPS complex influences telomere recombination remains unclear. A previous study by Downey et al. showed that mutation of the KEOPS complex decreased the amount of single-stranded telomeric DNA in the cdc13-1 mutant [29]. It is possible that the KEOPS complex facilitates the formation of the telomeric 3′-overhang and promotes recombination of TG-tracts. Coincidently, SUA5, a telomeric single-stranded DNA binding protein, is required for both Type II recombination and t6A modification of tRNA [18], [79]. It will be interesting to determine whether SUA5 is a downstream target of the KEOPS complex and if it functions in the same pathway in regulating telomere recombination. It is possible that Sua5 is a substrate of the Bud32 kinase.
In summary, our screen identified dozens of genes that regulate telomere recombination pathways. Because of the complexity of the recombination process, the molecular mechanisms of telomere recombination remain elusive. Our work not only provides important clues for beginning to understand how telomere recombination is coordinated, but also offers new insights into general DNA repair processes via homologous recombination.
Materials and Methods
Yeast strains and plasmids
All strains used in this work are summarized in Table S1 and Table S3. Gene deletions were carried out using standard procedures by genetic cross and homologous recombination. Systematic deletion strains are from EUROSCARF. We constructed CEN plasmids pRS316-PIF1, pRS313-KAE1 and pRS313-BUD32 by inserting fragments (from upstream 1000 bp to downstream 500 bp of genes' open reading frame) into the pRS316 or pRS313 vector. Point mutations were introduced using a site-directed mutagenesis method.
Cell viability assay
A single colony of the indicated yeast strains was inoculated into 5 ml yeast extract-peptone-dextrose (YPD) medium and grown at 30°C to saturation (OD600 ∼2.5 to 3.0). Then every 24 hours the cell density was measured by spectrometry (OD600) and the cell culture was diluted to the density at OD600 ∼0.02 with fresh YPD medium. This procedure was repeated for up to 14 times, unless the cell density is too low for dilution.
Single-colony streaking assay
A single colony of the indicated yeast strains was streaked on YPD plate and grown until emergency of single colonies (25 cell divisions) at 30°C. Individual colonies were restreaked repeatedly at least six times to allow survivors to generate.
Telomere Southern blot
Genomic DNA was prepared from each strain, digested with XhoI, separated on 1% gel, transferred to Hybond-N+ membrane (GE Healthcare) and then probed with TG1–3 telomere-specific probe or Y'-element probe [25]. The CDC15 probe was ∼263 bp sequence of CDC15 gene [50].
General recombination assays
Recombination assays for intrachromosomal and interchromosomal recombination in haploid and diploid strains were performed and recombination rates were determined as previously described [72], [73]. For each mutant, about 2×107 yeast cells were plated on solid selective medium. After growing at 30°C for 2–3 days, about 200 positive colonies would appear on the plate in wild-type haploid strain. Recombination rates were calculated and statistically analyzed by paired two-sample t-test.
Measurement of break-induced-replication efficiency
Break-induced-replication (BIR) efficiency was measured in a system developed by Lydeard et al [74]. Semi-quantitative PCR was conducted as previously described [74]. PCR products were quantified in Image Quant Software.
Supporting Information
Zdroje
1. McEachernMJ, KrauskopfA, BlackburnEH (2000) Telomeres and their control. Annu Rev Genet 34 : 331–358.
2. WellingerRJ, WolfAJ, ZakianVA (1993) Saccharomyces telomeres acquire single-strand TG1–3 tails late in S phase. Cell 72 : 51–60.
3. GreiderCW, BlackburnEH (1985) Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell 43 : 405–413.
4. LundbladV, BlackburnEH (1993) An alternative pathway for yeast telomere maintenance rescues est1 - senescence. Cell 73 : 347–360.
5. TengSC, ZakianVA (1999) Telomere-telomere recombination is an efficient bypass pathway for telomere maintenance in Saccharomyces cerevisiae. Mol Cell Biol 19 : 8083–8093.
6. LingnerJ, CechTR, HughesTR, LundbladV (1997) Three Ever Shorter Telomere (EST) genes are dispensable for in vitro yeast telomerase activity. Proc Natl Acad Sci U S A 94 : 11190–11195.
7. SingerMS, GottschlingDE (1994) TLC1: template RNA component of Saccharomyces cerevisiae telomerase. Science 266 : 404–409.
8. ChenXF, MengFL, ZhouJQ (2009) Telomere recombination accelerates cellular aging in Saccharomyces cerevisiae. PLoS Genet 5: e1000535 doi:10.1371/journal.pgen.1000535.
9. ShoreD, BianchiA (2009) Telomere length regulation: coupling DNA end processing to feedback regulation of telomerase. EMBO J 28 : 2309–2322.
10. BryanTM, ReddelRR (1997) Telomere dynamics and telomerase activity in in vitro immortalised human cells. Eur J Cancer 33 : 767–773.
11. NeumannAA, ReddelRR (2002) Telomere maintenance and cancer – look, no telomerase. Nat Rev Cancer 2 : 879–884.
12. LundbladV, SzostakJW (1989) A mutant with a defect in telomere elongation leads to senescence in yeast. Cell 57 : 633–643.
13. LendvayTS, MorrisDK, SahJ, BalasubramanianB, LundbladV (1996) Senescence mutants of Saccharomyces cerevisiae with a defect in telomere replication identify three additional EST genes. Genetics 144 : 1399–1412.
14. McEachernMJ, HaberJE (2006) Break-induced replication and recombinational telomere elongation in yeast. Annu Rev Biochem 75 : 111–135.
15. LeS, MooreJK, HaberJE, GreiderCW (1999) RAD50 and RAD51 define two pathways that collaborate to maintain telomeres in the absence of telomerase. Genetics 152 : 143–152.
16. ChenYB, YangCP, LiRX, ZengR, ZhouJQ (2005) Def1p is involved in telomere maintenance in budding yeast. J Biol Chem 280 : 24784–24791.
17. GrandinN, CharbonneauM (2003) Mitotic cyclins regulate telomeric recombination in telomerase-deficient yeast cells. Mol Cell Biol 23 : 9162–9177.
18. MengFL, ChenXF, HuY, TangHB, DangW, et al. (2010) Sua5p is required for telomere recombination in Saccharomyces cerevisiae. Cell Res 20 : 495–498.
19. LeeJY, MogenJL, ChavezA, JohnsonFB (2008) Sgs1 RecQ helicase inhibits survival of Saccharomyces cerevisiae cells lacking telomerase and homologous recombination. J Biol Chem 283 : 29847–29858.
20. PikeBL, HeierhorstJ (2007) Mdt1 facilitates efficient repair of blocked DNA double-strand breaks and recombinational maintenance of telomeres. Mol Cell Biol 27 : 6532–6545.
21. TsaiYL, TsengSF, ChangSH, LinCC, TengSC (2002) Involvement of replicative polymerases, Tel1p, Mec1p, Cdc13p, and the Ku complex in telomere-telomere recombination. Mol Cell Biol 22 : 5679–5687.
22. TengSC, ChangJ, McCowanB, ZakianVA (2000) Telomerase-independent lengthening of yeast telomeres occurs by an abrupt Rad50p-dependent, Rif-inhibited recombinational process. Mol Cell 6 : 947–952.
23. GatbontonT, ImbesiM, NelsonM, AkeyJM, RuderferDM, et al. (2006) Telomere length as a quantitative trait: genome-wide survey and genetic mapping of telomere length-control genes in yeast. PLoS Genet 2: e35 doi:10.1371/journal.pgen.0020035.
24. MengFL, HuY, ShenN, TongXJ, WangJ, et al. (2009) Sua5p a single-stranded telomeric DNA-binding protein facilitates telomere replication. Embo J 28 : 1466–1478.
25. TsukamotoY, MitsuokaC, TerasawaM, OgawaH, OgawaT (2005) Xrs2p regulates Mre11p translocation to the nucleus and plays a role in telomere elongation and meiotic recombination. Mol Biol Cell 16 : 597–608.
26. AskreeSH, YehudaT, SmolikovS, GurevichR, HawkJ, et al. (2004) A genome-wide screen for Saccharomyces cerevisiae deletion mutants that affect telomere length. Proc Natl Acad Sci U S A 101 : 8658–8663.
27. AzamM, LeeJY, AbrahamV, ChanouxR, SchoenlyKA, et al. (2006) Evidence that the S.cerevisiae Sgs1 protein facilitates recombinational repair of telomeres during senescence. Nucleic Acids Res 34 : 506–516.
28. BertuchAA, LundbladV (2004) EXO1 contributes to telomere maintenance in both telomerase-proficient and telomerase-deficient Saccharomyces cerevisiae. Genetics 166 : 1651–1659.
29. DowneyM, HoulsworthR, MaringeleL, RollieA, BrehmeM, et al. (2006) A genome-wide screen identifies the evolutionarily conserved KEOPS complex as a telomere regulator. Cell 124 : 1155–1168.
30. HayashiN, MurakamiS (2002) STM1, a gene which encodes a guanine quadruplex binding protein, interacts with CDC13 in Saccharomyces cerevisiae. Mol Genet Genomics 267 : 806–813.
31. PangTL, WangCY, HsuCL, ChenMY, LinJJ (2003) Exposure of Single-stranded Telomeric DNA Causes G2/M Cell Cycle Arrest in Saccharomyces cerevisiae. J Biol Chem 278 : 9318–9321.
32. ToogunOA, DezwaanDC, FreemanBC (2008) The hsp90 molecular chaperone modulates multiple telomerase activities. Mol Cell Biol 28 : 457–467.
33. YorkSJ, ArmbrusterBN, GreenwellP, PetesTD, YorkJD (2005) Inositol diphosphate signaling regulates telomere length. J Biol Chem 280 : 4264–4269.
34. YuEY, Steinberg-NeifachO, DandjinouAT, KangF, MorrisonAJ, et al. (2007) Regulation of telomere structure and functions by subunits of the INO80 chromatin remodeling complex. Mol Cell Biol 27 : 5639–5649.
35. ZhouJ, MonsonEK, TengSC, SchulzVP, ZakianVA (2000) Pif1p helicase, a catalytic inhibitor of telomerase in yeast. Science 289 : 771–774.
36. LahayeA, StahlH, Thines-SempouxD, FouryF (1991) PIF1: a DNA helicase in yeast mitochondria. Embo J 10 : 997–1007.
37. ChangM, DittmarJC, RothsteinR (2011) Long telomeres are preferentially extended during recombination-mediated telomere maintenance. Nat Struct Mol Biol 18 : 451–456.
38. BaoY, ShenX (2011) SnapShot: Chromatin remodeling: INO80 and SWR1. Cell 144 : 158–e152, 158-158, e152.
39. BouleJB, ZakianVA (2007) The yeast Pif1p DNA helicase preferentially unwinds RNA DNA substrates. Nucleic Acids Res 35 : 5809–5818.
40. SchulzVP, ZakianVA (1994) The saccharomyces PIF1 DNA helicase inhibits telomere elongation and de novo telomere formation. Cell 76 : 145–155.
41. ChengX, DunawayS, IvessaAS (2007) The role of Pif1p, a DNA helicase in Saccharomyces cerevisiae, in maintaining mitochondrial DNA. Mitochondrion 7 : 211–222.
42. FouryF, DyckEV (1985) A PIF-dependent recombinogenic signal in the mitochondrial DNA of yeast. Embo J 4 : 3525–3530.
43. Van DyckE, FouryF, StillmanB, BrillSJ (1992) A single-stranded DNA binding protein required for mitochondrial DNA replication in S. cerevisiae is homologous to E. coli SSB. Embo J 11 : 3421–3430.
44. BouleJB, VegaLR, ZakianVA (2005) The yeast Pif1p helicase removes telomerase from telomeric DNA. Nature 438 : 57–61.
45. BuddME, ReisCC, SmithS, MyungK, CampbellJL (2006) Evidence suggesting that Pif1 helicase functions in DNA replication with the Dna2 helicase/nuclease and DNA polymerase delta. Mol Cell Biol 26 : 2490–2500.
46. PikeJE, BurgersPM, CampbellJL, BambaraRA (2009) Pif1 helicase lengthens some Okazaki fragment flaps necessitating Dna2 nuclease/helicase action in the two-nuclease processing pathway. J Biol Chem 284 : 25170–25180.
47. IvessaAS, ZhouJQ, ZakianVA (2000) The Saccharomyces Pif1p DNA helicase and the highly related Rrm3p have opposite effects on replication fork progression in ribosomal DNA. Cell 100 : 479–489.
48. RibeyreC, LopesJ, BouleJB, PiazzaA, GuedinA, et al. (2009) The yeast Pif1 helicase prevents genomic instability caused by G-quadruplex-forming CEB1 sequences in vivo. PLoS Genet 5: e1000475 doi:10.1371/journal.pgen.1000475.
49. PaeschkeK, CapraJA, ZakianVA (2011) DNA replication through G-quadruplex motifs is promoted by the Saccharomyces cerevisiae Pif1 DNA helicase. Cell 145 : 678–691.
50. DewarJM, LydallD (2010) Pif1 - and Exo1-dependent nucleases coordinate checkpoint activation following telomere uncapping. EMBO J 29 : 4020–4034.
51. SingletonMR, DillinghamMS, WigleyDB (2007) Structure and mechanism of helicases and nucleic acid translocases. Annu Rev Biochem 76 : 23–50.
52. ShiratoriA, ShibataT, ArisawaM, HanaokaF, MurakamiY, et al. (1999) Systematic identification, classification, and characterization of the open reading frames which encode novel helicase-related proteins in Saccharomyces cerevisiae by gene disruption and Northern analysis. Yeast 15 : 219–253.
53. HuangP, PrydeFE, LesterD, MaddisonRL, BortsRH, et al. (2001) SGS1 is required for telomere elongation in the absence of telomerase. Curr Biol 11 : 125–129.
54. GrandinN, CharbonneauM (2003) The Rad51 pathway of telomerase-independent maintenance of telomeres can amplify TG1–3 sequences in yku and cdc13 mutants of Saccharomyces cerevisiae. Mol Cell Biol 23 : 3721–3734.
55. NugentCI, BoscoG, RossLO, EvansSK, SalingerAP, et al. (1998) Telomere maintenance is dependent on activities required for end repair of double-strand breaks. Curr Biol 8 : 657–660.
56. TongXJ, LiQJ, DuanYM, LiuNN, ZhangML, et al. (2011) Est1 protects telomeres and inhibits subtelomeric y'-element recombination. Mol Cell Biol 31 : 1263–1274.
57. GameJC, ChernikovaSB (2009) The role of RAD6 in recombinational repair, checkpoints and meiosis via histone modification. DNA Repair (Amst) 8 : 470–482.
58. RobzykK, RechtJ, OsleyMA (2000) Rad6-dependent ubiquitination of histone H2B in yeast. Science 287 : 501–504.
59. HwangWW, VenkatasubrahmanyamS, IanculescuAG, TongA, BooneC, et al. (2003) A conserved RING finger protein required for histone H2B monoubiquitination and cell size control. Mol Cell 11 : 261–266.
60. KroganNJ, DoverJ, KhorramiS, GreenblattJF, SchneiderJ, et al. (2002) COMPASS, a histone H3 (Lysine 4) methyltransferase required for telomeric silencing of gene expression. J Biol Chem 277 : 10753–10755.
61. HoegeC, PfanderB, MoldovanGL, PyrowolakisG, JentschS (2002) RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419 : 135–141.
62. RaoH, UhlmannF, NasmythK, VarshavskyA (2001) Degradation of a cohesin subunit by the N-end rule pathway is essential for chromosome stability. Nature 410 : 955–959.
63. MaoDY, NeculaiD, DowneyM, OrlickyS, HaffaniYZ, et al. (2008) Atomic structure of the KEOPS complex: an ancient protein kinase-containing molecular machine. Mol Cell 32 : 259–275.
64. Ben-AroyaS, CoombesC, KwokT, O'DonnellKA, BoekeJD, et al. (2008) Toward a comprehensive temperature-sensitive mutant repository of the essential genes of Saccharomyces cerevisiae. Mol Cell 30 : 248–258.
65. Kisseleva-RomanovaE, LopreiatoR, Baudin-BaillieuA, RousselleJC, IlanL, et al. (2006) Yeast homolog of a cancer-testis antigen defines a new transcription complex. Embo J 25 : 3576–3585.
66. SrinivasanM, MehtaP, YuY, PrugarE, KooninEV, et al. (2011) The highly conserved KEOPS/EKC complex is essential for a universal tRNA modification, t6A. Embo J 30 : 873–881.
67. HeckerA, LopreiatoR, GrailleM, CollinetB, ForterreP, et al. (2008) Structure of the archaeal Kae1/Bud32 fusion protein MJ1130: a model for the eukaryotic EKC/KEOPS subcomplex. Embo J 27 : 2340–2351.
68. BirrellGW, GiaeverG, ChuAM, DavisRW, BrownJM (2001) A genome-wide screen in Saccharomyces cerevisiae for genes affecting UV radiation sensitivity. Proc Natl Acad Sci U S A 98 : 12608–12613.
69. AndersenMP, NelsonZW, HetrickED, GottschlingDE (2008) A genetic screen for increased loss of heterozygosity in Saccharomyces cerevisiae. Genetics 179 : 1179–1195.
70. LiF, DongJ, PanX, OumJH, BoekeJD, et al. (2008) Microarray-based genetic screen defines SAW1, a gene required for Rad1/Rad10-dependent processing of recombination intermediates. Mol Cell 30 : 325–335.
71. WestmorelandTJ, WickramasekaraSM, GuoAY, SelimAL, WinsorTS, et al. (2009) Comparative genome-wide screening identifies a conserved doxorubicin repair network that is diploid specific in Saccharomyces cerevisiae. PLoS ONE 4: e5830 doi:10.1371/journal.pone.0005830.
72. KleinHL (1997) RDH54, a RAD54 homologue in Saccharomyces cerevisiae, is required for mitotic diploid-specific recombination and repair and for meiosis. Genetics 147 : 1533–1543.
73. AguileraA, KleinHL (1988) Genetic control of intrachromosomal recombination in Saccharomyces cerevisiae. I. Isolation and genetic characterization of hyper-recombination mutations. Genetics 119 : 779–790.
74. LydeardJR, JainS, YamaguchiM, HaberJE (2007) Break-induced replication and telomerase-independent telomere maintenance require Pol32. Nature 448 : 820–823.
75. VegaLR, MateyakMK, ZakianVA (2003) Getting to the end: telomerase access in yeast and humans. Nat Rev Mol Cell Biol 4 : 948–959.
76. JosephIS, KumariA, BhattacharyyaMK, GaoH, LiB, et al. (2010) An mre11 mutation that promotes telomere recombination and an efficient bypass of senescence. Genetics 185 : 761–770.
77. LusserA, KadonagaJT (2003) Chromatin remodeling by ATP-dependent molecular machines. Bioessays 25 : 1192–1200.
78. KatoY, KawasakiH, OhyamaY, MorishitaT, IwasakiH, et al. (2011) Cell polarity in Saccharomyces cerevisiae depends on proper localization of the Bud9 landmark protein by the EKC/KEOPS complex. Genetics 188 : 871–882.
79. El YacoubiB, LyonsB, CruzY, ReddyR, NordinB, et al. (2009) The universal YrdC/Sua5 family is required for the formation of threonylcarbamoyladenosine in tRNA. Nucleic Acids Res 37 : 2894–2909.
Štítky
Genetika Reprodukční medicína
Článek Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across PathogensČlánek TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association StudiesČlánek Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization inČlánek Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA ExpressionČlánek The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of GenesČlánek The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2013 Číslo 1
-
Všechny články tohoto čísla
- A Model of High Sugar Diet-Induced Cardiomyopathy
- Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across Pathogens
- Emerging Function of Fat Mass and Obesity-Associated Protein (Fto)
- Positional Cloning Reveals Strain-Dependent Expression of to Alter Susceptibility to Bleomycin-Induced Pulmonary Fibrosis in Mice
- Genetics of Ribosomal Proteins: “Curiouser and Curiouser”
- Transposable Elements Re-Wire and Fine-Tune the Transcriptome
- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- MAML1 Enhances the Transcriptional Activity of Runx2 and Plays a Role in Bone Development
- Predicting Mendelian Disease-Causing Non-Synonymous Single Nucleotide Variants in Exome Sequencing Studies
- A Systematic Mapping Approach of 16q12.2/ and BMI in More Than 20,000 African Americans Narrows in on the Underlying Functional Variation: Results from the Population Architecture using Genomics and Epidemiology (PAGE) Study
- Transcription of the Major microRNA–Like Small RNAs Relies on RNA Polymerase III
- Histone H3K56 Acetylation, Rad52, and Non-DNA Repair Factors Control Double-Strand Break Repair Choice with the Sister Chromatid
- Genome-Wide Association Study Identifies a Novel Susceptibility Locus at 12q23.1 for Lung Squamous Cell Carcinoma in Han Chinese
- Genetic Disruption of the Copulatory Plug in Mice Leads to Severely Reduced Fertility
- The [] Prion Exists as a Dynamic Cloud of Variants
- Adult Onset Global Loss of the Gene Alters Body Composition and Metabolism in the Mouse
- Fis Protein Insulates the Gene from Uncontrolled Transcription
- The Meiotic Nuclear Lamina Regulates Chromosome Dynamics and Promotes Efficient Homologous Recombination in the Mouse
- Genome-Wide Haplotype Analysis of Expression Quantitative Trait Loci in Monocytes
- TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association Studies
- Structural Basis of a Histone H3 Lysine 4 Demethylase Required for Stem Elongation in Rice
- The Ecm11-Gmc2 Complex Promotes Synaptonemal Complex Formation through Assembly of Transverse Filaments in Budding Yeast
- MCM8 Is Required for a Pathway of Meiotic Double-Strand Break Repair Independent of DMC1 in
- Comparative Genomic Analysis of the Endosymbionts of Herbivorous Insects Reveals Eco-Environmental Adaptations: Biotechnology Applications
- Integration of Nodal and BMP Signals in the Heart Requires FoxH1 to Create Left–Right Differences in Cell Migration Rates That Direct Cardiac Asymmetry
- Pharmacodynamics, Population Dynamics, and the Evolution of Persistence in
- A Hybrid Likelihood Model for Sequence-Based Disease Association Studies
- Aberration in DNA Methylation in B-Cell Lymphomas Has a Complex Origin and Increases with Disease Severity
- Multiple Opposing Constraints Govern Chromosome Interactions during Meiosis
- Transcriptional Dynamics Elicited by a Short Pulse of Notch Activation Involves Feed-Forward Regulation by Genes
- Dynamic Large-Scale Chromosomal Rearrangements Fuel Rapid Adaptation in Yeast Populations
- Heterologous Gln/Asn-Rich Proteins Impede the Propagation of Yeast Prions by Altering Chaperone Availability
- Gene Copy-Number Polymorphism Caused by Retrotransposition in Humans
- An Incompatibility between a Mitochondrial tRNA and Its Nuclear-Encoded tRNA Synthetase Compromises Development and Fitness in
- Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization in
- Single-Stranded Annealing Induced by Re-Initiation of Replication Origins Provides a Novel and Efficient Mechanism for Generating Copy Number Expansion via Non-Allelic Homologous Recombination
- Tbx2 Controls Lung Growth by Direct Repression of the Cell Cycle Inhibitor Genes and
- Suv4-20h Histone Methyltransferases Promote Neuroectodermal Differentiation by Silencing the Pluripotency-Associated Oct-25 Gene
- A Conserved Helicase Processivity Factor Is Needed for Conjugation and Replication of an Integrative and Conjugative Element
- Telomerase-Null Survivor Screening Identifies Novel Telomere Recombination Regulators
- Genome-Wide Analysis Reveals Selection for Important Traits in Domestic Horse Breeds
- Coordinated Degradation of Replisome Components Ensures Genome Stability upon Replication Stress in the Absence of the Replication Fork Protection Complex
- Nkx6.1 Controls a Gene Regulatory Network Required for Establishing and Maintaining Pancreatic Beta Cell Identity
- HIF- and Non-HIF-Regulated Hypoxic Responses Require the Estrogen-Related Receptor in
- Delineating a Conserved Genetic Cassette Promoting Outgrowth of Body Appendages
- The Telomere Capping Complex CST Has an Unusual Stoichiometry, Makes Multipartite Interaction with G-Tails, and Unfolds Higher-Order G-Tail Structures
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Loci Associated with -Glycosylation of Human Immunoglobulin G Show Pleiotropy with Autoimmune Diseases and Haematological Cancers
- Switchgrass Genomic Diversity, Ploidy, and Evolution: Novel Insights from a Network-Based SNP Discovery Protocol
- Centromere-Like Regions in the Budding Yeast Genome
- Sequencing of Loci from the Elephant Shark Reveals a Family of Genes in Vertebrate Genomes, Forged by Ancient Duplications and Divergences
- Mendelian and Non-Mendelian Regulation of Gene Expression in Maize
- Mutational Spectrum Drives the Rise of Mutator Bacteria
- Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA Expression
- The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of Genes
- Sex-Specific Signaling in the Blood–Brain Barrier Is Required for Male Courtship in
- A Newly Uncovered Group of Distantly Related Lysine Methyltransferases Preferentially Interact with Molecular Chaperones to Regulate Their Activity
- Is Required for Leptin-Mediated Depolarization of POMC Neurons in the Hypothalamic Arcuate Nucleus in Mice
- Unlocking the Bottleneck in Forward Genetics Using Whole-Genome Sequencing and Identity by Descent to Isolate Causative Mutations
- The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
- MTERF3 Regulates Mitochondrial Ribosome Biogenesis in Invertebrates and Mammals
- Downregulation and Altered Splicing by in a Mouse Model of Facioscapulohumeral Muscular Dystrophy (FSHD)
- NBR1-Mediated Selective Autophagy Targets Insoluble Ubiquitinated Protein Aggregates in Plant Stress Responses
- Retroactive Maintains Cuticle Integrity by Promoting the Trafficking of Knickkopf into the Procuticle of
- Phenome-Wide Association Study (PheWAS) for Detection of Pleiotropy within the Population Architecture using Genomics and Epidemiology (PAGE) Network
- Genetic and Functional Modularity of Activities in the Specification of Limb-Innervating Motor Neurons
- A Population Genetic Model for the Maintenance of R2 Retrotransposons in rRNA Gene Loci
- A Quartet of PIF bHLH Factors Provides a Transcriptionally Centered Signaling Hub That Regulates Seedling Morphogenesis through Differential Expression-Patterning of Shared Target Genes in
- A Genome-Wide Integrative Genomic Study Localizes Genetic Factors Influencing Antibodies against Epstein-Barr Virus Nuclear Antigen 1 (EBNA-1)
- Mutation of the Diamond-Blackfan Anemia Gene in Mouse Results in Morphological and Neuroanatomical Phenotypes
- Life, the Universe, and Everything: An Interview with David Haussler
- Alternative Oxidase Expression in the Mouse Enables Bypassing Cytochrome Oxidase Blockade and Limits Mitochondrial ROS Overproduction
- An Evolutionarily Conserved Synthetic Lethal Interaction Network Identifies FEN1 as a Broad-Spectrum Target for Anticancer Therapeutic Development
- The Flowering Repressor Underlies a Novel QTL Interacting with the Genetic Background
- Telomerase Is Required for Zebrafish Lifespan
- and Diversified Expression of the Gene Family Bolster the Floral Stem Cell Network
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Increased Maternal Genome Dosage Bypasses the Requirement of the FIS Polycomb Repressive Complex 2 in Arabidopsis Seed Development
- WNK1/HSN2 Mutation in Human Peripheral Neuropathy Deregulates Expression and Posterior Lateral Line Development in Zebrafish ()
- Synergistic Interaction of Rnf8 and p53 in the Protection against Genomic Instability and Tumorigenesis
- Dot1-Dependent Histone H3K79 Methylation Promotes Activation of the Mek1 Meiotic Checkpoint Effector Kinase by Regulating the Hop1 Adaptor
- A Heterogeneous Mixture of F-Series Prostaglandins Promotes Sperm Guidance in the Reproductive Tract
- Starvation, Together with the SOS Response, Mediates High Biofilm-Specific Tolerance to the Fluoroquinolone Ofloxacin
- Directed Evolution of a Model Primordial Enzyme Provides Insights into the Development of the Genetic Code
- Genome-Wide Screens for Tinman Binding Sites Identify Cardiac Enhancers with Diverse Functional Architectures
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



