-
Články
Reklama
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
ReklamaSingle-Stranded Annealing Induced by Re-Initiation of Replication Origins Provides a Novel and Efficient Mechanism for Generating Copy Number Expansion via Non-Allelic Homologous Recombination
Copy number expansions such as amplifications and duplications contribute to human phenotypic variation, promote molecular diversification during evolution, and drive the initiation and/or progression of various cancers. The mechanisms underlying these copy number changes are still incompletely understood, however. We recently demonstrated that transient, limited re-replication from a single origin in Saccharomyces cerevisiae efficiently induces segmental amplification of the re-replicated region. Structural analyses of such re-replication induced gene amplifications (RRIGA) suggested that RRIGA could provide a new mechanism for generating copy number variation by non-allelic homologous recombination (NAHR). Here we elucidate this new mechanism and provide insight into why it is so efficient. We establish that sequence homology is both necessary and sufficient for repetitive elements to participate in RRIGA and show that their recombination occurs by a single-strand annealing (SSA) mechanism. We also find that re-replication forks are prone to breakage, accounting for the widespread DNA damage associated with deregulation of replication proteins. These breaks appear to stimulate NAHR between re-replicated repeat sequences flanking a re-initiating replication origin. Our results support a RRIGA model where the expansion of a re-replication bubble beyond flanking homologous sequences followed by breakage at both forks in trans provides an ideal structural context for SSA–mediated NAHR to form a head-to-tail duplication. Given the remarkable efficiency of RRIGA, we suggest it may be an unappreciated contributor to copy number expansions in both disease and evolution.
Published in the journal: . PLoS Genet 9(1): e32767. doi:10.1371/journal.pgen.1003192
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1003192Summary
Copy number expansions such as amplifications and duplications contribute to human phenotypic variation, promote molecular diversification during evolution, and drive the initiation and/or progression of various cancers. The mechanisms underlying these copy number changes are still incompletely understood, however. We recently demonstrated that transient, limited re-replication from a single origin in Saccharomyces cerevisiae efficiently induces segmental amplification of the re-replicated region. Structural analyses of such re-replication induced gene amplifications (RRIGA) suggested that RRIGA could provide a new mechanism for generating copy number variation by non-allelic homologous recombination (NAHR). Here we elucidate this new mechanism and provide insight into why it is so efficient. We establish that sequence homology is both necessary and sufficient for repetitive elements to participate in RRIGA and show that their recombination occurs by a single-strand annealing (SSA) mechanism. We also find that re-replication forks are prone to breakage, accounting for the widespread DNA damage associated with deregulation of replication proteins. These breaks appear to stimulate NAHR between re-replicated repeat sequences flanking a re-initiating replication origin. Our results support a RRIGA model where the expansion of a re-replication bubble beyond flanking homologous sequences followed by breakage at both forks in trans provides an ideal structural context for SSA–mediated NAHR to form a head-to-tail duplication. Given the remarkable efficiency of RRIGA, we suggest it may be an unappreciated contributor to copy number expansions in both disease and evolution.
Introduction
Duplication or amplification of chromosomal segments is important for evolution, phenotypic variation, human genetic disorders, and cancer [1]–[5]. Many of these duplications or amplifications are arranged in direct tandem repeat and have homologous sequence elements at their boundary, suggesting they were formed through recombination between non-allelic homologous sequences. Evidence for such non-allelic homologous recombination (NAHR) events is found in the genomes of nearly all species, including humans, where almost half of the human genome is comprised of low or high copy number repeat sequences [6]–[8].
The mechanisms responsible for these duplications or amplifications have been difficult to discern because these events are usually too rare to characterize their molecular intermediates. Nonetheless, studies primarily in microorganisms have led to a number of models for how these duplications/amplifications might arise. The most established model assumes that these NAHR events occur through the same fundamental mechanism as allelic homologous recombination [9]–[11]. In this model NAHR is initiated by a simple DNA double-strand break (DSB) in a repeat sequence, which normally provokes a homology search for the intact allelic counterpart as a repair template. An imperfect search arising from misalignment of sister chromatids or homologs, however, would lead to establishment of a double Holliday junction structure between non-allelic homologous sequences that can resolve into an unequal crossover. Evidence of the reciprocal copy number expansions and contractions expected to arise from such unequal crossing over is limited, having only been observed in the context of large tandem arrays of rDNA [12], [13] or CUP1 repeats [14], at subtelomeric repeats [15], and in some human genetic disorders [16]. A recent variation on this model suggests that NAHR-mediated tandem duplications/amplifications may be generated by break-induced replication (BIR) [17]. In this model, a broken chromosomal end initiates strand invasion and replication fork assembly at a non-allelic homologous sequence. The fork then duplicates the chromosomal segment between the homologous sequences before proceeding to the end of the chromosome. Again, direct support for this model is minimal.
Recently, we demonstrated that re-replication of a chromosomal segment due to dysregulation of replication controls can efficiently induce NAHR-medicated tandem duplication/amplification of that segment in the budding yeast Saccharomyces cerevisiae [18]. Importantly, introduction of simple DSBs failed to induce duplication/amplification with similar efficiency. Our findings raise the possibility that an alternative mechanism initiated by the loss of replication control might be responsible for some NAHR-mediated tandem duplications/amplifications. We have thus been eager to elucidate the mechanism of re-replication induced gene amplification (RRIGA).
The initiation of eukaryotic DNA replication is controlled by a battery of overlapping mechanisms that prevent re-initiation of DNA replication from the hundreds to thousands of replication origins in eukaryotic genomes. Replication initiation normally occurs at these origins in a two-stage process [19], [20]. In G1 phase, origins are licensed for initiation by loading the Mcm2-7 core replicative helicase onto them, a process that requires the origin recognition complex (ORC), Cdc6, Cdt1, and Mcm2-7. During S-phase, licensed origins are triggered to initiate DNA replication. To ensure that none of these origins re-initiate, multiple mechanisms inhibit ORC, Cdc6, Cdt1, and Mcm2-7 to minimize the chance that origins will be relicensed after they have initiated [19]–[21]. Consistent with the non-redundant nature of these controls, experimentally inactivating increasing numbers of these mechanisms leads to progressively increasing amounts of re-initiation and re-replication in budding yeast [22], [23].
These replication controls are critical for cell viability and genome stability. When sufficient controls are disrupted to cause overt re-replication (i.e. an increase in genomic DNA content detectable by flow cytometry), extensive DNA damage and a major DNA damage response is observed [24]–[31]. While the source of damage is not well understood, the amount of damage apparently overwhelms the DNA damage response and leads to massive cell death. In budding yeast, we developed the ability to induce and detect much lower levels of re-replication compatible with cell viability [23]. Retention of viability allowed us to examine the effect of re-replication on genome stability [18]. We found that limited, transient re-replication of a chromosomal segment induced tandem duplication and occasionally higher order amplification of that segment at a rate approximating 10−2 per cell division, about five orders of magnitude higher than spontaneous duplication rates [17]. The tandem duplications were bounded by Ty retrotransposon elements, a class of repetitive elements scattered throughout the yeast genome [32].
Here we uncover the mechanism of this re-replication induced gene amplification. These studies show that re-replication induces DNA damage because re-replication forks are highly susceptible to breakage. Our data support a model for RRIGA in which the two forks of a re-replication bubble both proceed beyond repetitive sequence elements flanking the re-initiating origin and then break. Should these breaks occur in trans with respect to the chromosome axis, normal 5′ to 3′ strand resection at each break will expose complementary strands of the non-allelic repetitive elements, providing a ready substrate for recombination by a single strand annealing (SSA) mechanism. Such repair of these broken re-replication bubbles will result in a tandem duplication arranged in direct repeat. In this model both the susceptibility of re-replication forks to breakage and the special structural context provided by the re-replication bubble contribute to the extraordinary efficiency of RRIGA. Importantly, the critical event triggering the formation of these tandem direct duplications is re-initiation of DNA replication within the duplicated segment. The remarkably efficient channeling of these re-initiation events into tandem direct duplications raises the possibility that even rare spontaneous re-initiation events may be a potent source of copy number variation in evolution and disease.
Results
Homology at Amplicon Boundaries Is Necessary and Sufficient for RRIGA
RRIGA generates gene duplications and amplifications arrayed in head-to-tail orientation at the original chromosomal locus with boundaries corresponding to Ty retrotransposable elements [18]. We previously reported that the inter-amplicon junctions generated by RRIGA had hybrid sequences consistent with a non-allelic homologous recombination event between Ty retrotransposable elements that flank the re-initiating origin (Figure 1A(i)). The two Ty elements most frequently involved in our RRIGA experiments share a 1.3 kb region of 99% sequence identity where the recombination events occurred (Figure S1A).
Fig. 1. Homologous sequences are necessary and sufficient in cis to support RRIGA. 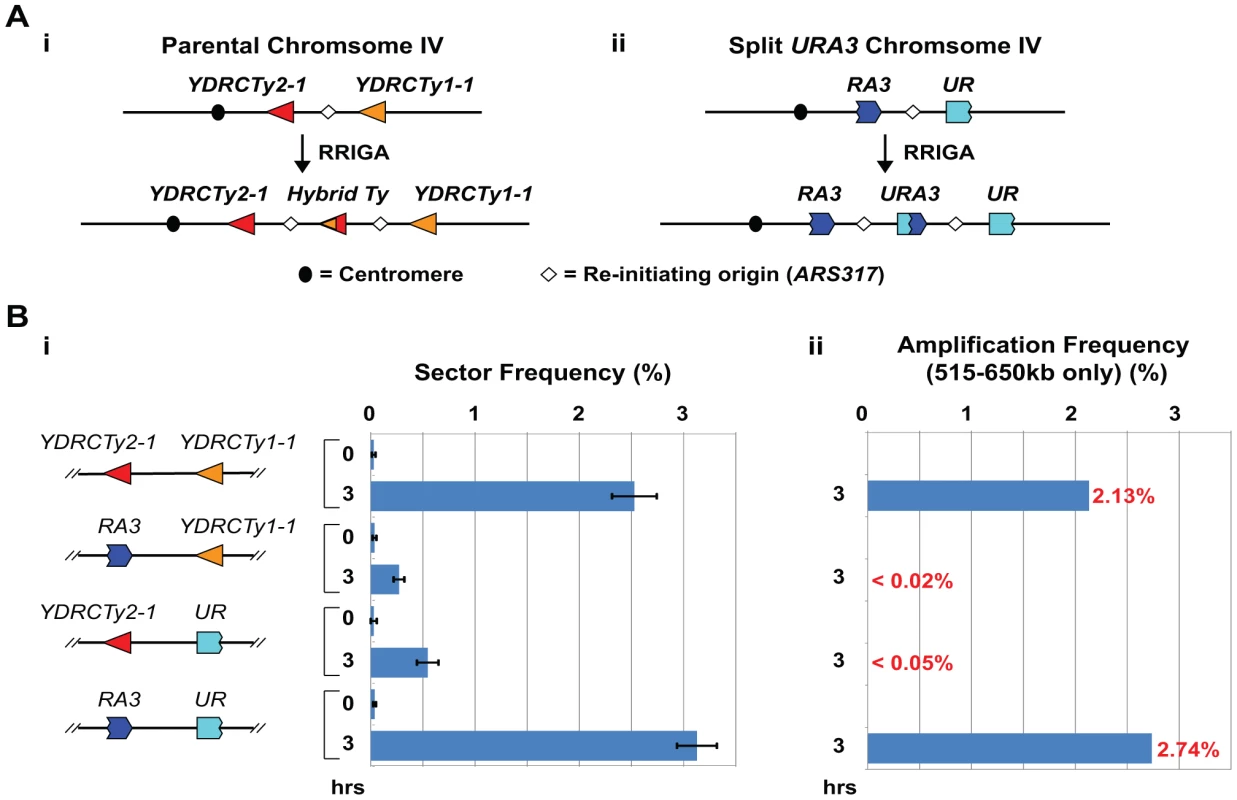
A) (i) Schematic of RRIGA arising from NAHR between YDRCTy2-1 at 515 kb and YDRCTy1-1 at 650 kb on Chromosome IV; (ii) Schematic of RRIGA arising from NAHR between 3′URA3 (RA3) and 5′URA3 (UR) fragments replacing YDRCTy2-1 and YDRCTy1-1, respectively (see Figure S1). Frequencies shown are for replacement of Ty and all adjacent LTR and tRNA sequences (version 2). B) (i) Sectoring frequencies (mean ± SEM, n = 3 to 5) before (0 hr) and after (3 hr) induction of re-replication in strains with endogenous Ty elements at 515 kb and 650 kb (YJL8100), with YDRCTy2-1 replaced by RA3 (YJL8355), with YDRCTy1-1 replaced by UR (YJL8359), or both Ty elements replaced with the respective URA3 fragments (YJL8363); (ii) Re-replication induced amplification frequency estimated by multiplying 3 hr sector frequency by fraction of sectors containing 515–650 kb amplification (see Figure S3). What we did not know was whether the homology between Ty elements is sufficient to promote RRIGA or whether other Ty-associated elements or biology are also important. Most Ty elements, including those involved in our RRIGA studies, are surrounded by tRNA genes and long terminal repeats (LTRs) in inverted orientation. These associated elements are known to cause replication forks to pause and possibly to break [33], [34], disruptions that could stimulate recombination. Hence, if such associated elements are important for RRIGA, it might constrain RRIGA to specific repetitive elements in budding yeast. On the other hand, if homology is sufficient for sequences to serve as RRIGA boundary elements, RRIGA could offer a potential mechanism for a broad range of NAHR-mediated copy number variations.
To address this question we used our previously described RRIGA assay, which exploits colony sectoring [35] to screen for amplification events [18]. In this assay, an origin particularly prone to re-initiate (ARS317) when Cdc6, Orc6, and the MCM complex are deregulated is integrated at 567 kb on Chromosome IV, along with a color based copy number reporter gene (ade3-2p). Cells with a single copy of ade3-2p are pink, while those with two or more copies are red. After transiently inducing re-initiation at ARS317 during a nocodazole arrest (G2/M), cells are plated for single colonies and possible amplification events are identified from pink colonies with red sectors that comprise 1/2–1/8 of the colony. We then verify and characterize amplifications in the red sectors by array Comparative Genomic Hybridization (aCGH). The vast majority of amplifications identified using this assay span the region from 515–650 kb on Chromosome IV, with YDRCTy2-1 and YDRCTy1-1 at the left and right boundaries, respectively. These are the closest Ty elements flanking the re-initiating ARS317 origin at 567 kb, and both are surrounded by tRNA genes and LTRs (Figure S1A).
To determine whether homology is sufficient to support RRIGA, we constructed strains in which: (1) YDRCTy2-1 was replaced by a 3′ portion of the URA3 gene; (2) YDRCTy1-1 was replaced by a 5′ portion of the URA3 gene; or (3) both Ty elements were replaced by their respective URA3 gene fragments (Figure S1). Two versions of these strains were generated. In version 1 some of the adjacent LTRs were replaced along with each Ty element, but tRNA genes and inverted LTR repeats were preserved (Figure S1B). In version 2, all of the adjacent tRNA genes and LTRs were replaced along with each Ty element (Figure S1C). Importantly, the URA3 fragments share a 390 bp overlapping region of 100% sequence identity. Thus, sequence homology was present at positions 515 kb and 650 kb on Chromosome IV in the strains in which both endogenous Ty elements were intact, as well as the strains in which both Ty elements were replaced by URA3 fragments. In contrast, no significant homology was present at these loci in strains in which only one Ty element was replaced by a URA3 fragment, and we refer to these as non-homologous boundary strains.
The non-homologous boundary strains showed a 5 - to 10-fold decrease in sector frequency (Figure 1B(i), Figure S2A, Table S1). Subsequent aCGH analysis of a dozen residual sectors induced in version 2 of each of these non-homologous boundary strains failed to detect any amplifications with endpoints at 515 kb and 650 kb on Chromosome IV (Figure S3, Table S2). Thus, when RRIGA frequencies for the 515–650 kb segment were estimated by multiplying sector frequencies by the percent of sectors that amplified this segment, there was at least a 50 - to 100-fold reduction in frequency (Figure 1B(ii)). We note that many red sectors derived from the strain without the right hand Ty element at 650 kb (YDRCTy1-1) did have an extra copy of the ade3-2p reporter, but achieved this either by using Ty elements further to the right as the right hand RRIGA boundary element, or through translocation or aneuploidy. In contrast, most red sectors derived from the strain without the left hand Ty element at 515 kb (YDRCTy2-1) did not have an extra copy of the ade3-2p reporter, presumably because there are no other Ty elements on Chromosome IV to serve as left-hand RRIGA boundaries (these red sectors presumably arose from other genomic changes that altered the rate of red pigment accumulation in ade3-2p cells). These findings confirmed that homology at the boundaries of amplicons is necessary for efficient RRIGA in budding yeast.
More importantly, when sequence homology was restored by replacing the remaining Ty element with the appropriate URA3 fragment, RRIGA frequencies were also restored. In strains with Ty elements at both 515 kb and 650 kb replaced by either version 1 or version 2 of the overlapping URA3 fragments, sectoring occurred at a frequency comparable to the strain with endogenous Ty elements intact (Figure 1B(i), Figure S2A, Table S1). Furthermore, most (14/16) of the red-sectors that were examined by aCGH bore an amplification of the 515 kb to 650 kb region of Chromosome IV (Figure S3, Table S2). Importantly, RRIGA frequency in the context of overlapping URA3 fragments was unaffected by the presence or absence of the tRNA genes or LTRs (compare Figure 1B and Figure S2A, Figure S2B). Thus, sequence homology at the boundaries of amplicons is sufficient to support RRIGA. Given the prevalence of homologous repetitive elements in eukaryotic genomes [36], this finding implies that these genomes are a potentially rich source of substrates for re-replication induced gene amplification.
Development of a Selection Assay for RRIGA
The fact that RRIGA in budding yeast results in NAHR between homologous sequences flanking a re-initiating origin allowed us to develop a more rapid and sensitive selection-based assay for quantifying RRIGA. We designed the URA3 fragments replacing the Ty elements at 515 kb and 650 kb such that NAHR between the fragments during RRIGA reconstitutes a full length, functional URA3 gene at the inter-amplicon junction (Figure 1A(ii)). Thus, in addition to scoring RRIGA between these two endpoints by colony sectoring, we could select for these events on media lacking uracil (Figure 2A, Table S1, Table S3). We note that the selection assay consistently gave a higher frequency than the sectoring assay, most likely because our visual criterion restricted the sectoring assay to capturing amplification events that occurred within 2–3 generations of cell plating (see Text S1).
Fig. 2. A selection-based assay for detecting RRIGA events. 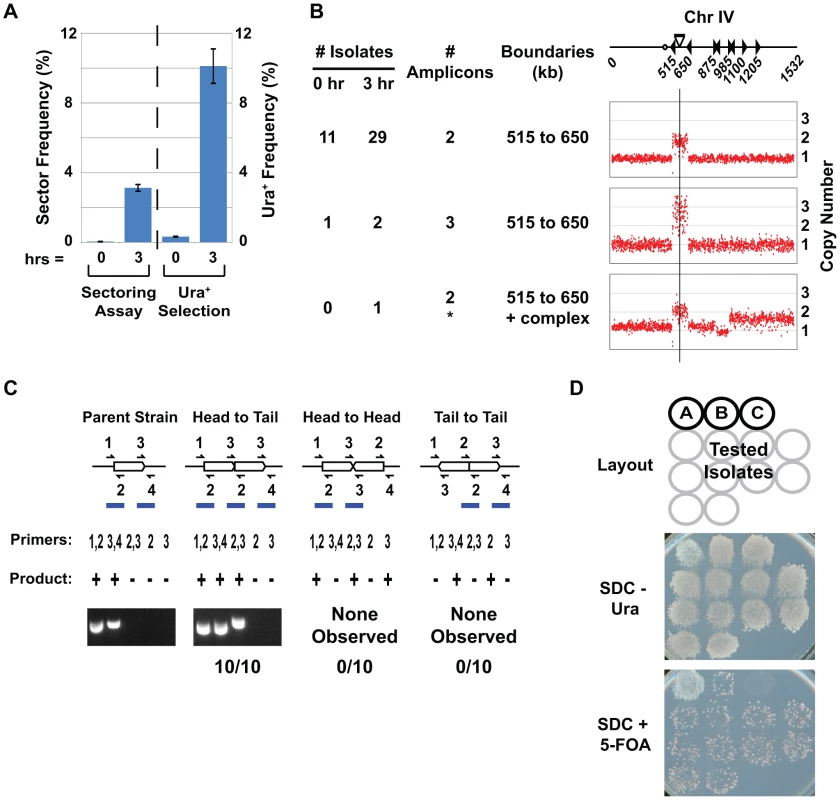
A) Comparison of RRIGA frequencies (mean ± SEM) measured using sectoring assay (n = 4) or URA3 selection assay (n = 5) in strains (YJL8363/8364) with YDRCTy2-1 and YDRCTy1-1 replaced by URA3 fragments as described for Figure 1B. B) Isolates from URA3 selection assay have amplifications spanning the segment between URA3 fragments (Chromosome IV 515–650 kb). aCGH copy number analysis of Chromosome IV shown for 12 isolates selected before (0 hr) and 32 isolates selected after (3 hr) re-replication from YJL8112/8113 and YJL8363/8364. Chromosome IV schematic shows position and orientation of Ty elements (triangles), centromere (circle), and ARS317-ade3-2p re-initiation cassette (bar and vertical line). C) Isolates from URA3 selection assay have amplifications tandemly arrayed in loco in direct repeat. The unamplified parental amplicon and three possible orientations for tandem duplications in loco are shown schematically. Predicted PCR junction fragments are shown for five sets of primers that flank amplicon boundaries (+, PCR product expected; −, no PCR product expected). Representative PCR products are shown for parental strain YJL8363 and 10 re-replication-induced isolates from B. D) Amplicons appear to be chromosomally integrated but excisable. The 10 isolates tested by PCR in C were grown on non-selective media (YEPD), then replica plated to media lacking uracil (SDC-Ura) or media containing 5-FOA (SDC+5-FOA). Patch A, YJL8698 with extrachromosomal copy of URA3. Patch B, YJL6974 with integrated but excisable copy of URA3. Patch C, YJL8344 with integrated and un-excisable URA3. We characterized the genetic alterations in the URA3 prototrophs recovered from our selection assay to ensure that they structurally resembled the RRIGA amplifications previously recovered from the sectoring assay. aCGH demonstrated that all prototrophs did indeed bear an amplification that spans the region from 515–650 kb on Chromosome IV (Figure 2B, Table S4). PCR across potential amplicon junctions confirmed that the original amplicon boundaries were intact and that the regenerated URA3 gene was created from a new head-to-tail amplicon junction (Figure 2C). Such a junction could arise from tandem intrachromosomal amplicons in head-to-tail orientation, as previously observed for RRIGA, but could also arise from circularization of an extrachromosomal amplicon via NAHR between the two URA3 fragments. These two possibilities can be distinguished by the spontaneous loss rate of the regenerated URA3 gene, because the latter will be lost at a much higher rate than the former. This loss rate can be estimated by the frequency of cells lacking URA3 (and thus resistant to the drug 5-fluoroorotic acid (5-FOA)) that accumulate in a population when selection for the gene is removed. As shown in Figure 2D, all the URA3 prototrophs obtained from our selection assay accumulated 5-FOA resistance at a frequency expected for an intrachromosomal amplification. Thus, the amplifications detected using the URA3 selection assay were structurally identical to those observed using the sectoring screen.
RRIGA Occurs through a Single-Strand Annealing Mechanism
Three major forms of homologous recombination have been characterized in budding yeast and shown to have distinct genetic dependencies (Figure 3A): gene conversion (GC), break induced replication (BIR), and single-strand annealing (SSA) [37]. BIR can be further subdivided into a form that requires the RecA homolog Rad51 and one that is independent of Rad51 [38]. We could thus narrow down the form of homologous recombination responsible for RRIGA by using the URA3 selection assay to quantify the dependence of RRIGA on various recombination genes. Because the frequency of RRIGA is dependent on the amount of induced re-replication, we normalized the measured frequency against the height of the induced re-replication peak (Figure S4A)
Fig. 3. RRIGA is primarily mediated by single-stranded annealing (SSA). 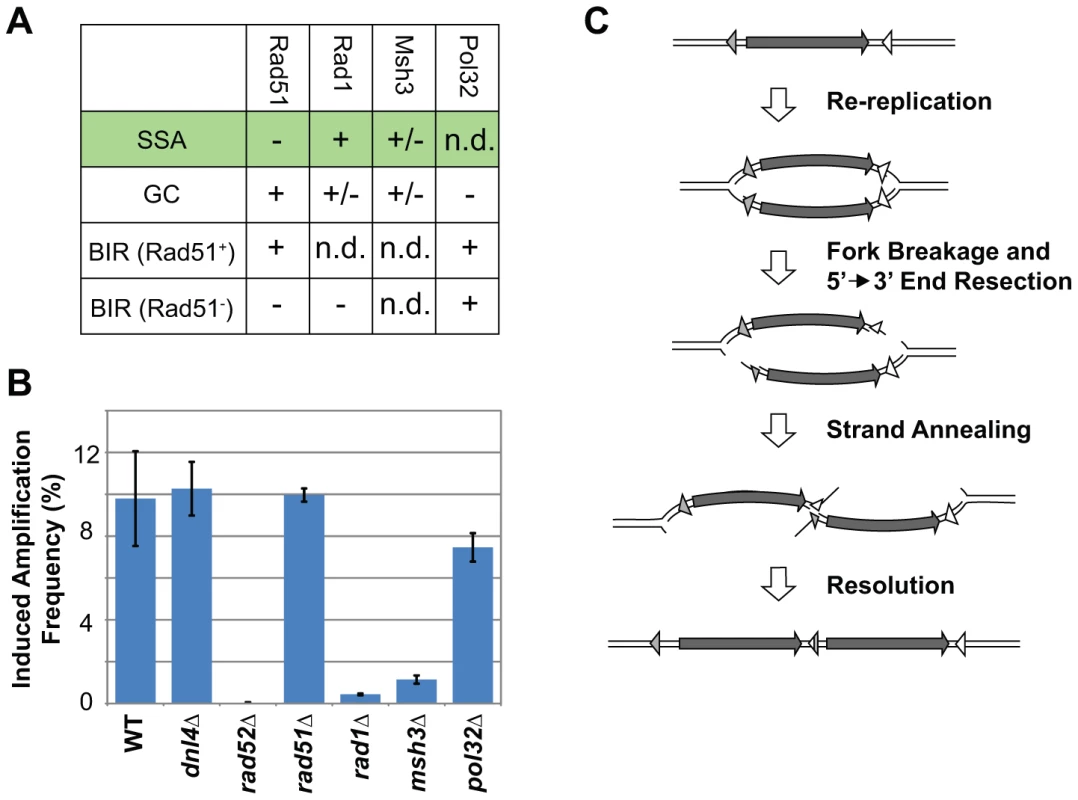
A) Summary of genetic requirements for the major sub-types of homologous recombination. SSA = Single Stranded Annealing; GC = Gene Conversion; BIR = Break Induced Replication. “+” = required; “−” = not required; “+/−” = required in some but not all cases; “n.d.” = not determined. B) RRIGA amplification frequencies for WT (YJL8363/8364), dnl4Δ (YJL8407/8408), rad52Δ (YJL8409/8410); rad51Δ (YJL8412/8413), rad1Δ (YJL8415/8416), msh3Δ (YJL8418/8419), and pol32Δ (YJL8421/8422) strains using the URA3 selection assay. Difference in frequency after 3 hr and 0 hr induction of re-replication was normalized for differences in the amount of re-initiation (see Text S1 and Figure S4). Data are presented as the mean ± combined error (see Text S1). C) Model for RRIGA involving SSA-mediated NAHR. Arrow, amplified segment. Triangles, non-allelic homologous sequences. The genetic dependencies for RRIGA most closely resembled those for SSA (Figure 3B, Table S3). First, RRIGA was independent of Rad51, which is required for strand invasion in GC and Rad51-dependent BIR but is not required for SSA [39]–[44]. Second RRIGA was dependent on Rad1 and Msh3. The former functions as part of the Rad1-Rad10 structure specific endonuclease, which removes non-homologous 3′ tails during SSA. The latter functions as part of the Msh2-Msh3 complex to stabilize the SSA structure that is recognized by Rad1-Rad10 [45]–[47]. Finally, RRIGA was independent of Pol32, a non-essential subunit of DNA Polymerase δ that is important for BIR [48]. Similar results for RAD51 and RAD1 were observed using the colony sectoring assay, although a partial dependence on Rad51 suggests that a subset of these RRIGA events may require this protein (Figure S4B, S4C, Table S1). Taken together, these results indicate that most of the NAHR observed in RRIGA is mediated by SSA.
Such a central role for SSA both restricts the possible mechanisms for RRIGA and expands the genetic alterations associated with SSA. SSA is almost always associated with deletion of chromosomal segments that lie between flanking homologous sequences [37]. A break between those sequences followed by 5′ end resection past both sequences allows them to anneal and initiate repair through NAHR, but at the cost of deleting the intervening segment. In the context of a re-replication bubble, however, SSA could generate a tandem duplication if both forks of the bubble travel beyond flanking homologous sequences and break in trans relative to the chromosome axis. The dual fork breaks would cleave the re-replicated sister chromatid in two, leaving a copy of its re-replicated portion at each broken end. Subsequent 5′ end resection back toward the re-replicated homologous sequences closest to each end would then expose complementary strands of these non-allelic sequences for annealing and SSA repair, resulting in a head-to-tail tandem duplication in loco (Figure 3C). Such a model provides the most straightforward explanation for how SSA can be responsible for RRIGA. Moreover, in this model, the special context provided by the re-replication bubble to exploit SSA for tandem duplications suggests one reason why re-replication is such a potent inducer of gene amplification.
Re-Replication Generates DSBs Distal to Both Flanking Repetitive Elements
A key requirement in our SSA model for RRIGA is that each re-replication fork must break origin-distal to the homologous sequence element that will undergo NAHR (Figure 3C). Although re-replication is known to induce double-strand breaks (DSBs), or at least a robust DNA damage response, in most cases the source of those breaks is unknown and actual breakage of re-replication forks has not been directly implicated [24]–[31]. We therefore asked whether there is a correspondence between the position of DSBs and re-replication forks and whether DSB do in fact arise distal to both flanking repetitive elements.
To map the location of DSBs that arise during re-replication from ARS317, we sized chromosomal fragments generated by these breaks using pulsed field gel electrophoresis (PFGE) (Figure 4, Figure S5). By preparing genomic DNA from cells embedded within agar plugs, this technique minimizes breakage from in vitro manipulations. Cells were harvested for PFGE after inducing re-replication for 0, 3, or 6 hr; as a control we harvested cells from a congenic non-re-replicating strain at the same time points. Prior to PFGE, chromosomal DNA was digested with the I-SceI endonuclease, which cuts a single unique I-SceI recognition site engineered very close to ARS317. This digest divides Chromosome IV into two fragments containing sequences to the left and right of ARS317, respectively. For those molecules that re-initiated from ARS317, the digestion will convert the resulting bubble intermediates into left and right Y-shaped chromosome fragments with ARS317 near the arm tips, telomere at the stem base, and re-replication fork at the branch point. Hence, a DSB in the re-replicated segment will cleave off an arm of the Y, generating a truncated chromosomal fragment whose length defines the position of the break relative to ARS317 (Figure 4A). After size separation by PFGE, these fragments were detected by southern analysis using probes just to the left or right of the I-SceI cut site (Figure 4B, Figure S5).
Fig. 4. Re-replication induces double-stranded DNA breaks distal to flanking repetitive elements. 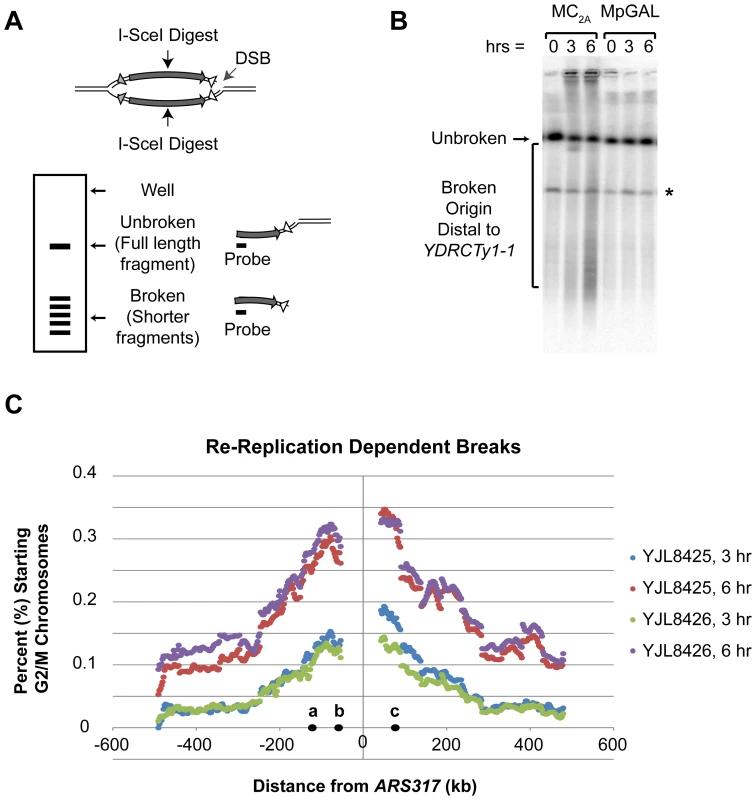
A) Strategy for mapping DSBs arising during re-replication using an I-SceI site near the re-initiating origin ARS317 as a physical reference point (see Text S1). The lengths of the small linear fragments generated by DSBs indicate the position of the DSBs relative to the I-SceI cut site. B) Representative Southern blot for DSBs induced by rightward moving re-replication forks. Re-replicating MC2A strains (YJL8425/YJL8426) were induced to re-replicate and at the indicated times chromosomal DNA was prepared, digested with I-SceI, size-separated by PFGE, Southern blotted, and probed for fragments extending rightward from the I-SceI site. Genomic DNA from non-re-replicating MpGAL control strains (YJL8427/8428) were processed in parallel. Unbroken: full-length Chromosome IV fragment from I-SceI site to right telomere. Bracket: fragments due to DSBs that map origin-distal to YDRCTy1-1. * unidentified DNA fragment present independent of re-replication. C) Distribution of DSBs induced by re-replication from ARS317 (see Text S1 and Figure S5). For each 2425 bp size range, the amount of re-replication induced fragmentation within that size range is displayed as a percent of the total amount of G2/M chromosomes before re-replication was induced. Positions are relative to ARS317 (at 0 kb) with positions of CEN4 (a), YDRCTy2-1 (b), and YDRCTy1-1 (c) indicated. Using this approach, we found that re-replication dependent DSBs did indeed arise with significant frequency on both sides of ARS317 (Figure 4B, Figure S5). Full-length right and left fragments from I-SceI-digested Chromosome IV were detected as discrete bands. Truncated fragments arising from DSBs migrated as a smear representing a range of sizes below the full-length fragments. These truncated fragments were specific to the re-replicating strain and became more abundant with longer induction of re-replication. Quantifying the amount of each fragment length relative to the starting amount of G2/M chromosomes before re-replication (0 hr) allowed us to estimate the percent of these chromosomes that acquired a DSB at each chromosomal position as a consequence of re-replication (see Text S1). Figure 4C shows a plot of this DSB percentage as a function of distance from ARS317. The distribution formed a broad peak centered about the origin similar to the distribution of re-replication forks around ARS317 (Figure S4A: WT). The similarity of these distributions is consistent with the notion that the DSBs arise from breakage of re-replication forks.
Importantly, many of the DSBs we mapped arose origin-distal to the two flanking repetitive Ty elements that are closest to ARS317 and that participate most frequently in RRIGA. We suspect our analysis undercounts DSB formation because some re-replication forks may not break until after the re-replication induction period, and some forks that break early in this period may already have been repaired. Nonetheless, the data provide a ballpark estimate of the percent of G2 chromosomes that acquire a double strand break beyond the most proximal Ty element as a consequence of re-replication. After 3 hr of re-replication this estimate is roughly 10–15% for either side of ARS317 (see Text S1). After 6 hr of re-replication the estimate is roughly 30–45%. Thus, these breaks are not rare, and there is a reasonable probability that a re-replication bubble will break at both forks at the positions needed to stimulate the use of homologous sequences in our model for SSA-mediated RRIGA.
Re-Replication Forks Must Proceed beyond Flanking Repetitive Elements for RRIGA
Our model predicts that the further away a homologous sequence is from the origin, the lower the frequency of RRIGA involving that sequence, as fewer re-replication forks will be able to reach that sequence and break beyond it. The fact that RRIGA amplicons preferentially arise from NAHR between the two closest Ty elements flanking the re-initiating origin as endpoints is consistent with this prediction. However, to test this prediction directly we used the URA3 selection assay to quantify the frequency of RRIGA in a series of strains where the flanking URA3 fragments were placed at increasing distance from ARS317 (Figure 5A).
Fig. 5. Flanking repetitive elements must be re-replicated in order for RRIGA to occur. 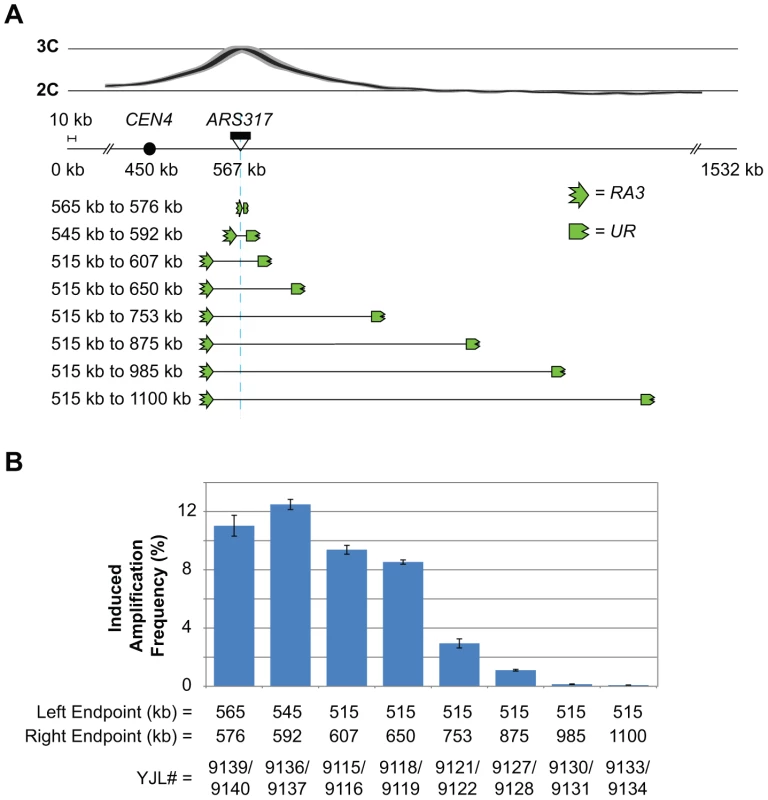
A) Schematic showing relocation of the RA3 and UR elements (3′ and 5′ portions of URA3, respectively) used in the URA3 RRIGA selection assay to change their position relative to the re-initiating origin (ARS317, light blue line) and the distribution of re-replication forks (as inferred from the re-replication profile). B) Induced amplification frequencies (mean ± SEM, n = 3) for strains with the indicated amplicon boundaries as defined by the position of the URA3 fragments. Induced frequency is frequency after 3 hr re-replication minus frequency after 0 hr re-replication. The overall trend supports the prediction. As the flanking URA3 fragments are moved further away from ARS317, RRIGA frequencies drop (Figure 5B, Table S3). RRIGA isolates from each starting strain were examined by aCGH to confirm that their amplicons did indeed extend from one URA3 fragment to the other (Figure S6A, Table S4). Importantly, despite the large range of amplicon sizes (11 kb to 585 kb) there was no detectable difference in the growth rates of these isolates (Figure S6B). Hence, the decrease in RRIGA frequencies cannot be explained by a decrease in fitness of those RRIGA isolates with larger amplicons. Instead these results support the requirement for forks to replicate and break beyond flanking homologous sequences.
RRIGA Proceeds Most Efficiently with the Re-Initiating Origin within the Amplicon
In the SSA model of RRIGA, after a re-replication fork breaks origin-distal to a homologous sequence element, 5′ end resection from the break must proceed back to the homologous sequence to make it available for SSA. With resection rates in S. cerevisiae estimated at 4 kb per hour [49], breaks that arise tens of kilobases past the homologous sequence will require many hours of resection before they can facilitate RRIGA, increasing the likelihood that the break will be repaired by an alternative mechanism or fail to occur before chromosomes finally segregate. Hence, one might expect some constraint on how far a fork break can occur beyond a homologous sequence and still stimulate the use of that sequence for RRIGA.
Such a constraint would influence the optimum position of a re-initiating origin relative to homologous boundaries of a potential amplicon. One would predict that RRIGA should be more efficient when the origin is within the amplicon than when it is outside (see Figure 6A). In the latter case, any distance traveled by the re-replication fork that initially moves away from the amplicon will have to be completely retraced during resection followed by further resection from the origin to the closest boundary.
Fig. 6. RRIGA proceeds most efficiently when the re-initiating origin lies within the amplicon. 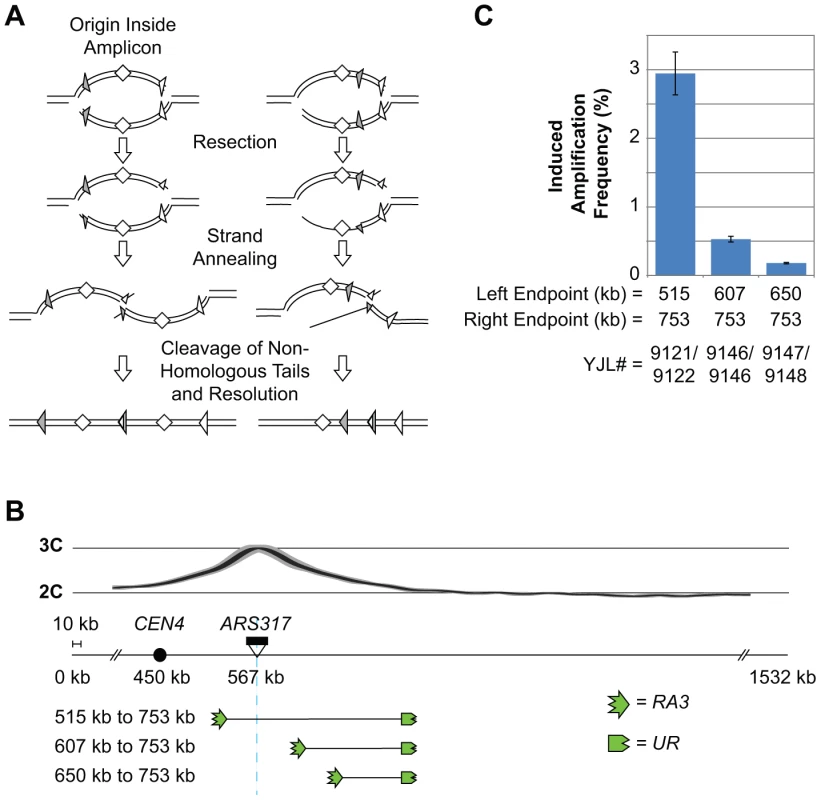
A) Schematic comparing SSA models for RRIGA when the re-initiating origin is within the amplicon versus when the origin is outside. The latter case requires long range strand resection back beyond the re-initiating origin in order to expose homologous sequences for NAHR. B) Schematic showing relocation of the RA3 element to change its position relative to the re-initiating origin (ARS317, light blue line). The re-replication profile is shown above. C) Induced amplification frequencies (mean ± SEM, n = 3) for strains with the indicated amplicon boundaries. Induced amplification frequency calculated as described for Figure 5B. To test this prediction, we generated a series of re-replicating strains in which the right amplicon boundary was held fixed while the left amplicon boundary lay either to the left of ARS317 (positioning the origin within the amplicon) or at two sites to the right of ARS317 (positioning the origin outside of the amplicon) (Figure 6B). In this series, the rightward re-replication fork has to travel the longest distance to reach the right homologous sequence boundary, and this distance is unchanged. In contrast, the leftward re-replication fork has little or no distance to travel to get past the left homologous sequence boundaries, but wherever it might break the resection distance back to those boundaries increases. In accordance with the prediction, the RRIGA frequency tracks inversely with the anticipated resection distance. The frequency is highest for the strain with the origin contained within the amplicon and becomes progressively lower as the left amplicon boundary is positioned further to the right of the origin (Figure 6C, Table S3).
Because the size of the amplicons varied in this series of strains, we also compared strains with relatively constant amplicon size in which the re-initiating origin was effectively repositioned outside of the amplicon (Figure S7A). In one set of strains, amplicon boundaries approximately 140 kb apart were moved to the right of ARS317, causing a precipitous drop in RRIGA (Figure S7B, Table S3). Although part of this drop can be attributed to the increased distance re-replication forks have to travel to the rightmost boundary (see Figure 5B, strains YJL9118/9119 versus strains YJL9121/9122), the remainder is likely due to the repositioning of the origin outside of the amplicon (compare Figure S7B strains YJL9145/9146 versus Figure 5B strains YJL9121/9122). Similarly, in strains where the amplicon size is maintained at ∼100 kb there is a dramatic decrease in RRIGA frequency when the origin is positioned outside of the amplicon (see Figure S7B, strains YJL9115/9116 versus YJL9147/9148). Thus, as expected from the SSA model for RRIGA, a re-initiating origin is most efficient at inducing amplification if the origin lies between the homologous sequences that define the amplicon boundaries.
Discussion
A Model for Re-Replication Induced Gene Amplification
We have previously shown that re-replication in budding yeast is remarkably efficient at inducing NAHR events that result in tandem gene amplifications oriented in direct repeat [18]. A transient, localized, limited pulse of re-replication from a single origin induced segmental amplifications on the order of 10−2 per cell per generation. This efficient amplification appeared to be specific to re-replication, as disruption of S-phase replication with mutant replication proteins or hydroxyurea did not induce equivalent amplification frequencies. In this paper, we propose a model for re-replication induced gene amplification (RRIGA) that helps explain why this amplification is so efficient and that provides a new mechanism for NAHR-mediated copy number variation. Such efficiency makes it conceivable that rare or sporadic re-replication events might contribute to DNA copy number changes observed during oncogenesis or evolution.
In our model for RRIGA (Figure 3C), bidirectional re-replication forks proceeding outward from a re-initiating origin can stimulate an efficient NAHR event between flanking homologous sequence elements by replicating beyond them and generating DSBs. Normal processing of these breaks will involve 5′ to 3′ single-strand resection back toward the homologous sequences. In those cases where the two forks break in trans, this resection can expose complementary sequences in non-allelic homologous sequences, resulting in annealing and repair of the break by an SSA mechanism. The result is a head-to-tail tandem duplication at the endogenous chromosomal locus. Such tandem duplications can provide a stepping stone for higher order amplifications [50]. Expansion of the duplication might occur readily without further re-replication, as the initially duplicated segments provide a much larger NAHR substrate. On the other hand, if re-replication were to recur in subsequent generations, it could stimulate a series of stepwise expansions that would lead to multi-copy amplification.
Re-Replication Fork Breakage Drives RRIGA
An important premise for our RRIGA model is the ability of re-replication to induce frequent chromosomal breaks. There are many reports associating the deregulation of replication initiation proteins with the generation of chromosomal breaks or the induction of a DNA damage response [24]–[31]. However, in most cases, this deregulation has been imposed constitutively throughout the cell cycle, making it hard to distinguish whether these breaks are due to re-replication per se or arise from possible disruption of S-phase replication. Because we induced re-replication after completion of an intact S-phase, the chromosomal breaks we observed and mapped can be specifically attributed to re-replication. Importantly, the correspondence between the distribution of breaks and the distribution of re-replication forks along the chromosome suggests that these forks are the source of these DSBs.
Formally, it is possible that the DSBs we mapped were actually the free ends of newly synthesized DNA fragments extruded by head-to-tail fork collisions during multiple rounds of re-replication, as has been proposed to explain induction of a DNA damage response during re-replication in Xenopus extracts [51]. However, this scenario is unlikely in our gene amplification studies, where we induced on average only half a round of re-replication near ARS317 (i.e. copy number increase from 2C to 3C). Hence, it appears that the re-replication forks themselves are breaking, leading to chromosome fragmentation.
The distribution of re-replication-induced breaks did not reveal any striking hotspots, indicating that these breaks do not depend on special DNA elements or structures that are suspected of potentiating DSB formation by promoting fork stalling and/or collapse [52]–[54]. The independence of these breaks from the inverted LTR repeats and multiple tRNA genes that often surround Ty elements is consistent with our ability to replace entire clusters of these elements with simple homologous sequences and still observe high frequency RRIGA.
Our results therefore raise the possibility that re-replication forks are particularly susceptible to breakage. Supporting this notion is our previous observation that the induction of re-replication can lead to a rapid and massive Rad9-dependent DNA damage response, a response that is not seen during unperturbed S-phase [24]. In fact, even when S-phase was subjected to prolonged disruption from hydroxyurea, it did not generate the type of chromosome fragmentation that was readily detected during re-replication [24]. Hence, unlike S-phase, where breakage among thousands of replication forks is a rare accident, during re-replication fork breakage may be the rule rather than the exception.
Clearly, an important future question will be why forks are so susceptible to breakage during re-replication. Nonetheless, the fact that they are increases the likelihood that re-initiation will lead to a bubble with a break at both forks. Our rough order of magnitude estimate of chromosome breakage frequencies on a single side of ARS317 (10–15% after 3 hr of re-replication and 30–45% after 6 hr) suggests that such dual fork breaks occur with significant frequency. Thus, even the slightest amount of re-replication may be a potent source of copy number variation.
The Structural Context Provided by the Re-Replication Bubble Also Facilitates RRIGA
Another key feature of our model that contributes to the efficiency of RRIGA is the structural context provided by the re-replication bubble. First, this structure provides an extra, non-essential chromosomal segment in close proximity to the endogenous segment. Second, when both forks happen to break in trans relative to the chromosome axis, this structure channels recombinational repair of the broken ends toward the formation of tandem duplications. Consistent with the importance of this structural context, simply inducing DSBs alone is insufficient to induce amplifications in our assay [18]. DSBs have been implicated in promoting NAHR events that result in the formation of deletions, translocations, and isochromosome formation, but they rarely lead to the type of intrachromosomal amplifications so efficiently induced by re-replication [10], [55]. DSBs are also capable of initiating a mechanism of gene amplification referred to as breakage-fusion-bridge (BFB), but the resulting amplification structure is very different from RRIGA, with amplicons oriented in inverted repeat and a terminal deletion beyond the amplified locus [56]–[59]. Thus, without the context of the re-replication bubble, DSBs do not show the same propensity for forming tandem duplications as observed during RRIGA.
The broader context in which re-replication bubbles appear may also contribute to the efficiency of RRIGA. The multiple overlapping layers of replication controls used by eukaryotic cells [18]–[20] ensure that only a limited number of origins will re-initiate when some of these controls are disrupted. The isolation of the resulting re-replication bubbles increases the likelihood that both forks of a bubble will eventually stall as they run into problems or simply reach the limits of their processivity. Without converging forks from nearby re-replication bubbles to rescue them, these stalled forks will be even more susceptible to breakage. Thus, we anticipate that a large proportion of re-initiation events will form the broken bubble intermediate that can be channeled into tandem direct amplifications.
In contrast, although S-phase replication bubbles are structurally identical to re-replication bubbles, the redundancy of active and backup origins in S-phase [60], [61] ensures that replication bubbles rarely arise in isolation and any fork that happens to stall is readily rescued by a converging fork from a neighboring replication bubble. This may explain why, despite the proposed link between S-phase accidents and various genomic rearrangements [62]–[64], inhibiting origin function or stressing S-phase forks throughout the genome with mutant replication proteins or DNA synthesis inhibitors does not generate RRIGA-like amplification frequencies [18]. To generate isolated replication bubbles, the origin failure or fork stress would have to be so severe that replication would be catastrophically and lethally disrupted. Examples of more tolerated localized replication fork stress have been identified through the discovery of common fragile sites (CFS) [65], [66]. However, individual CFSs can only affect one of the two forks from an expanding replication bubble and, hence, would not generate the broken bubble intermediate central to our RRIGA model. Consistent with this expectation, although some CFSs have been associated with amplifications, the amplifications are not arranged in tandem direct repeat like RRIGA. Instead they are arranged as inverted repeats and are thought to arise through a BFB mechanism initiated by a CFS break [67]–[69].
There is one example of a mutation that disrupts S-phase replication and induces segmental duplications structurally similar to RRIGA amplifications. Deletion of CLB5, one of the S-phase cyclins that triggers origin firing in budding yeast, reduces or delays origin activity primarily in the later replicating regions of the genome [70]. This deletion significantly stimulates tandem direct duplications involving NAHR at one chromosomal locus [17]. The mechanism by which clb5Δ stimulates these duplications is unknown, but whether or not it occurs through a RRIGA-like mechanism, the duplication rate is still more than one hundred fold lower (7.3×10−5 per cell division [17]) than the rate of RRIGA induced by ARS317 re-initiation. Altogether, these observations suggest that origin re-initiation may be particularly efficient at inducing amplifications in direct repeat because it is particularly efficient at generating isolated re-replication bubbles broken at both forks.
A Wider Set of Amplification Structures from Re-Replication?
In principle, once re-replication has generated re-replication bubbles broken at both forks in trans, subsequent formation of tandem direct duplications can occur by a variety of available repair mechanisms. In budding yeast, where homologous recombination predominates, we have shown that this repair occurs primarily through SSA-mediated NAHR. However, in other organisms, repair could conceivably proceed through alternative mechanisms that don't require extensive sequence homology, such as non-homologous end-joining (NHEJ) or microhomology mediated end-joining (MMEJ). Interestingly, numerous tandem duplications with little or no sequence homology at their interamplicon junctions have been observed in both normal and cancerous human genomes [16], [71], [72]. Hence, we are currently investigating whether re-replication can induce these types of duplications as well.
One can also imagine that if both forks of a re-replication bubble break, they may often break in cis, releasing one arm of the re-replication bubble and a full length chromosome. In that event, re-circularization of the arm, whether by SSA-mediated NAHR or some other repair mechanism, would generate a circular extrachromosomal amplicon, such as those frequently seen in human tumors [73]. We did not see evidence of extrachromosomal amplification in our system (Figure 2D), but our amplicons lacked centromeres, and in budding yeast, such amplicons would both be very unstable and preferentially accumulate to high, potentially toxic, copy number in mother cells [74]. It will thus be interesting to see if re-replication can also efficiently stimulate extrachromosomal amplification of centromere containing segments, such as the amplifications reported by Libuda and Winston [75].
Implications for Disease and Evolution
Our work provides a basis for understanding why re-replication arising from the loss of replication controls can generate duplications and amplifications with such remarkable efficiency. A much harder question to address is does re-replication contribute to gene amplifications and tandem duplications, such as those observed in cancers? Three observations suggest that this question is worth pursuing. First, dysregulation of replication initiation proteins has been observed in human cancer cells [4], [76]–[81]. Second, overexpression of replication initiation proteins in certain murine models can promote oncogenesis [81]–[83]. Third, a number of oncogene amplifications display amplicon structures with some or all of the features of yeast RRIGA structures: direct repeat, at the endogenous chromosomal locus, bounded by homologous sequence elements [84]–[87]. More recently, cancer genome sequencing efforts have detected the appearance of numerous tandem duplications in certain cancers [72]. Despite the lack of significant sequence homology at many of their boundaries, these duplications could still conceivably arise by some form of RRIGA, as discussed above. Importantly, the most popular model for gene amplification, the breakage-fusion-bridge mechanism [56], [57], cannot explain these amplifications/duplications in direct repeat, creating a need for alternative mechanisms.
Recently, it has been suggested that the deregulation of replication initiation might promote oncogenesis more directly through the induction of replication stress, resulting in extensive fork stalling, fork collapse, and DSBs [88]–[91]. Although the exact nature or source of this replication stress is not clear, our work documenting the extensive DNA damage and DSBs arising from re-replication forks makes re-replication a possible candidate.
Finally, we note that rare spontaneous tandem duplications and/or amplifications involving NAHR have been observed arising in yeast containing intact replication controls [17] (KJ Finn, unpublished results). These observations invite speculation that, despite intact controls, sporadic re-replication might still occur and cause gene copy number expansions that could promote evolution directly by generating phenotypic variation [11] or indirectly by removing constraints on molecular diversification [1]. It thus will be of interest to see whether these spontaneous copy number gains share some of the genetic dependencies of RRIGA established in this work.
Materials and Methods
Strains
All strains used in this study are listed in Table S5 are derived from YJL6974 and YJL6558 [18] using standard methods. Details of their construction, along with the plasmids (Table S6) and oligonucleotides (Table S5) used in their derivation, can be found in the Supplemental Material.
Strain Growth and Induction of Re-Replication
Yeast cells were grown as previously described [18]. Induction of re-replication was performed as previously described [18]. Full details are available in the Supplemental Material.
Colony Sectoring Assay
The colony sectoring assay was performed as previously described [18]. Full details are available in the Supplemental Material.
Uracil Prototrophy Assay
Following induction of re-replication ∼5,000 cfu were plated onto each SDC-Ura plate (to isolate uracil prototrophs) and ∼250 cfu onto each SDC plate (to determine an accurate cfu plated onto the SDC-Ura plates). Plates were incubated at 30°C for 3–5 days, then colonies were counted. The frequency of uracil prototrophs was determined by dividing the total number of colonies on the SDC-Ura plates by the number of cfus plated on the SDC-Ura plates. This frequency was measured in at least two independent experiments and the mean and standard error of the mean (when 3 or more trials were conducted), or the mean and the standard deviation (when only 2 trials were conducted) are reported.
Pulsed-Field Gel Electrophoresis and Southern Blotting
Cells were fixed and embedded in agarose for PFGE essentially as described [24], except that a proteinase K inactivation step with PMSF was included at the end of plug preparation. 1/3 of each plug was then treated with I-SceI to digest the embedded chromosomal DNA. Plugs were then loaded on a 1% SeaKem LE agarose (wt/vol) gel in 0.5× TBE. The gel was electrophoresed in 14°C 0.5× TBE on a CHEF DR-III system (Bio-Rad) with initial switch time of 50 sec, final switch time of 95 sec, run time of 26 hr, voltage of 6 V/cm, and angle of 120°. The DNA was transferred essentially as described [24], except UV-nicking was used instead of acid hydrolysis. The membrane was probed with a MAK21 probe generated by PCR from yeast genomic DNA with oligonucleotides OJL2449 and OJL2450, and a YOS9 probe generated by PCR from yeast genomic DNA with oligonucleotides OJL2231 and OJL2232. A Lambda probe was used to detect a sizing ladder. Images were collected using a Typhoon 9400 (GE Healthcare). Data analysis was carried out using Image J (NIH) and Excel (Microsoft Corp.) software. Full details are available in the Text S1.
aCGH
DNA used for aCGH was prepared as essentially described [18], [92]. Labeling, hybridization, data acquisition, and data analysis were performed as described [18]. Full details are available in the Supplemental Material.
Junction PCR
Primers used for junction PCR to determine amplicon orientation and preservation of parental junctions are listed in Table S7. PCR was performed using Phusion DNA polymerase (Finnzymes) according to the manufacturer's instructions. DNA used for junction PCR was prepared using a spheroplasting mini-prep method. Full details are available in the Supplemental Material.
Accession Numbers
All arrayCGH data from this study have been deposited in the Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.nih.gov/geo) database (Series Accession Number GSE41259).
Supporting Information
Zdroje
1. Ohno S (1970) Evolution by gene duplication. Springer-Verlag. 184 p.
2. SimmonsAD, CarvalhoCMB, LupskiJR (2012) What have studies of genomic disorders taught us about our genome? Methods Mol Biol 838 : 1–27 doi:10.1007/978-1-61779-507-7_1.
3. BeroukhimR, MermelCH, PorterD, WeiG, RaychaudhuriS, et al. (2010) The landscape of somatic copy-number alteration across human cancers. Nature 463 : 899–905 doi:10.1038/nature08822.
4. SantariusT, ShipleyJ, BrewerD, StrattonMR, CooperCS (2010) A census of amplified and overexpressed human cancer genes. Nat Rev Cancer 10 : 59–64 doi:10.1038/nrc2771.
5. GirirajanS, CampbellCD, EichlerEE (2011) Human copy number variation and complex genetic disease. Annu Rev Genet 45 : 203–226 doi:10.1146/annurev-genet-102209-163544.
6. VenterJC, AdamsMD, MyersEW, LiPW, MuralRJ, et al. (2001) The sequence of the human genome. Science 291 : 1304–1351 doi:10.1126/science.1058040.
7. LanderES, LintonLM, BirrenB, NusbaumC, ZodyMC, et al. (2001) Initial sequencing and analysis of the human genome. Nature 409 : 860–921 doi:10.1038/35057062.
8. LevyS, SuttonG, NgPC, FeukL, HalpernAL, et al. (2007) The diploid genome sequence of an individual human. PLoS Biol 5: e254 doi:10.1371/journal.pbio.0050254.
9. ChenJ-M, CooperDN, FérecC, Kehrer-SawatzkiH, PatrinosGP (2010) Genomic rearrangements in inherited disease and cancer. Semin Cancer Biol 20 : 222–233 doi:10.1016/j.semcancer.2010.05.007.
10. HoangML, TanFJ, LaiDC, CelnikerSE, HoskinsRA, et al. (2010) Competitive repair by naturally dispersed repetitive DNA during non-allelic homologous recombination. PLoS Genet 6: e1001228 doi:10.1371/journal.pgen.1001228.
11. HastingsPJ, LupskiJR, RosenbergSM, IraG (2009) Mechanisms of change in gene copy number. Nat Rev Genet 10 : 551–564 doi:10.1038/nrg2593.
12. SzostakJW, WuR (1980) Unequal crossing over in the ribosomal DNA of Saccharomyces cerevisiae. Nature 284 : 426–430 doi:10.1038/284426a0.
13. PetesTD (1980) Unequal meiotic recombination within tandem arrays of yeast ribosomal DNA genes. Cell 19 : 765–774 doi:10.1016/S0092-8674(80)80052-3.
14. WelchJW, MaloneyDH, FogelS (1990) Unequal crossing-over and gene conversion at the amplified CUP1 locus of yeast. Mol Gen Genet 222 : 304–310 doi:10.1007/BF00633833.
15. LouisEJ, HaberJE (1990) Mitotic recombination among subtelomeric Y′ repeats in Saccharomyces cerevisiae. Genetics 124 : 547–559.
16. LiuP, CarvalhoCM, HastingsP, LupskiJR (2012) Mechanisms for recurrent and complex human genomic rearrangements. Current Opinion in Genetics & Development 22 : 211–220 doi:10.1016/j.gde.2012.02.012.
17. PayenC, KoszulR, DujonB, FischerG (2008) Segmental duplications arise from Pol32-dependent repair of broken forks through two alternative replication-based mechanisms. PLoS Genet 4: e1000175 doi:10.1371/journal.pgen.1000175.
18. GreenBM, FinnKJ, LiJJ (2010) Loss of DNA replication control is a potent inducer of gene amplification. Science 329 : 943–946 doi:10.1126/science.1190966.
19. AriasEE, WalterJC (2007) Strength in numbers: preventing rereplication via multiple mechanisms in eukaryotic cells. Genes Dev 21 : 497–518 doi:10.1101/gad.1508907.
20. DiffleyJFX (2011) Quality control in the initiation of eukaryotic DNA replication. Philos Trans R Soc Lond, B, Biol Sci 366 : 3545–3553 doi:10.1098/rstb.2011.0073.
21. NguyenVQ, CoC, LiJJ (2001) Cyclin-dependent kinases prevent DNA re-replication through multiple mechanisms. Nature 411 : 1068–1073 doi:10.1038/35082600.
22. WilmesGM, ArchambaultV, AustinRJ, JacobsonMD, BellSP, et al. (2004) Interaction of the S-phase cyclin Clb5 with an “RXL” docking sequence in the initiator protein Orc6 provides an origin-localized replication control switch. Genes Dev 18 : 981–991 doi:10.1101/gad.1202304.
23. GreenBM, MorrealeRJ, OzaydinB, DerisiJL, LiJJ (2006) Genome-wide mapping of DNA synthesis in Saccharomyces cerevisiae reveals that mechanisms preventing reinitiation of DNA replication are not redundant. Mol Biol Cell 17 : 2401–2414 doi:10.1091/mbc.E05-11-1043.
24. GreenBM, LiJJ (2005) Loss of rereplication control in Saccharomyces cerevisiae results in extensive DNA damage. Mol Biol Cell 16 : 421–432 doi:10.1091/mbc.E04-09-0833.
25. MelixetianM, BallabeniA, MasieroL, GaspariniP, ZamponiR, et al. (2004) Loss of Geminin induces rereplication in the presence of functional p53. J Cell Biol 165 : 473–482 doi:10.1083/jcb.200403106.
26. ArchambaultV, IkuiAE, DrapkinBJ, CrossFR (2005) Disruption of mechanisms that prevent rereplication triggers a DNA damage response. Mol Cell Biol 25 : 6707–6721 doi:10.1128/MCB.25.15.6707-6721.2005.
27. VaziriC, SaxenaS, JeonY, LeeC, MurataK, et al. (2003) A p53-dependent checkpoint pathway prevents rereplication. Mol Cell 11 : 997–1008 doi:10.1016/S1097-2765(03)00099-6.
28. LovejoyCA, LockK, YenamandraA, CortezD (2006) DDB1 maintains genome integrity through regulation of Cdt1. Mol Cell Biol 26 : 7977–7990 doi:10.1128/MCB.00819-06.
29. JinJ, AriasEE, ChenJ, HarperJW, WalterJC (2006) A family of diverse Cul4-Ddb1-interacting proteins includes Cdt2, which is required for S phase destruction of the replication factor Cdt1. Mol Cell 23 : 709–721 doi:10.1016/j.molcel.2006.08.010.
30. ZhuW, DuttaA (2006) An ATR - and BRCA1-mediated Fanconi anemia pathway is required for activating the G2/M checkpoint and DNA damage repair upon rereplication. Mol Cell Biol 26 : 4601–4611 doi:10.1128/MCB.02141-05.
31. ZhuW, ChenY, DuttaA (2004) Rereplication by depletion of geminin is seen regardless of p53 status and activates a G2/M checkpoint. Mol Cell Biol 24 : 7140–7150 doi:10.1128/MCB.24.16.7140-7150.2004.
32. MieczkowskiPA, LemoineFJ, PetesTD (2006) Recombination between retrotransposons as a source of chromosome rearrangements in the yeast Saccharomyces cerevisiae. DNA Repair (Amst) 5 : 1010–1020 doi:10.1016/j.dnarep.2006.05.027.
33. DeshpandeAM, NewlonCS (1996) DNA replication fork pause sites dependent on transcription. Science 272 : 1030–1033 doi:10.1126/science.272.5264.1030.
34. VoineaguI, NarayananV, LobachevKS, MirkinSM (2008) Replication stalling at unstable inverted repeats: interplay between DNA hairpins and fork stabilizing proteins. Proc Natl Acad Sci USA 105 : 9936–9941 doi:10.1073/pnas.0804510105.
35. KoshlandD, KentJC, HartwellLH (1985) Genetic analysis of the mitotic transmission of minichromosomes. Cell 40 : 393–403 doi:10.1016/0092-8674(85)90153-9.
36. RichardG-F, KerrestA, DujonB (2008) Comparative genomics and molecular dynamics of DNA repeats in eukaryotes. Microbiol Mol Biol Rev 72 : 686–727 doi:10.1128/MMBR.00011-08.
37. PâquesF, HaberJE (1999) Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 63 : 349–404.
38. McEachernMJ, HaberJE (2006) Break-induced replication and recombinational telomere elongation in yeast. Annu Rev Biochem 75 : 111–135 doi:10.1146/annurev.biochem.74.082803.133234.
39. IraG, HaberJE (2002) Characterization of RAD51-independent break-induced replication that acts preferentially with short homologous sequences. Mol Cell Biol 22 : 6384–6392 doi:10.1128/MCB.22.18.6384-6392.2002.
40. SymingtonLS (2002) Role of RAD52 epistasis group genes in homologous recombination and double-strand break repair. Microbiol Mol Biol Rev 66 : 630–670 doi:10.1128/MMBR.66.4.630-670.2002.
41. LeS, MooreJK, HaberJE, GreiderCW (1999) RAD50 and RAD51 define two pathways that collaborate to maintain telomeres in the absence of telomerase. Genetics 152 : 143–152.
42. DavisAP, SymingtonLS (2004) RAD51-dependent break-induced replication in yeast. Mol Cell Biol 24 : 2344–2351 doi:10.1128/MCB.24.6.2344-2351.2004.
43. IvanovEL, SugawaraN, Fishman-LobellJ, HaberJE (1996) Genetic requirements for the single-strand annealing pathway of double-strand break repair in Saccharomyces cerevisiae. Genetics 142 : 693–704.
44. SugawaraN, IraG, HaberJE (2000) DNA length dependence of the single-strand annealing pathway and the role of Saccharomyces cerevisiae RAD59 in double-strand break repair. Mol Cell Biol 20 : 5300–5309 doi:10.1128/MCB.20.14.5300-5309.2000.
45. SugawaraN, PâquesF, ColaiácovoM, HaberJE (1997) Role of Saccharomyces cerevisiae Msh2 and Msh3 repair proteins in double-strand break-induced recombination. Proc Natl Acad Sci USA 94 : 9214–9219.
46. Fishman-LobellJ, HaberJE (1992) Removal of nonhomologous DNA ends in double-strand break recombination: the role of the yeast ultraviolet repair gene RAD1. Science 258 : 480–484 doi:10.1126/science.1411547.
47. LyndakerAM, AlaniE (2009) A tale of tails: insights into the coordination of 3′ end processing during homologous recombination. Bioessays 31 : 315–321 doi:10.1002/bies.200800195.
48. LydeardJR, JainS, YamaguchiM, HaberJE (2007) Break-induced replication and telomerase-independent telomere maintenance require Pol32. Nature 448 : 820–823 doi:10.1038/nature06047.
49. ZhuZ, ChungW-H, ShimEY, LeeSE, IraG (2008) Sgs1 helicase and two nucleases Dna2 and Exo1 resect DNA double-strand break ends. Cell 134 : 981–994 doi:10.1016/j.cell.2008.08.037.
50. RomeroD, PalaciosR (1997) Gene amplification and genomic plasticity in prokaryotes. Annu Rev Genet 31 : 91–111 doi:10.1146/annurev.genet.31.1.91.
51. DavidsonIF, LiA, BlowJJ (2006) Deregulated replication licensing causes DNA fragmentation consistent with head-to-tail fork collision. Mol Cell 24 : 433–443 doi:10.1016/j.molcel.2006.09.010.
52. CarrAM, PaekAL, WeinertT (2011) DNA replication: failures and inverted fusions. Semin Cell Dev Biol 22 : 866–874 doi:10.1016/j.semcdb.2011.10.008.
53. LabibK, HodgsonB (2007) Replication fork barriers: pausing for a break or stalling for time? EMBO Rep 8 : 346–353 doi:10.1038/sj.embor.7400940.
54. MirkinEV, MirkinSM (2007) Replication fork stalling at natural impediments. Microbiol Mol Biol Rev 71 : 13–35 doi:10.1128/MMBR.00030-06.
55. ArguesoJL, WestmorelandJ, MieczkowskiPA, GawelM, PetesTD, et al. (2008) Double-strand breaks associated with repetitive DNA can reshape the genome. Proc Natl Acad Sci USA 105 : 11845–11850 doi:10.1073/pnas.0804529105.
56. DebatisseM, MalfoyB (2005) Gene amplification mechanisms. Adv Exp Med Biol 570 : 343–361 doi:10.1007/1-4020-3764-3_12.
57. McClintockB (1942) The Fusion of Broken Ends of Chromosomes Following Nuclear Fusion. Proc Natl Acad Sci USA 28 : 458–463.
58. NarayananV, MieczkowskiPA, KimH-M, PetesTD, LobachevKS (2006) The pattern of gene amplification is determined by the chromosomal location of hairpin-capped breaks. Cell 125 : 1283–1296 doi:10.1016/j.cell.2006.04.042.
59. VanHulleK, LemoineFJ, NarayananV, DowningB, HullK, et al. (2007) Inverted DNA repeats channel repair of distant double-strand breaks into chromatid fusions and chromosomal rearrangements. Mol Cell Biol 27 : 2601–2614 doi:10.1128/MCB.01740-06.
60. DershowitzA, SnyderM, SbiaM, SkurnickJH, OngLY, et al. (2007) Linear derivatives of Saccharomyces cerevisiae chromosome III can be maintained in the absence of autonomously replicating sequence elements. Mol Cell Biol 27 : 4652–4663 doi:10.1128/MCB.01246-06.
61. BlowJJ, GeXQ (2009) A model for DNA replication showing how dormant origins safeguard against replication fork failure. EMBO Rep 10 : 406–412 doi:10.1038/embor.2009.5.
62. AguileraA, Gómez-GonzálezB (2008) Genome instability: a mechanistic view of its causes and consequences. Nat Rev Genet 9 : 204–217 doi:10.1038/nrg2268.
63. KolodnerRD, PutnamCD, MyungK (2002) Maintenance of genome stability in Saccharomyces cerevisiae. Science 297 : 552–557 doi:10.1126/science.1075277.
64. WeinertT, KaocharS, JonesH, PaekA, ClarkAJ (2009) The replication fork's five degrees of freedom, their failure and genome rearrangements. Curr Opin Cell Biol 21 : 778–784 doi:10.1016/j.ceb.2009.10.004.
65. DebatisseM, Le TallecB, LetessierA, DutrillauxB, BrisonO (2012) Common fragile sites: mechanisms of instability revisited. Trends Genet 28 : 22–32 doi:10.1016/j.tig.2011.10.003.
66. DurkinSG, GloverTW (2007) Chromosome fragile sites. Annu Rev Genet 41 : 169–192 doi:10.1146/annurev.genet.41.042007.165900.
67. CoquelleA, PipirasE, ToledoF, ButtinG, DebatisseM (1997) Expression of fragile sites triggers intrachromosomal mammalian gene amplification and sets boundaries to early amplicons. Cell 89 : 215–225 doi:10.1016/S0092-8674(00)80201-9.
68. HellmanA, ZlotorynskiE, SchererSW, CheungJ, VincentJB, et al. (2002) A role for common fragile site induction in amplification of human oncogenes. Cancer Cell 1 : 89–97 doi:10.1016/S1535-6108(02)00017-X.
69. CiulloM, DebilyM-A, RozierL, AutieroM, BillaultA, et al. (2002) Initiation of the breakage-fusion-bridge mechanism through common fragile site activation in human breast cancer cells: the model of PIP gene duplication from a break at FRA7I. Hum Mol Genet 11 : 2887–2894 doi:10.1093/hmg/11.23.2887.
70. McCuneHJ, DanielsonLS, AlvinoGM, CollingwoodD, DelrowJJ, et al. (2008) The temporal program of chromosome replication: genomewide replication in clb5{Delta} Saccharomyces cerevisiae. Genetics 180 : 1833–1847 doi:10.1534/genetics.108.094359.
71. MillsRE, WalterK, StewartC, HandsakerRE, ChenK, et al. (2011) Mapping copy number variation by population-scale genome sequencing. Nature 470 : 59–65 doi:10.1038/nature09708.
72. McBrideDJ, EtemadmoghadamD, CookeSL, AlsopK, GeorgeJ, et al. (2012) Tandem duplication of chromosomal segments is common in ovarian and breast cancer genomes. J Pathol 227 : 446–455 doi:10.1002/path.4042.
73. HahnPJ (1993) Molecular biology of double-minute chromosomes. Bioessays 15 : 477–484 doi:10.1002/bies.950150707.
74. MurrayAW, SzostakJW (1983) Pedigree analysis of plasmid segregation in yeast. Cell 34 : 961–970 doi:10.1016/0092-8674(83)90553-6.
75. LibudaDE, WinstonF (2006) Amplification of histone genes by circular chromosome formation in Saccharomyces cerevisiae. Nature 443 : 1003–1007 doi:10.1038/nature05205.
76. BondsL, BakerP, GupC, ShroyerKR (2002) Immunohistochemical localization of cdc6 in squamous and glandular neoplasia of the uterine cervix. Arch Pathol Lab Med 126 : 1164–1168 doi:10.1043/0003-9985(2002)126<1164:ILOCIS>2.0.CO;2.
77. BorladoLR, MéndezJ (2008) CDC6: from DNA replication to cell cycle checkpoints and oncogenesis. Carcinogenesis 29 : 237–243 doi:10.1093/carcin/bgm268.
78. KarakaidosP, TaravirasS, VassiliouLV, ZacharatosP, KastrinakisNG, et al. (2004) Overexpression of the replication licensing regulators hCdt1 and hCdc6 characterizes a subset of non-small-cell lung carcinomas: synergistic effect with mutant p53 on tumor growth and chromosomal instability–evidence of E2F-1 transcriptional control over hCdt1. Am J Pathol 165 : 1351–1365 doi:10.1016/S0002-9440(10)63393-7.
79. MurphyN, RingM, HeffronCCBB, KingB, KillaleaAG, et al. (2005) p16INK4A, CDC6, and MCM5: predictive biomarkers in cervical preinvasive neoplasia and cervical cancer. J Clin Pathol 58 : 525–534 doi:10.1136/jcp.2004.018895.
80. RenB, YuG, TsengGC, CieplyK, GavelT, et al. (2006) MCM7 amplification and overexpression are associated with prostate cancer progression. Oncogene 25 : 1090–1098 doi:10.1038/sj.onc.1209134.
81. LiontosM, KoutsamiM, SideridouM, EvangelouK, KletsasD, et al. (2007) Deregulated overexpression of hCdt1 and hCdc6 promotes malignant behavior. Cancer Res 67 : 10899–10909 doi:10.1158/0008-5472.CAN-07-2837.
82. ArentsonE, FaloonP, SeoJ, MoonE, StudtsJM, et al. (2002) Oncogenic potential of the DNA replication licensing protein CDT1. Oncogene 21 : 1150–1158 doi:10.1038/sj.onc.1205175.
83. SeoJ, ChungYS, SharmaGG, MoonE, BurackWR, et al. (2005) Cdt1 transgenic mice develop lymphoblastic lymphoma in the absence of p53. Oncogene 24 : 8176–8186 doi:10.1038/sj.onc.1208881.
84. KuwaharaY, TanabeC, IkeuchiT, AoyagiK, NishigakiM, et al. (2004) Alternative mechanisms of gene amplification in human cancers. Genes Chromosomes Cancer 41 : 125–132 doi:10.1002/gcc.20075.
85. HerrickJ, ContiC, TeissierS, ThierryF, CouturierJ, et al. (2005) Genomic organization of amplified MYC genes suggests distinct mechanisms of amplification in tumorigenesis. Cancer Res 65 : 1174–1179 doi:10.1158/0008-5472.CAN-04-2802.
86. O'NeilJ, TchindaJ, GutierrezA, MoreauL, MaserRS, et al. (2007) Alu elements mediate MYB gene tandem duplication in human T-ALL. J Exp Med 204 : 3059–3066 doi:10.1084/jem.20071637.
87. StroutMP, MarcucciG, BloomfieldCD, CaligiuriMA (1998) The partial tandem duplication of ALL1 (MLL) is consistently generated by Alu-mediated homologous recombination in acute myeloid leukemia. Proc Natl Acad Sci USA 95 : 2390–2395.
88. Di MiccoR, FumagalliM, CicaleseA, PiccininS, GaspariniP, et al. (2006) Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature 444 : 638–642 doi:10.1038/nature05327.
89. BartkovaJ, RezaeiN, LiontosM, KarakaidosP, KletsasD, et al. (2006) Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature 444 : 633–637 doi:10.1038/nature05268.
90. Dominguez-SolaD, YingCY, GrandoriC, RuggieroL, ChenB, et al. (2007) Non-transcriptional control of DNA replication by c-Myc. Nature 448 : 445–451 doi:10.1038/nature05953.
91. HalazonetisTD, GorgoulisVG, BartekJ (2008) An oncogene-induced DNA damage model for cancer development. Science 319 : 1352–1355 doi:10.1126/science.1140735.
92. HoffmanCS, WinstonF (1987) A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene 57 : 267–272.
Štítky
Genetika Reprodukční medicína
Článek Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across PathogensČlánek TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association StudiesČlánek Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization inČlánek Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA ExpressionČlánek The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of GenesČlánek The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2013 Číslo 1
-
Všechny články tohoto čísla
- A Model of High Sugar Diet-Induced Cardiomyopathy
- Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across Pathogens
- Emerging Function of Fat Mass and Obesity-Associated Protein (Fto)
- Positional Cloning Reveals Strain-Dependent Expression of to Alter Susceptibility to Bleomycin-Induced Pulmonary Fibrosis in Mice
- Genetics of Ribosomal Proteins: “Curiouser and Curiouser”
- Transposable Elements Re-Wire and Fine-Tune the Transcriptome
- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- MAML1 Enhances the Transcriptional Activity of Runx2 and Plays a Role in Bone Development
- Predicting Mendelian Disease-Causing Non-Synonymous Single Nucleotide Variants in Exome Sequencing Studies
- A Systematic Mapping Approach of 16q12.2/ and BMI in More Than 20,000 African Americans Narrows in on the Underlying Functional Variation: Results from the Population Architecture using Genomics and Epidemiology (PAGE) Study
- Transcription of the Major microRNA–Like Small RNAs Relies on RNA Polymerase III
- Histone H3K56 Acetylation, Rad52, and Non-DNA Repair Factors Control Double-Strand Break Repair Choice with the Sister Chromatid
- Genome-Wide Association Study Identifies a Novel Susceptibility Locus at 12q23.1 for Lung Squamous Cell Carcinoma in Han Chinese
- Genetic Disruption of the Copulatory Plug in Mice Leads to Severely Reduced Fertility
- The [] Prion Exists as a Dynamic Cloud of Variants
- Adult Onset Global Loss of the Gene Alters Body Composition and Metabolism in the Mouse
- Fis Protein Insulates the Gene from Uncontrolled Transcription
- The Meiotic Nuclear Lamina Regulates Chromosome Dynamics and Promotes Efficient Homologous Recombination in the Mouse
- Genome-Wide Haplotype Analysis of Expression Quantitative Trait Loci in Monocytes
- TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association Studies
- Structural Basis of a Histone H3 Lysine 4 Demethylase Required for Stem Elongation in Rice
- The Ecm11-Gmc2 Complex Promotes Synaptonemal Complex Formation through Assembly of Transverse Filaments in Budding Yeast
- MCM8 Is Required for a Pathway of Meiotic Double-Strand Break Repair Independent of DMC1 in
- Comparative Genomic Analysis of the Endosymbionts of Herbivorous Insects Reveals Eco-Environmental Adaptations: Biotechnology Applications
- Integration of Nodal and BMP Signals in the Heart Requires FoxH1 to Create Left–Right Differences in Cell Migration Rates That Direct Cardiac Asymmetry
- Pharmacodynamics, Population Dynamics, and the Evolution of Persistence in
- A Hybrid Likelihood Model for Sequence-Based Disease Association Studies
- Aberration in DNA Methylation in B-Cell Lymphomas Has a Complex Origin and Increases with Disease Severity
- Multiple Opposing Constraints Govern Chromosome Interactions during Meiosis
- Transcriptional Dynamics Elicited by a Short Pulse of Notch Activation Involves Feed-Forward Regulation by Genes
- Dynamic Large-Scale Chromosomal Rearrangements Fuel Rapid Adaptation in Yeast Populations
- Heterologous Gln/Asn-Rich Proteins Impede the Propagation of Yeast Prions by Altering Chaperone Availability
- Gene Copy-Number Polymorphism Caused by Retrotransposition in Humans
- An Incompatibility between a Mitochondrial tRNA and Its Nuclear-Encoded tRNA Synthetase Compromises Development and Fitness in
- Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization in
- Single-Stranded Annealing Induced by Re-Initiation of Replication Origins Provides a Novel and Efficient Mechanism for Generating Copy Number Expansion via Non-Allelic Homologous Recombination
- Tbx2 Controls Lung Growth by Direct Repression of the Cell Cycle Inhibitor Genes and
- Suv4-20h Histone Methyltransferases Promote Neuroectodermal Differentiation by Silencing the Pluripotency-Associated Oct-25 Gene
- A Conserved Helicase Processivity Factor Is Needed for Conjugation and Replication of an Integrative and Conjugative Element
- Telomerase-Null Survivor Screening Identifies Novel Telomere Recombination Regulators
- Genome-Wide Analysis Reveals Selection for Important Traits in Domestic Horse Breeds
- Coordinated Degradation of Replisome Components Ensures Genome Stability upon Replication Stress in the Absence of the Replication Fork Protection Complex
- Nkx6.1 Controls a Gene Regulatory Network Required for Establishing and Maintaining Pancreatic Beta Cell Identity
- HIF- and Non-HIF-Regulated Hypoxic Responses Require the Estrogen-Related Receptor in
- Delineating a Conserved Genetic Cassette Promoting Outgrowth of Body Appendages
- The Telomere Capping Complex CST Has an Unusual Stoichiometry, Makes Multipartite Interaction with G-Tails, and Unfolds Higher-Order G-Tail Structures
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Loci Associated with -Glycosylation of Human Immunoglobulin G Show Pleiotropy with Autoimmune Diseases and Haematological Cancers
- Switchgrass Genomic Diversity, Ploidy, and Evolution: Novel Insights from a Network-Based SNP Discovery Protocol
- Centromere-Like Regions in the Budding Yeast Genome
- Sequencing of Loci from the Elephant Shark Reveals a Family of Genes in Vertebrate Genomes, Forged by Ancient Duplications and Divergences
- Mendelian and Non-Mendelian Regulation of Gene Expression in Maize
- Mutational Spectrum Drives the Rise of Mutator Bacteria
- Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA Expression
- The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of Genes
- Sex-Specific Signaling in the Blood–Brain Barrier Is Required for Male Courtship in
- A Newly Uncovered Group of Distantly Related Lysine Methyltransferases Preferentially Interact with Molecular Chaperones to Regulate Their Activity
- Is Required for Leptin-Mediated Depolarization of POMC Neurons in the Hypothalamic Arcuate Nucleus in Mice
- Unlocking the Bottleneck in Forward Genetics Using Whole-Genome Sequencing and Identity by Descent to Isolate Causative Mutations
- The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
- MTERF3 Regulates Mitochondrial Ribosome Biogenesis in Invertebrates and Mammals
- Downregulation and Altered Splicing by in a Mouse Model of Facioscapulohumeral Muscular Dystrophy (FSHD)
- NBR1-Mediated Selective Autophagy Targets Insoluble Ubiquitinated Protein Aggregates in Plant Stress Responses
- Retroactive Maintains Cuticle Integrity by Promoting the Trafficking of Knickkopf into the Procuticle of
- Phenome-Wide Association Study (PheWAS) for Detection of Pleiotropy within the Population Architecture using Genomics and Epidemiology (PAGE) Network
- Genetic and Functional Modularity of Activities in the Specification of Limb-Innervating Motor Neurons
- A Population Genetic Model for the Maintenance of R2 Retrotransposons in rRNA Gene Loci
- A Quartet of PIF bHLH Factors Provides a Transcriptionally Centered Signaling Hub That Regulates Seedling Morphogenesis through Differential Expression-Patterning of Shared Target Genes in
- A Genome-Wide Integrative Genomic Study Localizes Genetic Factors Influencing Antibodies against Epstein-Barr Virus Nuclear Antigen 1 (EBNA-1)
- Mutation of the Diamond-Blackfan Anemia Gene in Mouse Results in Morphological and Neuroanatomical Phenotypes
- Life, the Universe, and Everything: An Interview with David Haussler
- Alternative Oxidase Expression in the Mouse Enables Bypassing Cytochrome Oxidase Blockade and Limits Mitochondrial ROS Overproduction
- An Evolutionarily Conserved Synthetic Lethal Interaction Network Identifies FEN1 as a Broad-Spectrum Target for Anticancer Therapeutic Development
- The Flowering Repressor Underlies a Novel QTL Interacting with the Genetic Background
- Telomerase Is Required for Zebrafish Lifespan
- and Diversified Expression of the Gene Family Bolster the Floral Stem Cell Network
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Increased Maternal Genome Dosage Bypasses the Requirement of the FIS Polycomb Repressive Complex 2 in Arabidopsis Seed Development
- WNK1/HSN2 Mutation in Human Peripheral Neuropathy Deregulates Expression and Posterior Lateral Line Development in Zebrafish ()
- Synergistic Interaction of Rnf8 and p53 in the Protection against Genomic Instability and Tumorigenesis
- Dot1-Dependent Histone H3K79 Methylation Promotes Activation of the Mek1 Meiotic Checkpoint Effector Kinase by Regulating the Hop1 Adaptor
- A Heterogeneous Mixture of F-Series Prostaglandins Promotes Sperm Guidance in the Reproductive Tract
- Starvation, Together with the SOS Response, Mediates High Biofilm-Specific Tolerance to the Fluoroquinolone Ofloxacin
- Directed Evolution of a Model Primordial Enzyme Provides Insights into the Development of the Genetic Code
- Genome-Wide Screens for Tinman Binding Sites Identify Cardiac Enhancers with Diverse Functional Architectures
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



