-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaCondensin II Promotes the Formation of Chromosome Territories by Inducing Axial Compaction of Polyploid Interphase Chromosomes
The eukaryotic nucleus is both spatially and functionally partitioned. This organization contributes to the maintenance, expression, and transmission of genetic information. Though our ability to probe the physical structure of the genome within the nucleus has improved substantially in recent years, relatively little is known about the factors that regulate its organization or the mechanisms through which specific organizational states are achieved. Here, we show that Drosophila melanogaster Condensin II induces axial compaction of interphase chromosomes, globally disrupts interchromosomal interactions, and promotes the dispersal of peri-centric heterochromatin. These Condensin II activities compartmentalize the nucleus into discrete chromosome territories and indicate commonalities in the mechanisms that regulate the spatial structure of the genome during mitosis and interphase.
Published in the journal: . PLoS Genet 8(8): e32767. doi:10.1371/journal.pgen.1002873
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002873Summary
The eukaryotic nucleus is both spatially and functionally partitioned. This organization contributes to the maintenance, expression, and transmission of genetic information. Though our ability to probe the physical structure of the genome within the nucleus has improved substantially in recent years, relatively little is known about the factors that regulate its organization or the mechanisms through which specific organizational states are achieved. Here, we show that Drosophila melanogaster Condensin II induces axial compaction of interphase chromosomes, globally disrupts interchromosomal interactions, and promotes the dispersal of peri-centric heterochromatin. These Condensin II activities compartmentalize the nucleus into discrete chromosome territories and indicate commonalities in the mechanisms that regulate the spatial structure of the genome during mitosis and interphase.
Introduction
It is widely accepted that interphase chromosomes occupy discrete sub-nuclear domains, known as chromosome territories (CTs). CTs have been described in animals, fungi, and plants [1]–[4], suggesting that they may have functional relevance for all eukaryotes. The spatial arrangement of these compartments is non-random and correlates with the functional compartmentalization of the genome, in which actively expressed genes are spatially segregated from silent genes and repetitive DNA sequences [5]–[8]. In light of recent findings, several models of the nucleus posit that CT structure plays a central role in regulating long distance DNA-DNA interactions, heterochromatic gene silencing, DNA repair, and genomic stability [9]–[11]. Despite this, little is known about the factors that regulate the higher levels of chromosome structure or the spatial relationships between different chromosomes.
Several aspects of nuclear architecture can vary across species and cell types, but a few general organizational patterns appear to be highly conserved. Within the nuclei of many mammalian, chicken, and Arabidopsis cells, CTs are roughly spherical and have dimensions that are much smaller than the nuclear diameter [4], [12]–[14]. Alternatively, the chromosomes of budding yeast, cereals, salamanders, and Ascaris often exist in polarized, elongated domains known as the Rabl configuration [3], [15]–[17]. This pattern resembles the organization of chromosomes during anaphase: the centromeres of all chromosomes are clustered together at one pole of the nucleus, telomeres are oriented toward the opposite pole, and chromosome arms are similar in length to the diameter of the nucleus [17], [18]. Although chromosomes in the Rabl configuration do occupy discrete regions of the nucleus (i.e. territories), we reserve the term CT to refer specifically to globular structures, such as those of typical mammalian nuclei.
In Drosophila, multiple chromosome organization schemes have been well characterized. The Rabl configuration is observed in many diploid nuclei such as those of the early embryo and the larval central nervous system [19], [20]. Polyploid-polytene chromosomes, such as those of the larval salivary gland, also exhibit features of the Rabl configuration [21], whereas spermatocytes have been shown to form non-Rabl, globular CTs in the G2-phase of meiosis I [22]. The polyploid nurse cells of the Drosophila ovary are exceptional in several aspects of their nuclear organization and dynamics. Their chromosomes appear polytene during the first few rounds of DNA replication, but they undergo a developmentally regulated reorganization. After the fifth endocycle, homology-dependent pairing interactions are disrupted [23]. In this arrangement known as the “5-blob stage”, each of the 5 major chromosome arms occupies a globular territory, similar in appearance to mammalian CTs. Since nurse cells also have some of the largest nuclei ever described, their structural features are readily observable by light microscopy, making them an attractive model for the study of nuclear architecture.
The molecular mechanisms that control chromosome organization are poorly understood; however, it is clear that Condensin complexes play a central role. In Drosophila, there are believed to be two distinct Condensin Complexes. The SMC2/SMC4 heterodimer is common to both Condensin I and Condensin II. Condensin I also contains Cap-G, Cap-D2, and the Kleisin subunit Cap-H, whereas Condensin II contains Cap-D3 and Cap-H2 [24]. While the canonical vertebrate Condensin II also has a Cap-G2 subunit, no Cap-G2 ortholog has been identified in Drosophila.
Condensins are ATPases that can alter DNA topology in vitro and have been shown to regulate chromosome structure during mitosis and meiosis [24]–[26]. Recently, it has been shown that Condensins also contribute to the structure of interphase chromosomes [27]. Since Condensin II is required to disrupt nurse cell polytene chromosome pairing near the transition to the 5-blob stage [28], we hypothesized that this complex may also contribute to the forces that drive the formation of globular CTs.
Results
Cap-H2 is required for nurse cell chromosome territory formation
To assess the impact of Condensin II on the spatial relationships between heterologous chromosomes, we performed FISH (fluorescence in situ hybridization) on wild type and Cap-H2Z3-0019/Df(3R)Exel6159 mutant ovaries using probes to simultaneously mark 12 unique loci: 5 probes spanning the X chromosome were labeled in green and 7 probes on the 2nd chromosome were labeled in red. On average, these probes were spaced ∼5 Mb apart. Confocal microscopy was then used to capture 3D images of nurse cell nuclei. Consistent with previous findings [23], wild type stage 4 nurse cell nuclei appeared polytene and tended to display a single fluorescent focus for each probe (Figure 1A). At stage 5, the chromatids that composed the polytene chromosomes began to unpair from one another and each set of homologous chromosomes became confined to a compact spherical CT (Figure 1B). Initially, each territory was distinguishable by DAPI staining alone due to the presence of a DNA-devoid region called the interchromatin compartment (Figure 1B). At later stages, territories were not apparent by DAPI staining but FISH revealed that CT structure was largely maintained throughout the subsequent endocycles and expansion of the nuclear volume (Figure 1C). In nurse cells carrying mutations in Cap-H2 (Cap-H2Z3-0019/Df(3R)Exel6159), a Condensin II subunit, a polytene organization was evident throughout development and compact spherical CTs were not visually apparent at any stage (Figure 1D–1F).
Fig. 1. The formation of compact chromosome territories is Condensin II dependent. 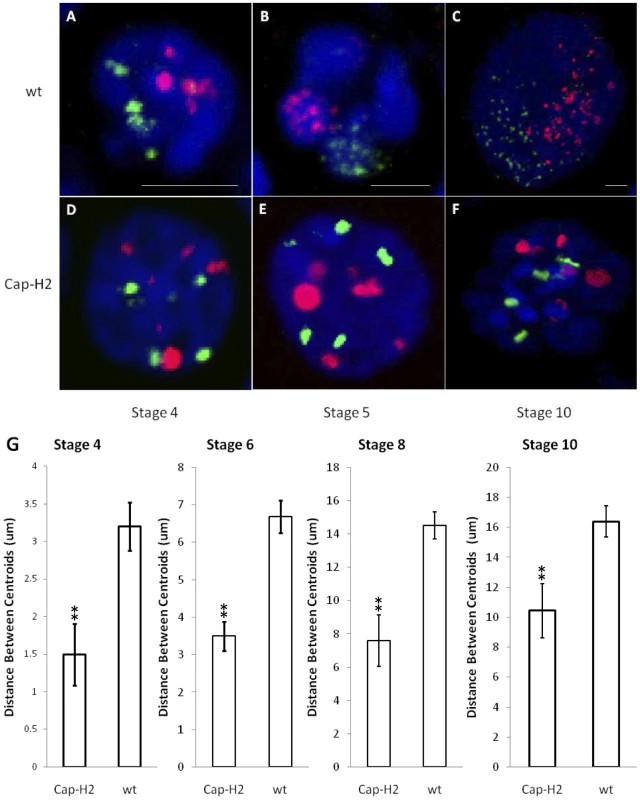
3D FISH was performed on nurse cell nuclei to mark the X chromosome in green and the 2nd chromosome in red. DAPI (DNA) is in blue. Representative nuclei from the following genotypes and developmental stages are depicted: A–C) wild type (y[1] w[67c23]) stages 4, 5, and 10; D–F) Cap-H2Z3-0019/Df(3R)6159 stages 4, 5, and 10. All scale bars are equal to 5 µm. G) Mean distances between X chromosome and 2nd chromosome centroids at stages 4, 6, 8, and 10. Error bars correspond to standard error. One asterisk signifies p<.05 and two asterisks signifies p<.01. Within each category, n = 15. The dispersed FISH signal in wild type nurse cells facilitated the visualization of CT boundaries. In Cap-H2Z3-0019/Df(3R)Exel6159 nurse cell nuclei, this signal was concentrated to a smaller number of foci (Figure 1). Thus, it remained possible that CTs were also present in this genotype but not readily visible. To address this, we computationally segmented the FISH signal from our 3D reconstructions and determined the geometric center of all FISH foci (centroid) for each chromosome-specific set of probes. The distances observed between the centroids of the X and 2nd chromosome probes provided a quantitative measure of the mean spatial separation between these chromosomes. At all stages of development, we found that the X and 2nd chromosomes were significantly more distant from each other in wild type nurse cells than in Cap-H2Z3-0019/Df(3R)Exel6159 (Figure 1G, p<0.01 for each stage). Based on this, we concluded that the compact CT structure observed in wild type nurse cells was dependent upon the presence of Condensin II.
Condensin II is required to inhibit the Rabl configuration and to disperse heterologous centromeres
We next sought to determine if nurse cell CTs were consistent with the Rabl configuration in which the centromeric regions of all chromosomes cluster together near the nuclear periphery and the telomeres cluster near the opposite pole. We performed FISH using 8 different probes (3 at a time) that targeted loci along the length of the X chromosome. For each of these loci, we determined its mean radial distance from the center of the nucleus. The center of each nucleus was determined based on the geometric center of all DAPI signal. The nuclear radius was then estimated based on the total volume of DAPI signal assuming that each nucleus was a sphere. These estimates correlated well with manual measurements of nurse cell nuclei (R2 = .87) and we were unable to detect any differences in total nuclear size in any Condensin II mutant backgrounds (Figure S1). All radial distances are reported as the distance from the center of nucleus divided by the nuclear radius.
In wild type nurse cell nuclei from stage 6, 8, and 10 egg chambers, the loci closest to the centromere were consistently near the maximal distance from the center of the nucleus (Figure 2A, Figure S2). Similarly, we observed that the X-chromosome heterochromatin FISH signal in Cap-H2Z3-0019/Df(3R)Exel6159, Cap-D3EY00456/Df(2L)Exel7023 and SMC4k08819/+; Cap-H2Z3-0019/+ nurse cells consistently localized near the nuclear periphery while loci near the middle of the chromosome arm were more centrally located and also exhibited a larger amount of variability in their radial positions (Figure 2A). We next performed 3D-FISH with sets of probes that differentially marked the peri-centric heterochromatin of the X, 2nd, 3rd, and 4th chromosomes. Again, we observed that peri-centric heterochromatin sequences consistently localized to the nuclear periphery in wild type and Cap-H2 mutant nuclei (Figure 2B–2E).
Fig. 2. Condensin II inhibits the Rabl conformation. 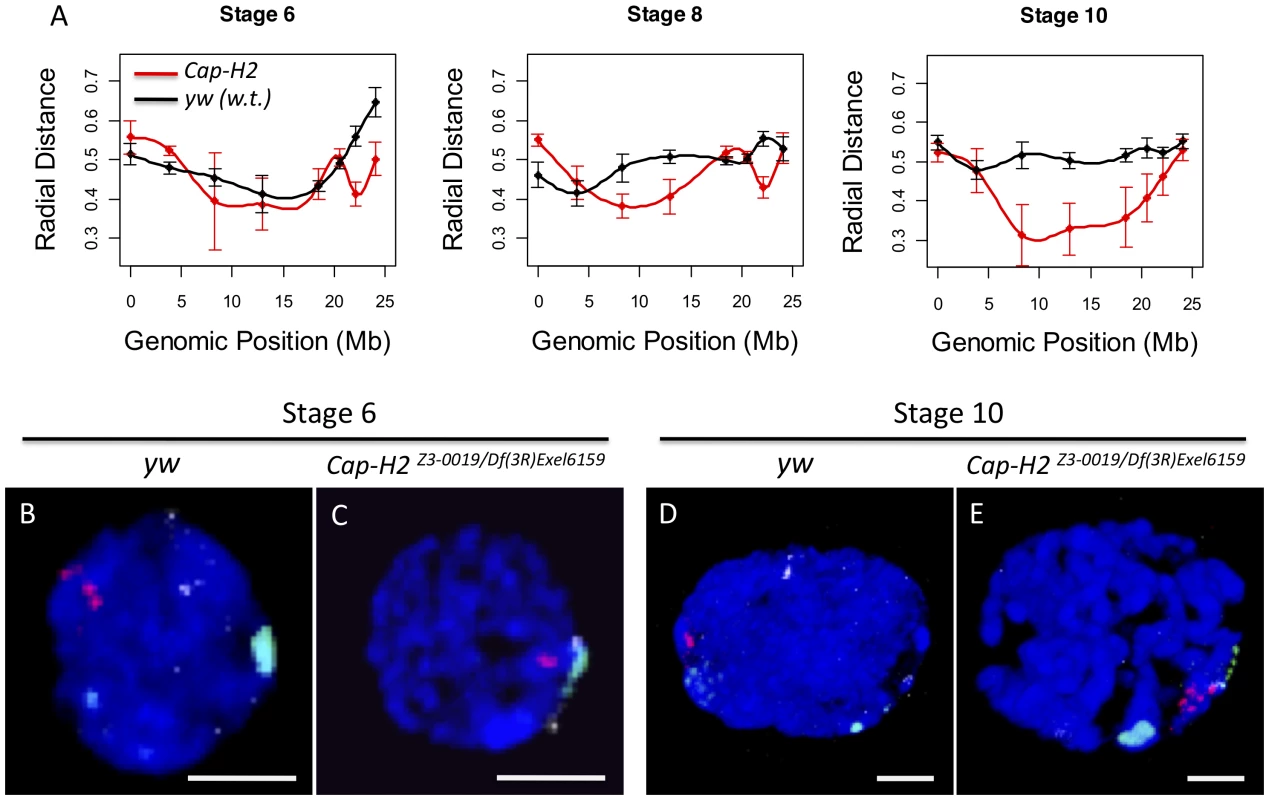
A) For each X chromosome probe, the radial distance from the center of nurse cell nuclei was measured at stages 6, 8, and 10. For each nucleus, the nuclear radius was estimated based on the volume of DAPI staining and the assumption that each nucleus was a sphere. All radial distances are reported as a fraction of the estimated nuclear radius with 0 corresponding to the center of the nucleus and 1 corresponding to the nuclear periphery. Genotypes are indicated by the legend in the bottom panel. Error bars correspond to standard error. B–E) Images of representative nuclei from wild type and Cap-H2Z3-0019/Df(3R)6159 mutant nurse cells. Probes label the pericentric heterochromatin of the X (green), 2nd (red), and 3rd (white) chromosomes. Interestingly, we also observed that while wild type cells had heterochromatic sequences that were dispersed to several loci, the pericentric heterochromatin from all chromosomes was always clustered toward one side of the nucleus in Cap-H2Z3-0019/Df(3R)Exel6159 nurse cells (Figure 2B–2E). Measurement of the distances between these probe centroids confirmed that heterochromatic sequences from different chromosomes remained in close proximity in a Cap-H2Z3-0019/Df(3R)Exel6159 background while these sequences were well separated in wild type nurse cells (Figure 2B–2E, Figure 3). This trend was clear throughout development and was also observed in other allelic combinations of Condensin II subunits indicating that Cap-H2 was functioning in the context of the Condensin II complex (Figure 3).
Fig. 3. Condensin II promotes the dispersal of heterologous centromeres. 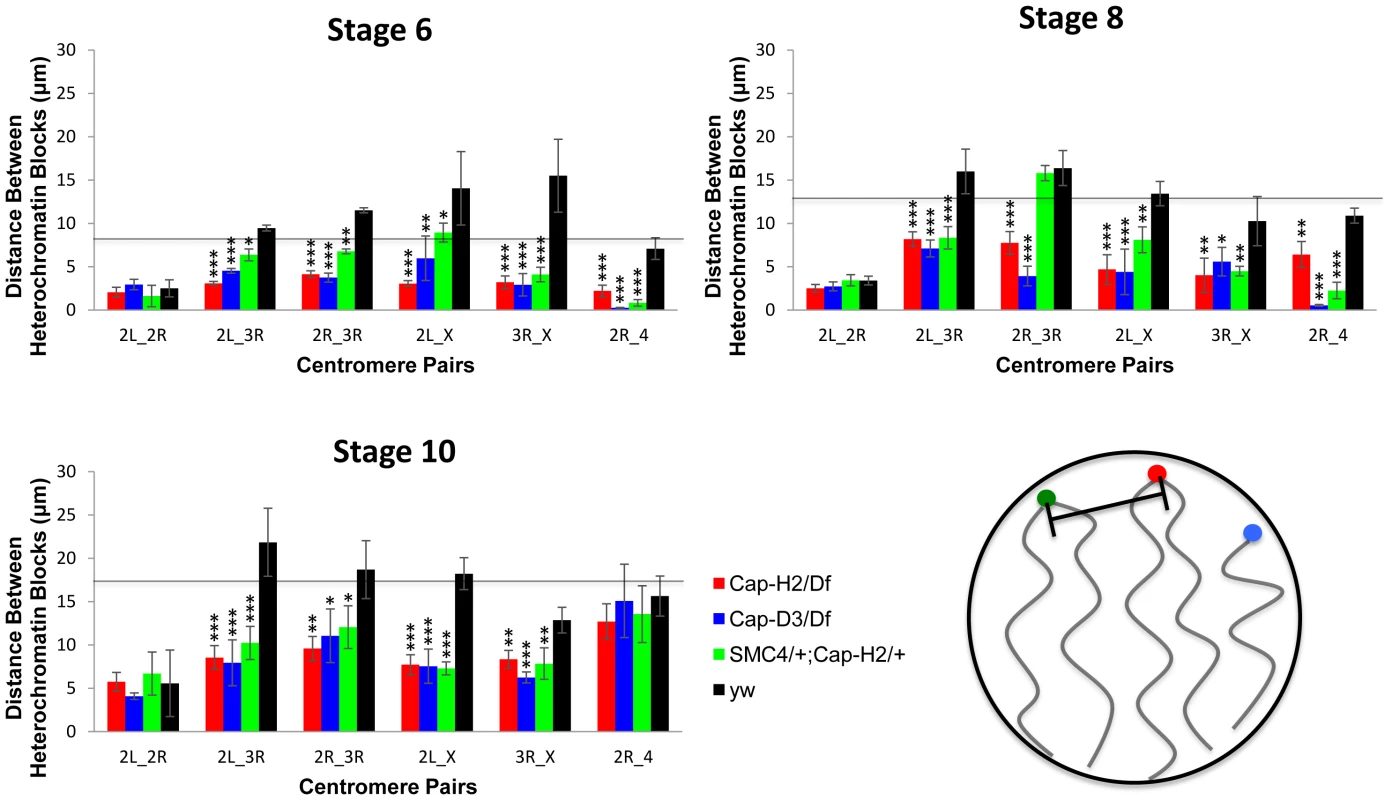
The mean distances between the peri-centric microsatellite sequences of the X, 2R, 2L, 3R and 4 chromosome arms at various stages of nurse cell development are shown. The labels below each set of four bars indicate which pair of loci are being compared. The genotypes assayed were Cap-H2Z3-0019/Df(3R)Exel6159, y[1] w[67c23]; P(w[+mC] = lacW)glu[k08819]/+; Cap-H2Z3-0019/+, Cap-D3EY00456/Df(2R)Exel7023, and wild type (y[1] w[67c23]). Error bars correspond to standard errors. One asterisk signifies p<.05, two asterisks signifies p<.01, and three asterisks signifies p<.001. The cartoon in the lower right panel illustrates the distance measurements being made between different heterochromatin FISH signals. Based on these data, we concluded that wild type nurse cell chromosomes exhibit dispersed peri-centric heterochromatin and centromeric regions at several distinct loci near the periphery of the nucleus and this organizational pattern is dependent upon Condensin II. By contrast, in Condensin II mutant nurse cells, all of the heterochromatic sequences and centromeres are clustered together at a single pole, consistent with the Rabl conformation. We lacked the resolution to determine whether the Rabl conformation is present prior to stage 4 so it was not possible to determine if Condensin II disrupts the Rabl configuration or simply prevents it from developing. However, if the Rabl configuration is a vestige of anaphase, then it would be present after the final cell division and require disruption.
Condensin II is required for axial compaction of nurse cell chromosomes
Since Condensins have been implicated in the axial compaction of chromosomes during mitosis and meiosis [29]–[31], we hypothesized that a similar compaction activity was contributing to the formation of nurse cell CTs. To test this, we measured the distances between pairs of locus centroids on the X chromosome in nurse cells with varying degrees of Condensin II activity. We observed that the mean distance between any two X chromosome loci were substantially increased in Cap-H2Z3-0019/Df(3R)Exel6159 nurse cells compared to their wild type counterparts. This trend was consistent and statistically significant for every pair of loci, and every developmental stage, that we analyzed (Figure 4, p<.01). Measurement of the distances between two pairs of probes on the 2nd and 3rd chromosomes showed that these chromosomes were also more compact in the presence of wild-type Condensin II relative to condenisn II mutants (Figure S3, p<0.01). This trend was also evident in double heterozygous SMC408819/+; Cap-H2Z3-0019/+ flies and was consistent when compared to two additional wild type strains, Oregon-R and Canton-S (Figure S3).
Fig. 4. Condensin II promotes chromosome axial compaction. 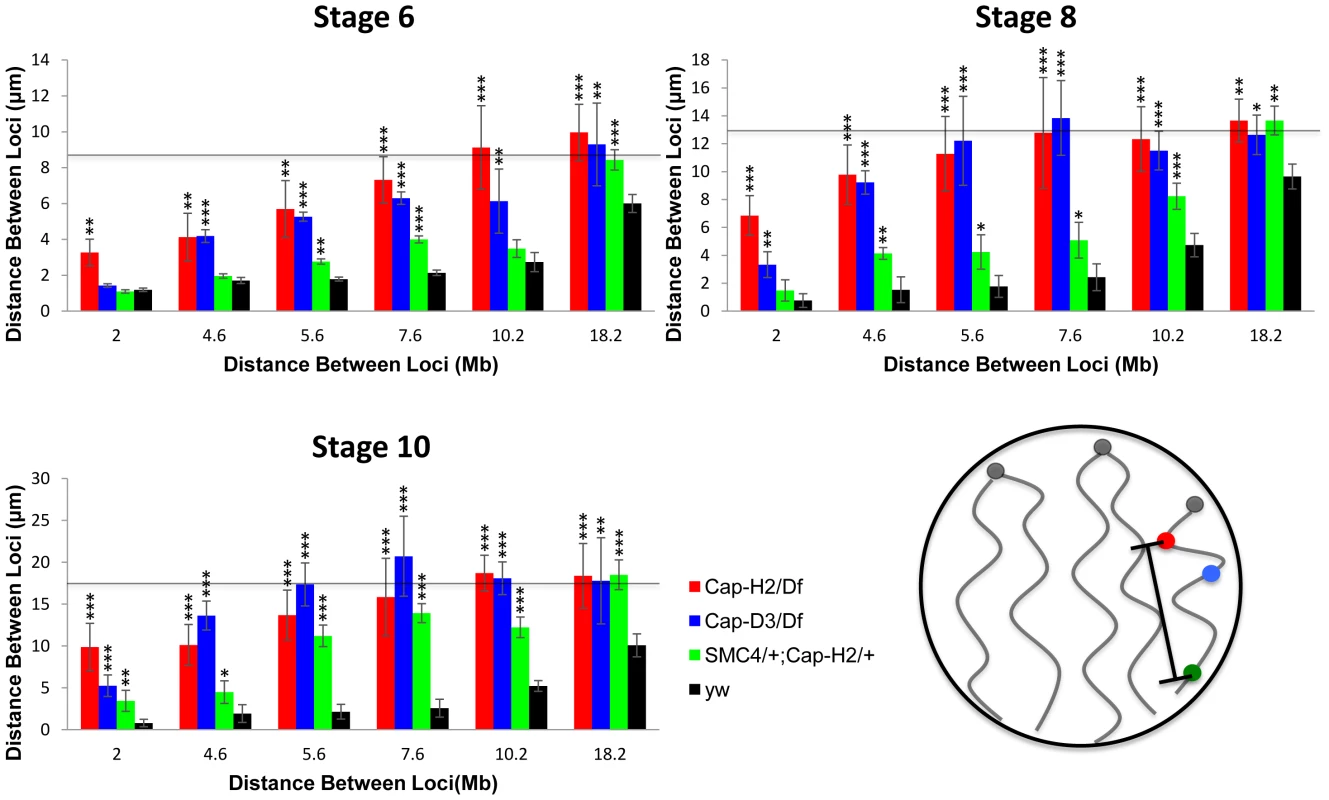
For developmental stage 6, 8, and 10, distances were measured between pairs of loci on the X chromosome. The X-axis indicates the distance between the loci in megabases and the Y-axis indicates their mean spatial separation in microns. The horizontal line indicates the average nuclear radius which is also a good approximation for the mean distance between two random points within a sphere. The following genotypes were assayed: Cap-H2Z3-0019/Df(3R)Exel6159, Cap-D3EY00456/Df(2R)Exel7023, y[1] w[67c23]; P(w[+mC] = lacW)glu[k08819]/+; Cap-H2Z3-0019/+, wild type (y[1] w[67c23]). Bars represent standard errors. One asterisk signifies p<.05, two asterisks signifies p<.01, and three asterisks signifies p<.001. The cartoon in the lower right panel illustrates the axial length measurements being made between different loci. It is interesting to note that the different levels of compaction in different Condensin II allelic combinations correlates well with their respective severity in polytene unpairing phenotypes (Figure 4, Figures S4 and S5). These data indicated that Condensin II functions to promote axial compaction in nurse cell chromosomes and that this compaction activity is concomitant with its polytene unpairing activity.
We next sought to estimate the magnitude of chromosome axial compaction induced by Condensin II. Since the distance between two loci is strictly limited by the diameter of the nucleus, which is much smaller than the length of a nurse cell polytene chromosome, the most accurate estimates of actual chromosome length come from pairs of probes that delimit short chromosomal segments. At stage 4, two loci on the X-chromosome spanning 2 Mb were separated by an average distance of 1.18±0.12 µm in wild type nurse cells. In later developmental stages, the mean distance between these probes remained fairly constant despite a dramatic increase in the size of the nucleus. By stage 10, the distance between these probes actually appeared to have been reduced to 0.80±0.44 µm, though this difference was not statistically significant (Figure 4). In Cap-H2Z3-0019/Df(3R)Exel6159 nurse cells, chromosome length consistently increased throughout development. At stage 4 the same 2 loci were found to be 2.17±0.22 µm apart, on average. By stage 10, this distance had increased to 9.88±2.82 µm (Figure 4) indicating that Condensin II was responsible for at least a 10-fold reduction of chromosome length. In a few of the Cap-H2Z3-0019/Df(3R)Exel6159 nuclei, it was possible to trace the path of the polytene chromosome between the two probes. Direct measurement of the curved path determined its length to be 12.87±0.46 µm (n = 3) (Figure S6). Based on this, we estimate that Condensin II can induce a 12 - to 16-fold reduction in nurse cell chromosome length.
Extrapolation from this estimate of compaction would mean that the entire X chromosome may be as short as 10 µm in wild type stage 10 nurse cells. This is only twice the length of the mitotic X chromosome, which is approximately 5 µm long in diploid cells (data not shown). This suggested that Condensin II was inducing compaction of interphase nurse cell chromosomes, just as it has been proposed to do during mitosis. However, it remained a formal possibility that Condensin II was indirectly contributing to chromosomal compaction or functioning to maintain chromosome length rather than reduce it, per se.
Cap-H2 overexpression in salivary glands is sufficient to induce axial compaction and CT formation
To further assess the ability of Condensin II to compact interphase chromosomes, we ectopically expressed Cap-H2 in the larval salivary glands. Larvae carrying multiple, heat shock inducible, Cap-H2 transgenes exhibit completely unpaired salivary gland chromosomes after a 24 hour heat shock [28]. Since this phenotype resembled the dispersal observed in wild type nurse cell chromosomes, we suspected that it was also associated with chromosome axial compaction, just as in nurse cells. In flies carrying a single Cap-H2 transgene, we found that polytene unpairing was limited while chromosome length was substantially reduced compared to wild type flies, even without heat shock (Figure 5A–5B). This is presumably due to weak expression of Cap-H2 at 25°C. To quantify this phenotype, we crossed in chromosomes that contained insertions of LacO arrays at cytological bands 50F (near the center of chromosome arm 2R) and 60F (near the telomere of 2R) as well as a LacI-GFP fusion protein that was also heat shock inducible. The result was a fly line that displayed two fluorescent chromosomal loci and expressed Cap-H2 in response to heat shock.
Fig. 5. Expression of Cap-H2 in salivary glands induces axial shortening of chromosomes. 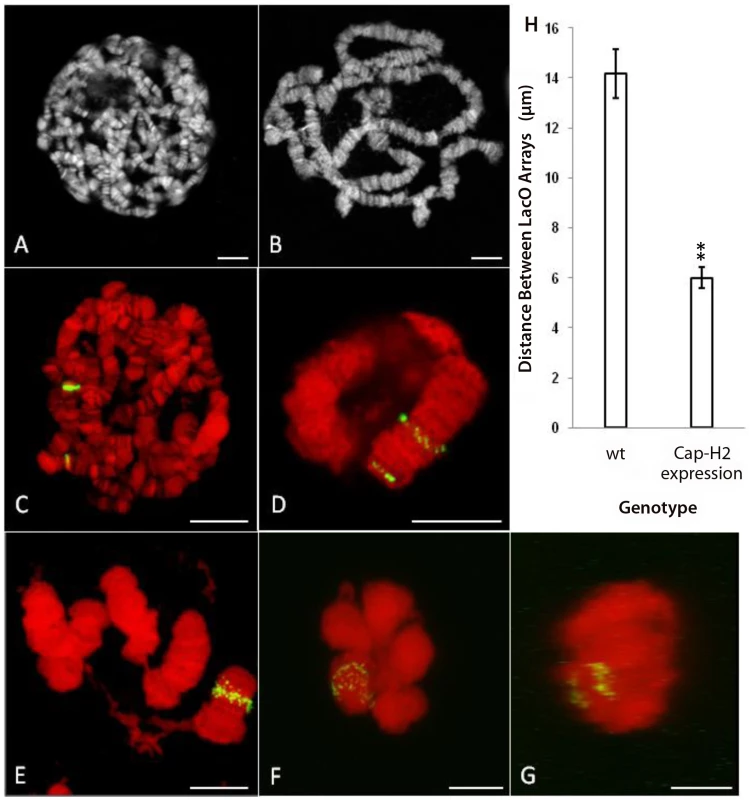
A) DAPI stained, unspread salivary gland chromosome squash from wild type larva raised at 25°C. B) DAPI stained, unspread salivary gland squash from hsp70>Gal4/UAS>Cap-H2 raised at 25°C. C) Projection of salivary gland nucleus from Hs83>LacI-GFP/+; 50F-LacO/60F-LacO with 1 hr, 37°C heat shock. DAPI is shown in red and GFP signal is shown in green. D) Projection of salivary gland nucleus from Hs83>LacI-GFP/+; 50F-LacO/60F-LacO; hsp70>Gal4/UAS>Cap-H2 with 1 hr, 37°C heat shock. E) A unique example (same treatment as in D) in which the nuclear envelope has ruptured revealing details of chromosome structure. F) Projection of salivary gland nucleus from Hs83>LacI-GFP/+; 50F-LacO/60F-LacO; hsp70>GAL4/UAS>Cap-H2 with 1 hr, 37°C heat shock. Both LacO arrays remain confined within a common territory during the dispersal process. G) A projection of the same nucleus in F through the Y-axis showing the separation between the two LacO arrays. H) Measurements of the distance between LacO insertions in cases similar to C and D. All scale bars correspond to 10 µm. In a control line that contained the LacI-GFP transgene and the two LacO arrays without a Cap-H2 transgene, a 1-hour heat shock resulted in polytene salivary gland chromosomes with two fluorescent bands that were separated by a mean distance of 14.2±1.0 µm (Figure 5C, 5H). With the addition of the Cap-H2 transgene and identical treatment, most GFP bands started to become unpaired but the two LacO arrays typically remained spatially distinct from each other (Figure 5D). In cases such as these, the mean distance between the LacO arrays was reduced to 6.0±0.4 µm (Figure 5H). Extrapolation from this figure suggested that the entire right arm of chromosome 2 was close to 12 µm long after expression of Cap-H2. In some instances, entire salivary gland chromosome arms could be resolved and measured, supporting this estimate (Figure 5E). A previous study determined that the entire right arm of chromosome 2 was 144.3±7.8 µm long in wild type salivary glands [21]. Based on this, we concluded that Condensin II is capable of inducing at least 12-fold compaction of salivary gland chromosomes.
Taken together, our observations in nurse cells and salivary glands indicated that Condensin II can induce a tremendous amount of axial compaction in interphase chromosomes. Moreover, this compaction coincides with the formation of compact CTs in both cell types (Figure 1B, Figure 5F–5G). Prior to the dispersal of salivary gland polytene chromosomes, each chromosome arm became compacted into a discrete nuclear compartment, similar to the 5-blob stage in nurse cells (Figure 1B, Figure 5F–5G). Even after substantial dispersal had occurred, GFP signal from both LacO insertion sites remained confined within a fraction of the nuclear volume (Figure 5F–5G). The finding that Condensin II can produce such remarkably similar phenotypes in two unrelated cell types suggests that this complex is likely to regulate chromosome length and CT structure in other contexts as well. This idea is further supported by the observation that meiotic CT formation in non-polyploid Drosophila primary spermatocytes is Condensin II dependent [26].
Discussion
We have shown that in the polyploid nurse cells of Drosophila, Condensin II activity is required to unpair polytene chromosomes and disperse heterochromatic sequences. Interestingly, while dispersal of heterologous peri-centric heterochromatin blocks on different chromosomes was dependent on Condensin II activity, homologous (or allelic) heterochromatic sequences tended to be paired and localized to the nuclear periphery in both wild-type and Condensin II mutant flies (Figure 2, Figure 3, Figures S2 and S4). Although Condensin II promotes the unpairing of euchromatic homologous sequences, unpaired homologs and sister chromatids were sequestered into discrete territories within the nucleus (Figure 1, Figure S5D). We also observed that axial compaction of chromosomes in nurse cells required Condensin II activity (Figure 4, Figure S3) and overexpression of Cap-H2 in salivary gland cells could induce shortening of chromosome arms (Figure 5). These observations are consistent with previous work showing that Condensin II serves to individualize homologous chromosomes by disrupting pairing interactions in polytene, diploid, and meiotic cells [26], [28]. In addition, a recent whole genome RNAi screen in cultured Drosophila cells also identified Cap-H2, Cap-D3 and SMC2 as homolog pairing disruptors, further supporting an anti-pairing activity for Condensin II [32].
If the activity of Condensin II spatially separates homologous chromosomes from each other, how does it simultaneously keep all homologous chromosomes of a polyploid nucleus confined within a common territory? One likely reason is that unpairing of homologs and sister chromatids is incomplete in polyploid cells. When we measured the unpairing of several loci in nurse cells, we found that pairing was disrupted at all euchromatic loci we examined, but heterochromatic regions near the telomere and centromere retained some associations with their allelic sequences (Figures S4, S7). We speculate that this incomplete dispersal of nurse cell polytene chromosomes is due, in part, to under-replicated heterochromatic regions associated with telomeres and centromeres [23], [33]–[34]. In addition to under-replicated regions, it is also possible that nurse cell pairing of homologous peri-centric heterochromatin is actively maintained and insensitive to Condensin II activities, while clustering of heterologous peri-centric heterochromatin is inhibited by Condensin II activities.
Previous studies have found that centromeric and telomeric sequences tend to localize to the nuclear periphery in a number of Drosophila cell types as well as in other organisms [20]–[21], [35]. We found that the same is true in nurse cells (Figure 2). Interestingly, in wild type nurse cells, all loci on the X chromosome tended to move toward the nuclear periphery with developmental progression (Figure 2A). In the polytene nurse cell chromosomes of Cap-H2Z3-0019/Df(3R)Exel6159 mutants, heterochromatic sequences remained near the periphery but euchromatic loci were always located more centrally (Figure 2A). If heterochromatic loci are physically tethered to the nuclear envelope, then one would expect chromosomal compaction to pull linked, euchromatic loci toward the nuclear periphery as well. Centromeres and telomeres on the same chromosome would likely move toward each other by sliding along the nuclear envelope (Figure 6).
Fig. 6. Model for Condensin II–mediated chromosome territory formation. 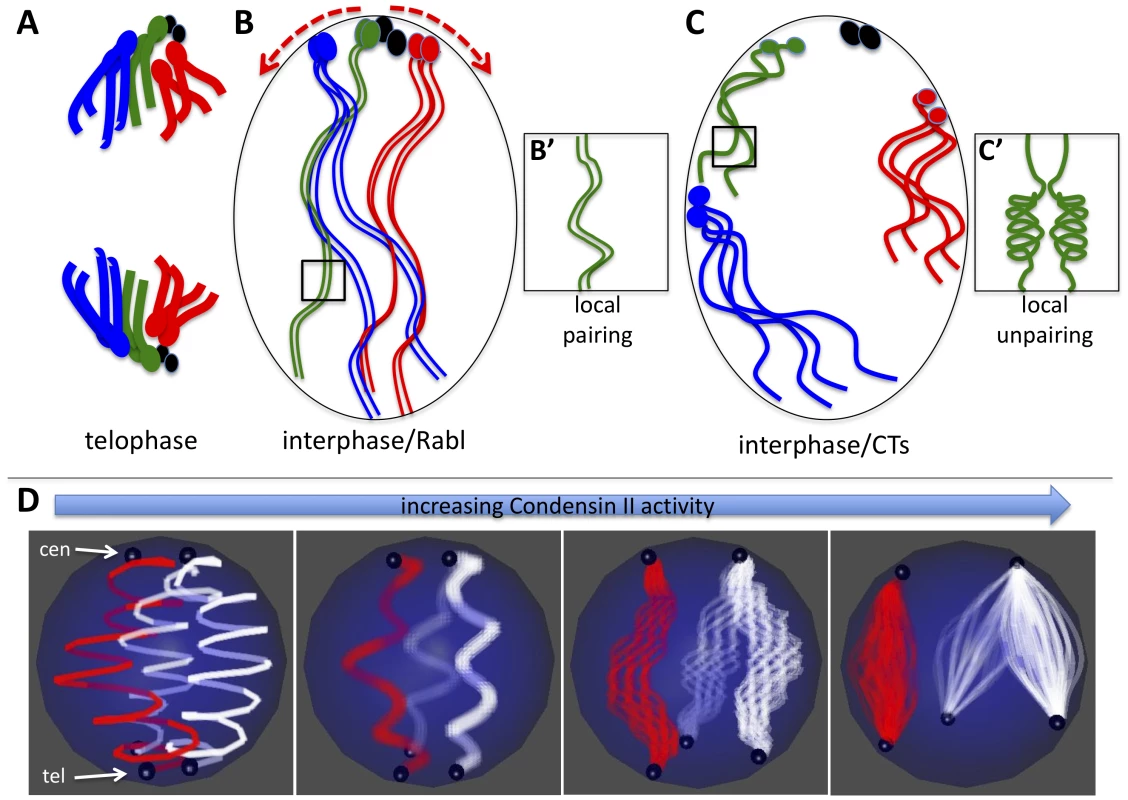
A) Condensed mitotic chromosomes are pulled to opposite poles and centromeres cluster at the poles in telophase. Four chromosomes are shown where the ovals represent centromeres and/or peri-centric heterochromatin. The small fourth chromosomes are shown in black. B) In interphase, decondensed chromosomes adopt the Rabl configuration with clustered centromeres simply because of their previous organization in telophase. B') Long and relaxed interphase homologous chromosomes are paired. C) Activation of Condensin II in interphase nuclei has two direct consequences: First it induces axial compaction of the chromosome arms. Second, in order to accommodate the shorting distance between telomeres and centromeres, heterochromatin slides along the nuclear periphery as the axial length of the arms decrease (see red arrows in B). Because heterochromatin proteins interact with the nuclear periphery, we speculate that as euchromatic regions of chromosome arms shorten they are drawn to the nuclear periphery. Heterochromatin and other chromatin interactions with the nuclear periphery combined with the Condensin mediated shortening of chromosome arms forces chromosomes into discrete territories. C') Chromosome arm compaction is most likely accompanied by chromatin folding and coiling that occludes homologous sequences from interacting in trans. D) The model in A–C depicts a diploid cells scenario, but the same mechanical principles can apply to a polyploid nucleus that forms CTs. In the panel on the left, two polytene chromosomes (the X chromosome in red and an autosome in white) are shown where centromeres (cen) are clustered at the North pole and telomeres (tel) are clustered at the South pole of the nucleus. As Condensin II activity increases (left to right) the chromatin fibers within polytenes begin to unpair as the chromosome arms shorten. Note that although euchromatic sequences unpair, homologous heterochromatin blocks remain paired and move in concert. By contrast, heterologous heterochromatin blocks disperse to different parts of the nuclear periphery. As above (B–C), Condensin II mediated compaction drives globular CT formation at the nuclear periphery as a consequence of chromatin interactions with nuclear envelope proteins. We propose a general model for the effects of Condensin II activity on interphase chromosome structure and the spatial organization of chromosomes within the nucleus: Interphase chromosomes may first adopt a Rabl configuration upon exit from mitotic telophase, simply because all centromeres are pulled to the spindle pole body and thus are clustered (Figure 6A–6B). In interphase cells, Condensin II promotes a dramatic axial compaction of chromosomes (Figure 4, Figure 5) and this compaction is likely associated with topological changes in DNA. This Condensin II mediated compaction of chromatin (Figure 6C) sequesters DNA into discrete chromosomal structures, as proposed for mitotic chromosome condensation [36].
Sequestering of sequences into intrachromosomal folds disfavors interchromosomal interactions between homologous loci (Figure 6C'), which disrupts polytene chromosome structure and trans-sensitive processes such as transvection [28]. Pairing between allelic, heterochromatic loci in polyploid nurse cells is unaffected by Condensin II and these sequences localize to the nuclear periphery in nurse cells (Figure 2, Figure S7). The combination of Condensin II mediated chromosome compaction and heterochromatic interactions with the nuclear periphery drive each chromosome into a globular CT that is associated with the nuclear envelope (Figure 6D). Because homologous heterochromatic sequences in polyploid nurse cells are under-replicated we speculate that this keeps the unpaired chromatids confined together within the CT. Thus, we propose that Condensin II serves to spatially separate individual chromosomes by inhibiting homology dependent pairing interactions as well as interactions between heterologous chromosomes. Many aspects of this model may be applicable to the roles of Condensins in the nuclear reorganization that occurs during mitosis or meiosis, since it seems to contribute similarly to the processes of chromosome compaction and individualization in all of these contexts [26], [28].
The relationship between chromosome individualization and compaction is currently unclear. Since Condensin II always appears to promote both processes in nurse cells and salivary gland cells, it may be that they both stem from a common enzymatic activity. It is tempting to propose that compaction drives individualization since the former is always detectable first. However, FISH does not provide sufficient resolution to distinguish between DNA sequences that are physically interacting and those that are no longer interacting at the molecular level but remain in close proximity simply because they were paired in the recent past. Perhaps DNA-DNA interactions in cis and in trans compete with each other, as proposed by Wu and colleagues [37]–[38]. In this paradigm, Condensin II could function to shift the equilibrium away from trans and toward cis interactions. This model would not only explain the temporal coincidence of compaction and polytene dispersal, but also the curious observation that polytene chromosome length increases with ploidy (Figure 4) [21]. In the latter case, homologous chromosomes interact with each other in trans (i.e. pair) and thus occlude intrachromosomal interactions that would otherwise restrict chromosome length. More copies of homologous sequences would shift the equilibrium in favor of more pairing interactions, thus further increasing chromosome length.
With regards to the generality of the proposed model, it is important to consider the cell types that it is based on. While nurse cells are germline derived and salivary glands are somatic, both tissues are polyploid. Though polyploidy is the rule rather than the exception in terminally differentiated Drosophila tissues, it may be that certain aspects of this model do not apply to diploid cells. We have not yet been able to compare chromosome axial lengths in wild-type and Condensin II mutant diploid cells, but efforts are underway to achieve this. The strongest evidence for the generality of our model comes from observations in spermatogenesis. During prophase I, wild-type primary spermatocytes form very well separated CTs, each of which is associated with the nuclear envelope. However, this organization is completely absent in Condensin II mutants which appear to contain a chromocenter-like structure of dense DAPI signal [26]. The phenotypes in this diploid cell type are strikingly similar to our observations in nurse cells suggesting that similar mechanisms may regulate genomic architecture regardless of ploidy.
Though the cytological effects of Condensin II on chromosome structure are clear, the molecular mechanisms that produce these changes remain enigmatic. In vitro evidence suggests that it mostly likely involves changes in DNA topology such as DNA supercoiling or looping [25]. Given that Condensin I and Condensin II share a common set of SMC subunits, it can be difficult to tease apart their effects. Our results agree well with recent reports that Condensin II is an important regulator of mitotic chromosome axial compaction while condensin I controls lateral compaction [39]–[40]. It seems likely that the phenotypes displayed after overexpression of Cap-H2 in salivary glands are the result of shifting an equilibrium toward Condensin II at the expense of Condensin I since Cap-H2 is specific to Condensin II.
Regardless, we believe this is the first report of any gene that directly controls interphase CT structure in any system. It will be of great interest to determine if Condensins function to regulate the formation or maintenance of CTs in other eukaryotes. A recent report described structural defects in the interphase nuclei of mouse embryonic stem cells when Condensin subunits were knocked down by RNAi [27]. Perhaps these cells exhibit disorganized CT structure or increased chromosome length.
Though it is clear that the spatial positions of genes, within a nucleus, can influence their expression patterns [41]–[42], it is not well understood how CTs contribute to gene regulation. We have now described two reciprocal systems in which we can prevent or induce Condensin II mediated CT formation and observe the effects. Future studies will be directed at elucidating the causal relationships between nuclear organization and transcriptional activity.
Considering the effects that Condensin II has on large scale chromosome conformations and nuclear organization, perhaps it would promote long distance interactions between sequences on the same chromosome while precluding interactions between elements on different chromosomes. This is supported by the finding that transvection is enhanced in Condensin II mutants and suppressed when Cap-H2 is overexpressed [28]. It is known that the transcriptional profiles of many genes correlate with those of their chromosomal neighbors [43]. We speculate that Condensin II may contribute to the mechanisms of co-regulation in gene neighborhoods as it is likely to promote the spatial co-localization of genetically linked genes and/or long-distance enhancer-promoter cis interactions.
Materials and Methods
Fly strains
The following fly strains were used in this study: y[1] w[1118] (wild type control for all nurse cell studies, unless otherwise noted), y[1] w[67c23]; P(w[+mC] y[+mDint2] = EPgy2)Cap-D3[EY00456]/Df (2L)Exel7023 (Cap-D3 mutant), y[1] w[67c23]; P(w[+mC] = lacW)glu[k08819]/+; Cap-H2Z3-0019/+ (SMC4, Cap-H2 double heterozygote), Cap-H2Z3-0019/Df(3R)Exel6159 (Cap-H2 mutant), Oregon-R-S (wild type control for all salivary gland studies), w[*]; P(w[+mC] = GAL4-nos.NGT)40 P(w[+mC] = lacO.256x)43 P(lacO.256x)50F P(lacO.256x)57A P(lacO.256x)60AB/CyO; P(w[+mC] = UAS-GFP.lacI)1.2/TM3, Sb[1] (lacO array 50F insertion line), w[*]; P(w[+mC] = lacO.256x)60F (lacO array 60F insertion line). Canton-S and Oregon-R (OrR) wild type controls were also used in Figure S2. All flies were maintained at 25°C on standard cornmeal molasses media.
Labeling of probes
BAC clones were selected to span the X chromosome at regular intervals. The following BACs were ordered from CHORI BACPAC Resources: BACR25p24, BACR18c23, BACR23m08, BACR32h11, BACR33k15, BACR32l12. These map approximately to 3.9, 8.3, 12.9, 18.5, 20.5, and 22.1 Mb, respectively. To mark the heterochromatic region of the X chromosome we synthesized PCR products to the 359 bp repeat sequence using the primers 5′-CGGTCATCAAATAATCATTTATTTTGC-3′ and 5′-CGAAATTTGGAAAAACAGACTCTGC-3′. We also used the following oligonucleotides to label the 2L, 2R, 3R, and 4 heterochromatic regions, respectively: (AATAG)5, (AACAC)5, 5′-CCCGTACTGGTCCCGTACTGGTCCCGTACTCGGTCCCGTACTCGGT-3′, and (AATAT)5. Telomeric probes were made by PCR amplification of plasmids containing a 2 kb ApaI–ApaI fragment of the 3′ UTR of HeT-A and the 23Zn-1 fragment containing ORF1+ORF2 of HeT-A.
Midipreps were performed on bacterial cultures to isolate BAC DNA. In some cases, probes were made by whole genome amplification (Sigma WGA kit) of previously isolated BAC DNA. 20 µg of BAC DNA or amplified DNA was digested overnight with the restriction enzymes AluI, HaeIII, MseI, MspI, and RsaI. DNA fragments were purified using a Qiagen PCR cleanup column, ethanol precipitated, and end labeled with aminoallyl dUTP (Invitrogen ARES Alexa Fluor DNA Labeling Kit) using terminal deoxytransferase as previously described [28]. DNA fragments were purified again on a PCR cleanup column (Qiagen) and ethanol precipitated. These labeled DNA fragments were then conjugated to Alexa488, Alexa546, or Alexa647 dyes as described in the ARES DNA labeling kits. Labeled probes were purified on a PCR cleanup column, ethanol precipitated, and resuspended in 20 µL hybridization buffer.
Fluorescence in situ hybridization and imaging
FISH protocol was adapted primarily from Dernburg et al., 2000. 0–2 day old female flies were fattened on yeast with males at 25°C for 2 days. 5 ovary pairs were dissected in Grace's medium and fixed within 15 minutes. Fixation was performed for 4 minutes in 100 mM Sodium cacodylate, 100 mM sucrose, 40 mM Sodium acetate, 10 mM EGTA with 3.7% formaldehyde. Ovaries were washed twice for 5 minutes each in 2X SSCT and individual ovarioles were separated in 2X SSCT. Ovarioles were washed for 10 minutes each in 2X SSCT with 20%, 40%, and 50% formamide at room temperature and once more for 2 hours in 2X SSCT with 50% formamide at 30°C (for AT rich heterochromatic probes) or 37°C. 200–500 ng of probe was mixed into a total of 40 µL hybridization buffer, denatured at 95°C, and snap frozen in liquid nitrogen. Formamide mixture was aspirated from ovarioles and replaced with probe/hybridization buffer mixture. This suspension was mixed gently and incubated at 30 or 37C for 10 minutes prior to a 2 min denaturation at 92°C. Tubes were immediately returned to hybridization oven and left overnight at the specified temperature.
The next day, ovarioles were washed 4 times for 30 minutes each in 2X SSCT with 50% formamide at the hybridization temperature. Ovarioles were then washed 10 minutes each in 2X SSCT with 40% and 20% formamide. Three 5 minute washes with 2X SSCT were performed to remove the remaining formamide. The ovarioles were then stained with 10 ng/mL DAPI in 2X SSCT for 10 minutes and washed twice with 2X SSCT for 10 minutes each. Ovarioles were mounted in vectashield. Pieces of number 1½ coverslips were used as spacers between the actual coverslip and slide to prevent flattening of the egg chambers. Slides were imaged on a Zeiss 510 meta confocal microscope using a 1.4 NA, 63× objective.
Nurse cell image analysis
Individual channels were separated from 3D image stacks and analyzed using the 3D Object Counter plugin for imageJ. The spatial coordinate and intensity measurements were then analyzed in the R programming environment. Intensity thresholds were set as follows: 25 for DAPI, 35 for Alexa488 and Alexa647, 55 for Alexa546. Objects smaller than 10 voxels were ignored. Spot counts for each probe are simply the number of objects detected. For each probe in each nucleus, the centroid was defined as the center of total fluorescence intensity. Principal component analysis was performed using point weights equal to the total fluorescence intensity of each object. The center of fluorescence intensity in the DAPI channel was set as the nuclear center and radial distances for each object were measured in relation to this point.
Salivary gland imaging and measurements
Transgenic lines carrying a 256-repeat array of the Lac-O sequence at chromosomal position 50F were crossed to lines carrying the same Lac-O array inserted at position 60F with transgenes Hsp70>Gal4 and UAS>Cap-H2, as previously described [28]. These lines also had the ability to express a GFP-LacI fusion protein that binds to the LacO arrays and marks the chromosomal insertion site of the LacO array. Expression of GFP-LacI and Cap-H2 was controlled with heatshock at 37°C, as detailed in the text and figure legends. GFP-LacI in salivary glands was imaged as previously described [28]. 3D distances between the two Lac-O arrays were measured as described for nurse cell images.
Supporting Information
Zdroje
1. CremerT, CremerM, DietzelS, MullerS, SoloveiI, et al. (2006) Chromosome territories–a functional nuclear landscape. Curr Opin Cell Biol 18 : 307–16.
2. GotoB, OkazakiK, NiwaO (2001) Cytoplasmic microtubular system implicated in de novo formation of a Rabl-like orientation of chromosomes in fission yeast. J Cell Sci 114 : 2427–35.
3. BystrickyK, LarocheT, van HouweG, BlaszczykM, GasserSM (2005) Chromosome looping in yeast: telomere pairing and coordinated movement reflect anchoring efficiency and territorial organization. J Cell Biol 168 : 375–87.
4. PecinkaA, SchubertV, MeisterA, KrethG, KlatteM, et al. (2004) Chromosome territory arrangement and homologous pairing in nuclei of Arabidopsis thaliana are predominantly random except for NOR-bearing chromosomes. Chromosoma 113 : 258–69.
5. TanabeH, MullerS, NeusserM, von HaseJ, CalcagnoE, et al. (2002) Evolutionary conservation of chromosome territory arrangements in cell nuclei from higher primates. Proc Natl Acad Sci U S A 99 : 4424–9.
6. RajapakseI, PerlmanMD, ScalzoD, KooperbergC, GroudineM, et al. (2009) The emergence of lineage-specific chromosomal topologies from coordinate gene regulation. Proc Natl Acad Sci USA 106 : 6679–84.
7. Lieberman-AidenE, van BerkumNL, WilliamsL, ImakaevM, RagoczyT, et al. (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326 : 289–93.
8. SextonT, YaffeE, KenigsbergE, BantigniesF, LeblancB, et al. (2012) Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148 : 458–72.
9. CremerT, CremerM (2010) Chromosome territories. Cold Spring Harb Perspect Biol 2: a003889.
10. BártováE, KozubekS (2006) Nuclear architecture in the light of gene expression and cell differentiation studies. Biol Cell 98 : 323–36.
11. MisteliT (2005) Concepts in nuclear architecture. Bioessays 27 : 477–87.
12. ZinkD, CermerT, SaffrichR, FischerR, TrendelenburgMF, et al. (1998) Structure and dynamics of human interphase chromosome territories in vivo. Hum Genet 102 : 241–51.
13. StadlerS, SchnappV, MayerR, SteinS, CermerC, et al. (2004) The architecture of chicken chromosome territories changes during differentiation. BMC Cell Biol 5 : 44.
14. MisteliT (2007) Beyond the sequence: cellular organization of genome function. Cell 128 : 787–800.
15. van DrielR, FranszP (2004) Nuclear architecture and genome functioning in plants and animals: what can we learn from both? Exp Cell Res 296 : 86–90.
16. ShawPJ, AbranchesR, Paula SantosA, BevenAF, StogerE, et al. (2002) The architecture of interphase chromosomes and nucleolar transcription sites in plants. J Struct Biol 140 : 31–8.
17. RablC (1885) Uber Zelltheilung. Morphol. Jahrbuch 10 : 214–330.
18. CremerT, CremerC, BaumannH, LuedtkeEK, SperlingK, et al. (1982) Rabl's model of the interphase chromosome arrangement tested in Chinese hamster cells by premature chromosome condensation and laser-UV-microbeam experiments. Hum Genet 60 : 46–56.
19. HiraokaY, DernburgAF, ParmeleeSJ, RykowskiMC, et al. (1993) The onset of homologous chromosome pairing during Drosophila melanogaster embryogenesis. J Cell Biol 120 : 591–600.
20. MarshallWF, DernburgAF, HarmonB, AgardDA, SedatJW (1996) Specific interactions of chromatin with the nuclear envelope: positional determination within the nucleus in Drosophila melanogaster. Mol Biol Cell 7 : 825–42.
21. HochstrasserM, SedatJW (1987) Three-dimensional organization of Drosophila melanogaster interphase nuclei. I. Tissue-specific aspects of polytene nuclear architecture. J Cell Biol 104 : 1455–70.
22. VazquezJ, BelmontAS, SedatJW (2002) The dynamics of homologous chromosome pairing during male Drosophila meiosis. Curr. Biol 12 : 1473–83.
23. DejKJ, SpradlingAC (1999) The endocycle controls nurse cell polytene chromosome structure during Drosophila oogenesis. Development 126 : 293–303.
24. KimuraK, HiranoT (1997) ATP-dependent positive supercoiling of DNA by 13S Condensin: a biochemical implication for chromosome condensation. Cell 90 : 625–34.
25. HiranoT (2005) Condensins: organizing and segregating the genome. Curr Biol 15 : 265–75.
26. HartlTA, SweeneySJ, KneplerPJ, BoscoG (2008) Condensin II resolves chromosomal associations to enable anaphase I segregation in Drosophila male meiosis. PLoS Genet 4: e1000228 doi:10.1371/journal.pgen.1000228.
27. FazzioTG, PanningB (2010) Condensin complexes regulate mitotic progression and interphase chromatin structure in embryonic stem cells. J Cell Biol 188 : 491–503.
28. HartlTA, SmithHF, BoscoG (2008) Chromosome alignment and transvection are antagonized by Condensin II. Science 322 : 1384–7.
29. HirotaT, GerlichD, KochB, EllenbergJ, PetersJM (2004) Distinct functions of Condensin I and II in mitotic chromosome assembly. J Cell Sci 117 : 6435–45.
30. YuHG, KoshlandDE (2003) Meiotic Condensin is required for proper chromosome compaction, SC assembly, and resolution of recombination-dependent chromosome linkages. J Cell Biol 163 : 937–47.
31. MetsDG, MeyerBJ (2009) Condensins regulate meiotic DNA break distribution, thus crossover frequency, by controlling chromosome structure. Cell 139 : 73–86.
32. JoyceEF, WilliamsBR, XieT, WuCT (2012) Identification of Genes That Promote or Antagonize Somatic Homolog Pairing Using a High-Throughput FISH-Based Screen. PLoS Genet 8: e1002667 doi:10.1371/journal.pgen.1002667.
33. LillyMA, SpradlingAC (1996) The Drosophila endocycle is controlled by cyclin E and lacks a checkpoint ensuring S-phase completion. Genes Dev 19 : 2514–26.
34. RoyzmanI, Hayashi-HagiharaA, DeiKJ, BoscoG, LeeJT, et al. (2002) The E2F cell cycle regulator is required for Drosophila nurse cell DNA replication and apoptosis. Mech. Dev 199 : 225–37.
35. DernburgAF, BromanKW, FungJC, MarshallWF, PhilipsJ, et al. (1996) Perturbation of nuclear architecture by long-distance chromosome interactions. Cell 85 : 745–59.
36. TumbarT, BelmontAS (2001) Interphase movements of a DNA chromosome region modulated by VP16 transcriptional activator. Nat Cell Biol 3 : 134–9.
37. CavalliG (2007) Chromosome kissing. Curr Opin Genet Dev 17 : 443–50.
38. BoutanaevAM, KalmykovaAI, ShevelyovYY, NurminskyDI (2002) Large clusters of co-expressed genes in the Drosophila genome. Nature 420 : 666–9.
39. HiranoT (2006) At the heart of the chromosome: SMC proteins in action. Nat. Rev. Mol. Cell Bio 5 : 311–22.
40. LeeAM, WuCT (2006) Enhancer-promoter communication at the yellow gene of Drosophila melanogaster: diverse promoters participate in and regulate trans interactions. Genetics 4 : 1867–80.
41. MorrisJR, PetrovDA, LeeAM, WuCT (2004) Enhancer choice in cis and in trans in Drosophila melanogaster: role of the promoter. Genetics 4 : 1739–47.
42. ShintomiK, HiranoT (2011) The relative ratio of condensin I to II determines chromosome shapes. Genes Dev 25(14):1464–9.
43. GreenLC, KalitsisP, ChangTM, CipeticM, KimJH, et al. (2012) Contrasting Roles of condensing I and condensing II in mitotic chromosome formation J Cell Sci. 125(Pt6):1591–604.
Štítky
Genetika Reprodukční medicína
Článek Mutational Signatures of De-Differentiation in Functional Non-Coding Regions of Melanoma GenomesČlánek Rescuing Alu: Recovery of Inserts Shows LINE-1 Preserves Alu Activity through A-Tail ExpansionČlánek Genetics and Regulatory Impact of Alternative Polyadenylation in Human B-Lymphoblastoid CellsČlánek Retrovolution: HIV–Driven Evolution of Cellular Genes and Improvement of Anticancer Drug ActivationČlánek The Mi-2 Chromatin-Remodeling Factor Regulates Higher-Order Chromatin Structure and Cohesin DynamicsČlánek Identification of Human Proteins That Modify Misfolding and Proteotoxicity of Pathogenic Ataxin-1
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2012 Číslo 8
-
Všechny články tohoto čísla
- Mutational Signatures of De-Differentiation in Functional Non-Coding Regions of Melanoma Genomes
- Rescuing Alu: Recovery of Inserts Shows LINE-1 Preserves Alu Activity through A-Tail Expansion
- Genetics and Regulatory Impact of Alternative Polyadenylation in Human B-Lymphoblastoid Cells
- Chromosome Territories Meet a Condensin
- It's All in the Timing: Too Much E2F Is a Bad Thing
- Fine-Mapping and Initial Characterization of QT Interval Loci in African Americans
- Genome Patterns of Selection and Introgression of Haplotypes in Natural Populations of the House Mouse ()
- A Combinatorial Amino Acid Code for RNA Recognition by Pentatricopeptide Repeat Proteins
- Advances in Quantitative Trait Analysis in Yeast
- Experimental Evolution of a Novel Sexually Antagonistic Allele
- Variation of Contributes to Dog Breed Skull Diversity
- , a Gene Involved in Axonal Pathfinding, Is Mutated in Patients with Kallmann Syndrome
- A Single Origin for Nymphalid Butterfly Eyespots Followed by Widespread Loss of Associated Gene Expression
- Cryptocephal, the ATF4, Is a Specific Coactivator for Ecdysone Receptor Isoform B2
- Retrovolution: HIV–Driven Evolution of Cellular Genes and Improvement of Anticancer Drug Activation
- The PARN Deadenylase Targets a Discrete Set of mRNAs for Decay and Regulates Cell Motility in Mouse Myoblasts
- A Sexual Ornament in Chickens Is Affected by Pleiotropic Alleles at and , Selected during Domestication
- Use of Allele-Specific FAIRE to Determine Functional Regulatory Polymorphism Using Large-Scale Genotyping Arrays
- Novel Loci for Metabolic Networks and Multi-Tissue Expression Studies Reveal Genes for Atherosclerosis
- The Genetic Basis of Pollinator Adaptation in a Sexually Deceptive Orchid
- Uncovering the Genome-Wide Transcriptional Responses of the Filamentous Fungus to Lignocellulose Using RNA Sequencing
- Inheritance Beyond Plain Heritability: Variance-Controlling Genes in
- The Metabochip, a Custom Genotyping Array for Genetic Studies of Metabolic, Cardiovascular, and Anthropometric Traits
- Reprogramming to Pluripotency Can Conceal Somatic Cell Chromosomal Instability
- Condensin II Promotes the Formation of Chromosome Territories by Inducing Axial Compaction of Polyploid Interphase Chromosomes
- PTEN Negatively Regulates MAPK Signaling during Vulval Development
- A Dynamic Response Regulator Protein Modulates G-Protein–Dependent Polarity in the Bacterium
- Population Genomics of the Facultatively Mutualistic Bacteria and
- Components of a Fanconi-Like Pathway Control Pso2-Independent DNA Interstrand Crosslink Repair in Yeast
- Polysome Profiling in Liver Identifies Dynamic Regulation of Endoplasmic Reticulum Translatome by Obesity and Fasting
- Stromal Liver Kinase B1 [STK11] Signaling Loss Induces Oviductal Adenomas and Endometrial Cancer by Activating Mammalian Target of Rapamycin Complex 1
- Reprogramming of H3K27me3 Is Critical for Acquisition of Pluripotency from Cultured Tissues
- Transgene Induced Co-Suppression during Vegetative Growth in
- Hox and Sex-Determination Genes Control Segment Elimination through EGFR and Activity
- A Quantitative Comparison of the Similarity between Genes and Geography in Worldwide Human Populations
- Minibrain/Dyrk1a Regulates Food Intake through the Sir2-FOXO-sNPF/NPY Pathway in and Mammals
- Comparative Analysis of Regulatory Elements between and by Genome-Wide Transcription Start Site Profiling
- Simple Methods for Generating and Detecting Locus-Specific Mutations Induced with TALENs in the Zebrafish Genome
- S Phase–Coupled E2f1 Destruction Ensures Homeostasis in Proliferating Tissues
- Cell-Nonautonomous Signaling of FOXO/DAF-16 to the Stem Cells of
- The Mi-2 Chromatin-Remodeling Factor Regulates Higher-Order Chromatin Structure and Cohesin Dynamics
- Comparative Analysis of the Genomes of Two Field Isolates of the Rice Blast Fungus
- Role of Mex67-Mtr2 in the Nuclear Export of 40S Pre-Ribosomes
- Genetic Modulation of Lipid Profiles following Lifestyle Modification or Metformin Treatment: The Diabetes Prevention Program
- HAL-2 Promotes Homologous Pairing during Meiosis by Antagonizing Inhibitory Effects of Synaptonemal Complex Precursors
- SLX-1 Is Required for Maintaining Genomic Integrity and Promoting Meiotic Noncrossovers in the Germline
- Phylogenetic and Transcriptomic Analysis of Chemosensory Receptors in a Pair of Divergent Ant Species Reveals Sex-Specific Signatures of Odor Coding
- Reduced Prostasin (CAP1/PRSS8) Activity Eliminates HAI-1 and HAI-2 Deficiency–Associated Developmental Defects by Preventing Matriptase Activation
- Dissecting the Gene Network of Dietary Restriction to Identify Evolutionarily Conserved Pathways and New Functional Genes
- Identification of Human Proteins That Modify Misfolding and Proteotoxicity of Pathogenic Ataxin-1
- and Link Transcription of Phospholipid Biosynthetic Genes to ER Stress and the UPR
- CDK9 and H2B Monoubiquitination: A Well-Choreographed Dance
- Rare Copy Number Variations in Adults with Tetralogy of Fallot Implicate Novel Risk Gene Pathways
- Ccdc94 Protects Cells from Ionizing Radiation by Inhibiting the Expression of
- NOL11, Implicated in the Pathogenesis of North American Indian Childhood Cirrhosis, Is Required for Pre-rRNA Transcription and Processing
- Human Developmental Enhancers Conserved between Deuterostomes and Protostomes
- A Luminal Glycoprotein Drives Dose-Dependent Diameter Expansion of the Hindgut Tube
- Melanophore Migration and Survival during Zebrafish Adult Pigment Stripe Development Require the Immunoglobulin Superfamily Adhesion Molecule Igsf11
- Dynamic Distribution of Linker Histone H1.5 in Cellular Differentiation
- Combining Comparative Proteomics and Molecular Genetics Uncovers Regulators of Synaptic and Axonal Stability and Degeneration
- Chemical Genetics Reveals a Specific Requirement for Cdk2 Activity in the DNA Damage Response and Identifies Nbs1 as a Cdk2 Substrate in Human Cells
- Experimental Relocation of the Mitochondrial Gene to the Nucleus Reveals Forces Underlying Mitochondrial Genome Evolution
- Rates of Gyrase Supercoiling and Transcription Elongation Control Supercoil Density in a Bacterial Chromosome
- Mutations in a P-Type ATPase Gene Cause Axonal Degeneration
- A General G1/S-Phase Cell-Cycle Control Module in the Flowering Plant
- Multiple Roles and Interactions of and in Development of the Respiratory System
- UNC-40/DCC, SAX-3/Robo, and VAB-1/Eph Polarize F-Actin during Embryonic Morphogenesis by Regulating the WAVE/SCAR Actin Nucleation Complex
- Epigenetic Remodeling of Meiotic Crossover Frequency in DNA Methyltransferase Mutants
- Modulating the Strength and Threshold of NOTCH Oncogenic Signals by
- Loss of Axonal Mitochondria Promotes Tau-Mediated Neurodegeneration and Alzheimer's Disease–Related Tau Phosphorylation Via PAR-1
- Acetyl-CoA-Carboxylase Sustains a Fatty Acid–Dependent Remote Signal to Waterproof the Respiratory System
- ATXN2-CAG42 Sequesters PABPC1 into Insolubility and Induces FBXW8 in Cerebellum of Old Ataxic Knock-In Mice
- Cohesin Rings Devoid of Scc3 and Pds5 Maintain Their Stable Association with the DNA
- The MicroRNA Inhibits Calcium Signaling by Targeting the TIR-1/Sarm1 Adaptor Protein to Control Stochastic L/R Neuronal Asymmetry in
- Rapid-Throughput Skeletal Phenotyping of 100 Knockout Mice Identifies 9 New Genes That Determine Bone Strength
- The Genes Define Unique Classes of Two-Partner Secretion and Contact Dependent Growth Inhibition Systems
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Dissecting the Gene Network of Dietary Restriction to Identify Evolutionarily Conserved Pathways and New Functional Genes
- It's All in the Timing: Too Much E2F Is a Bad Thing
- Variation of Contributes to Dog Breed Skull Diversity
- The PARN Deadenylase Targets a Discrete Set of mRNAs for Decay and Regulates Cell Motility in Mouse Myoblasts
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



