-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaIntracellular Growth Is Dependent on Tyrosine Catabolism in the Dimorphic Fungal Pathogen
Fungi that infect humans are a major health problem, especially for those with compromised immune systems. Many fungal infections are extremely difficult to cure and if left untreated are fatal. For successful infection to occur, the fungal pathogen must be able to grow by acquiring and utilising the available nutrient sources within the host whilst evading or tolerating the host’s defence systems. Expression profiling in several pathogenic fungal species has revealed that genes required for tyrosine catabolism are induced specifically in the pathogenic cell type at 37°C. As well as enabling the fungus to acquire carbon and nitrogen intermediates from proteins within the host, tyrosine is also an important precursor in the formation of two different types of melanin, which protects cells against the host’s defence systems. This study shows that the ability to catabolise tyrosine and produce tyrosine derived melanin is not required for the initial stages of fungal infection. However, a novel role for hpdA, which encodes the enzyme which catalyses the second step of tyrosine catabolism, was identified during growth in host cells.
Published in the journal: . PLoS Pathog 11(3): e32767. doi:10.1371/journal.ppat.1004790
Category: Research Article
doi: https://doi.org/10.1371/journal.ppat.1004790Summary
Fungi that infect humans are a major health problem, especially for those with compromised immune systems. Many fungal infections are extremely difficult to cure and if left untreated are fatal. For successful infection to occur, the fungal pathogen must be able to grow by acquiring and utilising the available nutrient sources within the host whilst evading or tolerating the host’s defence systems. Expression profiling in several pathogenic fungal species has revealed that genes required for tyrosine catabolism are induced specifically in the pathogenic cell type at 37°C. As well as enabling the fungus to acquire carbon and nitrogen intermediates from proteins within the host, tyrosine is also an important precursor in the formation of two different types of melanin, which protects cells against the host’s defence systems. This study shows that the ability to catabolise tyrosine and produce tyrosine derived melanin is not required for the initial stages of fungal infection. However, a novel role for hpdA, which encodes the enzyme which catalyses the second step of tyrosine catabolism, was identified during growth in host cells.
Introduction
A number of pathogenic microbes reside within phagocytic cells of the host innate immune system, a strategy that minimizes exposure to the adaptive immune response but which requires subversion or escape from the cytotoxic capacity of innate immune cells. The acquisition of nutrients for growth is a key challenge faced by these intracellular pathogens, which have to scavenge nutrients from the relatively nutrient poor environment of the macrophage. Expression profiling studies using a number of intracellular pathogens has revealed distinct changes to metabolism upon phagocytosis, specifically for genes involved in carbon assimilation from host proteins and amino acids [1, 2, 3, 4]. Many fungi can readily assimilate amino acids as both carbon and nitrogen sources and proteins are relatively abundant in host cells. There is good evidence that amino acids are an important nutritional source for pathogenic fungi. For example, genes required for tyrosine catabolism are induced under infection conditions in the fungal pathogens Paracoccidioides brasiliensis, Histoplasma capsulatum, Penicillium marneffei (recently renamed Talaromyces marneffei) and Aspergillus fumigatus, suggesting that tyrosine may provide an important source of carbon and/or nitrogen during infectious growth [2, 4, 5, 6]. Tyrosine is catabolized via a pathway that is conserved across the kingdoms. Garrod originally identified this pathway in his classic work on inborn errors of metabolism in humans and the full pathway was uncovered by elegant studies in the model fungus Aspergillus nidulans [7, 8]. The catabolism of tyrosine provides the fungus with nitrogen from a transamination reaction with α-ketoglutarate to produce glutamate and carbon via production of fumarate and acetoacetate, which is further catabolised to acetyl-CoA, which feed into the TCA cycle [7].
As well as enabling the fungus to acquire carbon and nitrogen intermediates from proteins within the host, tyrosine is also an important precursor in the formation of two different types of melanin. Melanins are a large group of pigment macromolecules which although chemically diverse, display similar chemical properties and function to protect cells from environmental stress. Melanins are broadly classified into one of three groups: pheomelanins, eumelanins and allomelanins (pyomelanins and DHN-melanins) [9]. Tyrosine can be hydroxylated by tyrosinases and/or laccases to produce DOPA that can then be used for the formation of the eumelanin DOPA melanin (L-3, 4-dihydroxyphenylalanine melanin), while catabolism of tyrosine produces the pathway intermediate homogentisate that can be used to generate the allomelanin pyomelanin through oxidation and polymerization [10, 11]. The two most commonly identified melanins in fungi are DOPA-melanin and DHN-melanin (1,8-dihydroxynaphthalene melanin); the latter being synthesised from polyketides made from acetate precursors. Both DHN - and DOPA-melanin have been shown to protect fungal cells from RNS and ROS derived from host macrophages [12, 13, 14, 15]. Mutants which block the biosynthesis of DOPA-melanin in Cryptococcus neoformans and DHN-melanin in A. fumigatus, P. marneffei and Wangiella dermatitidis show reduced virulence in murine models of infection [12, 13, 14, 15, 16]. Melanisation has also been shown to influence phagocytosis, phagolysosomal maturation and the release of proinflammatory cytokines during infection [17, 18, 19, 15, 20, 21, 22].
A number of important human fungal pathogens are dimorphic and switch from a non-pathogenic multicellular hyphal form found outside the host to a unicellular yeast growth form during infection. Microarray-based expression profiling in P. marneffei, P. brasiliensis and H. capsulatum, to identify differential expression between the two growth types, has revealed that genes required for tyrosine catabolism are induced specifically in the pathogenic cell type at 37°C [2, 4, 6]. The melanin synthesized ex vivo by these pathogens has been previously postulated to be either or both of the two most commonly identified melanins, DHN-melanin and DOPA-melanin [23, 24, 25, 26]. However, this raises the possibility that these fungal pathogens may be producing the third type of melanin, pyomelanin, via the oxidation and polymerization of homogentisate produced during tyrosine catabolism [10, 11]. To investigate this possibility, and the role of tyrosine catabolism during infectious growth, the role of genes required for the catabolism of tyrosine was investigated in P. marneffei. The genes required for tyrosine catabolism are located within a conserved gene cluster. This study shows that the expression of genes within the cluster is both positively and negatively regulated by a gene (hmgR) encoding a C6 binuclear cluster DNA binding motif transcription factor, present within the cluster, in response to nitrogen source availability. The expression of tyrosine catabolic cluster genes is also under the regulation of the global GATA-type transcription factor AreA. Deletion of genes of the tyrosine catabolic cluster reveals the requirement of the cluster for growth on tyrosine as both a nitrogen and carbon source at both 25°C during hyphal growth and at 37°C during yeast growth. This study also shows that P. marneffei produces pyomelanin when grown on medium containing tyrosine at 37°C and this melanisation requires genes of the tyrosine catabolic cluster. Deletion of the genes of the tyrosine catabolic cluster, with the exception of hpdA, does not affect infectious growth in macrophages. This result suggests that tyrosine catabolism and pyomelanin formation are not required for the initial stages of infection, however, HpdA has an additional novel role during ex vivo infectious growth of P. marneffei.
Results
HpdA is essential for ex vivo yeast cell production during infection
Microarray-based expression profiling in P. marneffei revealed that hpdA, encoding 4-hydroxyphenylpyruvate dioxygenase which catalyses the conversion of 4-hydroxyphenylpyruvate to homogentisate during tyrosine catabolism, is induced specifically in the pathogenic cell type at 37°C [6]. To assess if hpdA is required for ex vivo yeast growth, wildtype conidia were used to infect murine J774 macrophages in the absence and presence of the 4-hydroxyphenylpyruvate dioxygenase (HpdA) chemical inhibitor NTBC (2-(2-nitro-4-trifluoromethylbenzoyl)-cyclohexane-1, 3-dione) and examined 24 hours post-infection [2, 27, 28]. After 24 hours, wildtype conidia within macrophages have completed isotropic expansion followed by elongation into a cylindrical yeast cell. Macrophages infected with wildtype conidia contain numerous yeast cells, some of which are dividing by fission (Fig. 1A and B). In contrast, predominately ungerminated conidia were observed in macrophages in the presence of NTBC (Fig. 1A and B). This suggests that tyrosine catabolism plays an important role during ex vivo yeast cell production in P. marneffei.
Fig. 1. hpdA is essential for yeast cell production during P. marneffei ex vivo growth in macrophages. 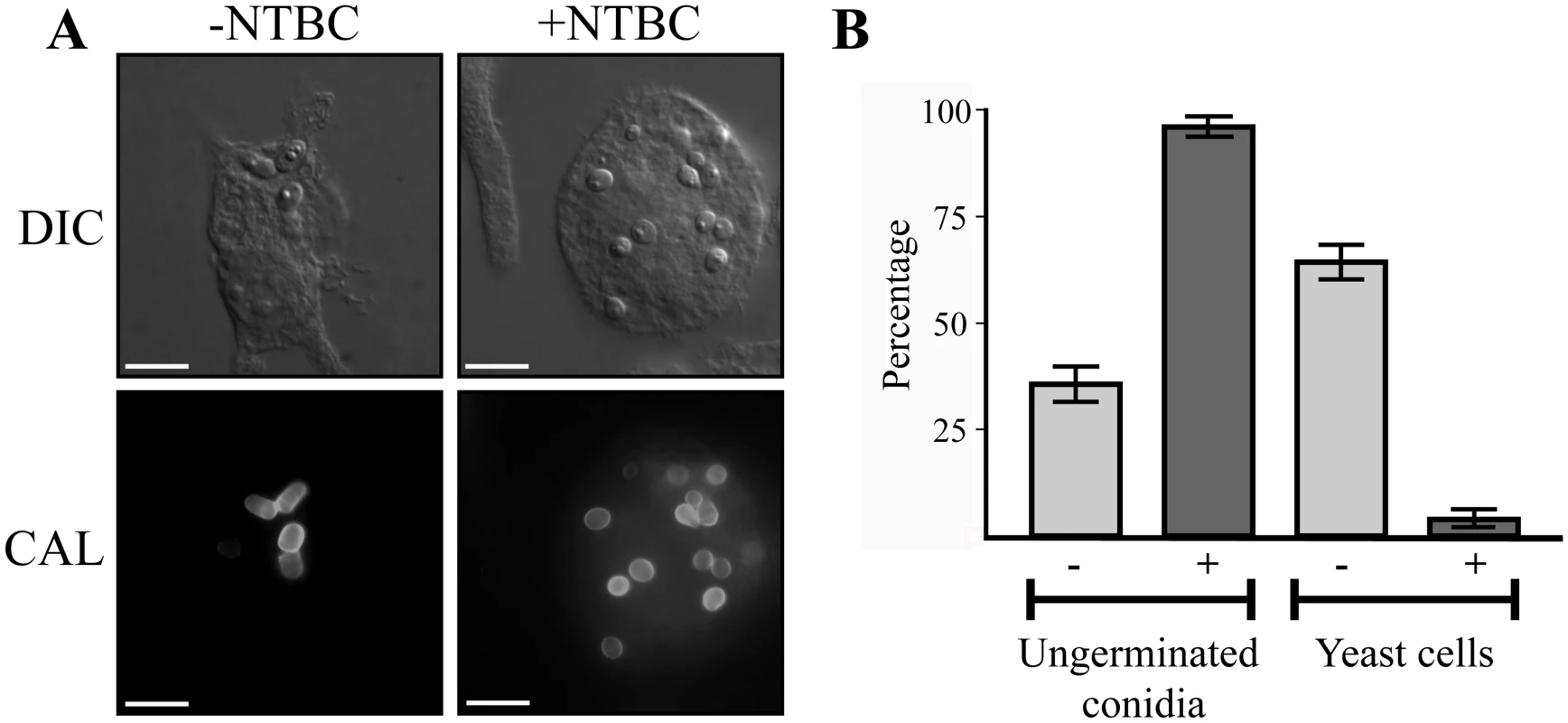
A. Macrophages infected with wildtype conidia without (-) or with (+) 400μg mL-1 of the HpdA inhibitor NTBC added to the macrophage media at the time of infection. After 24 hours, macrophages infected with wildtype conidia without NTBC added contain numerous yeast cells dividing by fission. In contrast, the addition of NTBC results in predominately ungerminated conidia remaining in macrophages 24 hours post-infection. B. Quantitation of the numbers of ungerminated conidia and yeast cells after 24 hours without (-) or with (+) NTBC. Genes required for tyrosine catabolism are located in a conserved gene cluster
Tyrosine is assimilated via a conserved catabolic pathway that provides the fungus with both nitrogen, from a transamination reaction with α-ketoglutarate to produce glutamate, and carbon via production of fumarate and acetoacetate, which is used to generate acetyl-CoA that can then feed into the TCA cycle (Fig. 2A). Microarray-based expression profiling identified the hpdA gene as highly upregulated in yeast cells compared to hyphal cells [6]. Annotation of the 30Kb genomic region which encompasses P. marneffei hpdA shows the genes encoding additional enzymes required for the catabolism of tyrosine (hmgA, maiA and fahA) are located in a gene cluster (Fig. 2B). Tyrosine catabolism genes are also clustered in Aspergillus species [7, 8, 11]. The P. marneffei gene cluster also contains a gene (hmgR) encoding a Zn(II)2-Cys6 binuclear cluster transcription factor which has been shown to be required for expression of genes required for tyrosine catabolism in A. fumigatus, and a conserved gene, hmgX, which is postulated to act as an accessory factor to HpdA [5](Fig. 2B). The P. marneffei tyrosine cluster contains an additional ORF PMAA031970, we have named hypW, which is absent in the Aspergilli and other dimorphic pathogens. hypW is predicted to encode a hypothetical protein of unknown function which lacks any predicted domains (Fig. 2B). The P. marneffei tyrosine catabolism gene cluster also contains an additional gene, mfpA (PMAA_032010), encoding a putative alpha-1, 2-mannosidase family protein which is conserved in other fungi but located elsewhere in the genome (Fig. 2B). Blast searches of the P. marneffei genome also revealed the presence of paralogous genes scattered throughout the genome. A comparison of other fungal genomes revealed this is common to many species (Table 1). The P. marneffei genome contains a hpdA paralogue (PMAA_089170), four hmgA paralogues (PMAA_102000, PMAA_054730, PMAA_035450 and PMAA_080510) and two fahA paralogues (PMAA_080500 and PMAA_099050).
Fig. 2. The catabolism of tyrosine. 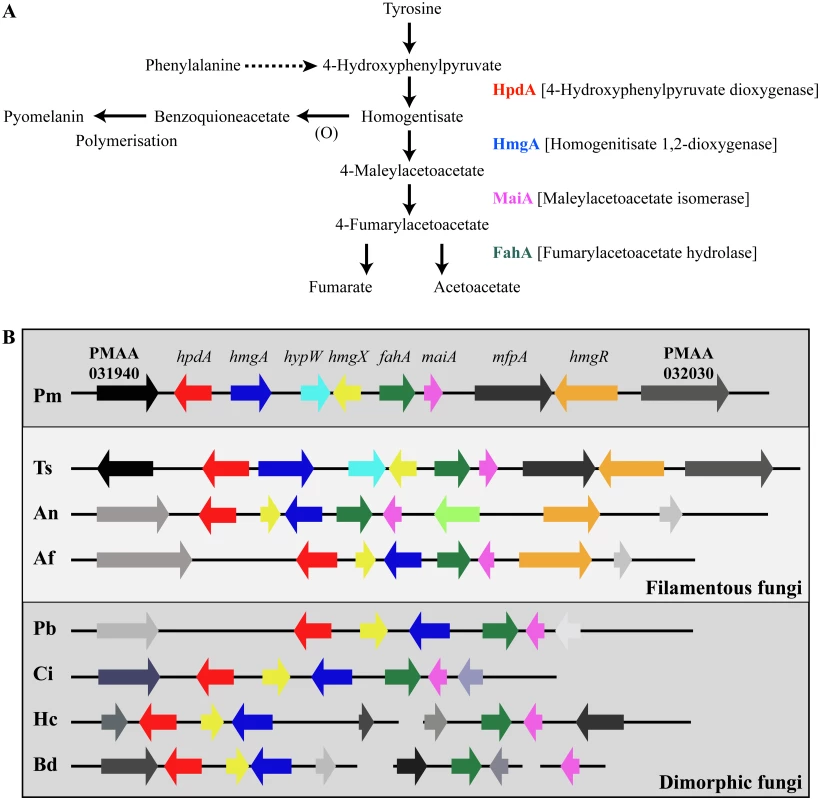
A. Tyrosine is catabolised to the toxic compound 4-hydroxyphenylpyruvate by an unknown mechanism. Phenylalanine can be degraded by the prephenate pathway to 4-hydroxyphenylpyruvate. 4-hydroxyphenylpyruvate dioxygenase (HpdA) catalyses the conversion of 4-hydroxyphenylpyruvate to homogentisate. Homogentisate can either be oxidized and polymerized to form the brown pigment pyomelanin or metabolised to 4-maleylacetoacetate by homogentisate 1,2-dioxygenase (HmgA). 4-maleylacetoacetate is metabolised to 4-fumarylacetoacetate by maleylacetoacetate isomerase (MaiA). Fumarlacetoacetate hydrolase (FahA) catalyses the conversion of 4-fumarylacetoacetate to acetoacetate and fumarate, which can be utilized as carbon sources via the TCA cycle. B. Genes required for the catabolism of tyrosine are located in a gene cluster which is conserved in filamentous (Talaromyces stipitatus; Ts, Aspergillus nidulans; An and Aspergillus fumigatus; Af, light grey box) and dimorphic (P. marneffei; Pm, Paracoccidioides brasiliensis; Pb, Coccidioides immitis; Ci, Histoplasma capsulatum; Hc and Blastomyces dermatitidis; Bd, dark grey boxes) fungi. Tyrosine catabolism genes are coloured as follows: hpdA; red, hmgA; blue, hypW; aqua, hmgX; yellow, fahA; green, maiA; pink and hmgR; orange. Flanking genes (PMAA_031940, PMAA_032030, TSTA_065640, TSTA_065560, AN1900, AN1892, AFUA_2G04190, AFUA_2G04270, PADG_08469, PADG_08463, CIMG_01309, CIMG_01315, HCEG_08529, HCEG_08533, HCEG_03254, HCEG_03257, BDDG_05746, BDDG_05738, BDDG_08624 and BDDG_12986) with no characterized role in tyrosine catabolism are shown in grey (same shade if orthologous). hypW is present only in P. marneffei (PMAA_032010) and T. stipitatus (TSTA_065580). A. nidulans contains an internal gene (AN1894) with no characterized role in tyrosine catabolism (light green). T. stipitatus fahA and maiA are misannotated in the Talaromyces stipitatus ATCC 10500 genomic database as a single fused gene named fahA (TSTA_065590). The cluster has been divided into two in H. capsulatum and B. dermatitidis. P. brasiliensis, C. immitis, H. capsulatum and B. dermatitidis lack a hmgR orthologue. Gene orthologues used to generate this Figure are as follows: hpdA (PMAA_031950, TSTA_065630, AN1899, AFUA_2G04200, PADG_08468, CIMG_01310, HCEG_08530 and BDDG_05744), hmgA (PMAA_031960, TSTA_065620, AN1897, AFUA_2G04220, PADG_08466, CIMG_01312, HCEG_08532 and BDDG_05741), hypW (PMAA_031970 and TSTA_065610), hmgX (PMAA_031980, TSTA_065600, AN1898, AFUA_2G04210, PADG_08467, CIMG_01311, HCEG_08531 and BDDG_05742), fahA (PMAA_031990, TSTA_065590, AN1896, AFUA_2G04230, PADG_08465, CIMG_01313, HCEG_03255 and BDDG_08623), maiA (PMAA_032000, AN1895, AFUA_2G04240, PADG_08464, CIMG_01314, HCEG_03256 and BDDG_08618), mfpA (PMAA_031010, TSTS_065580), and hmgR (PMAA_032020, TSTA_065570, AN1893 and AFUA_2G04262). Tab. 1. Paralogues of tyrosine catabolic cluster genes. 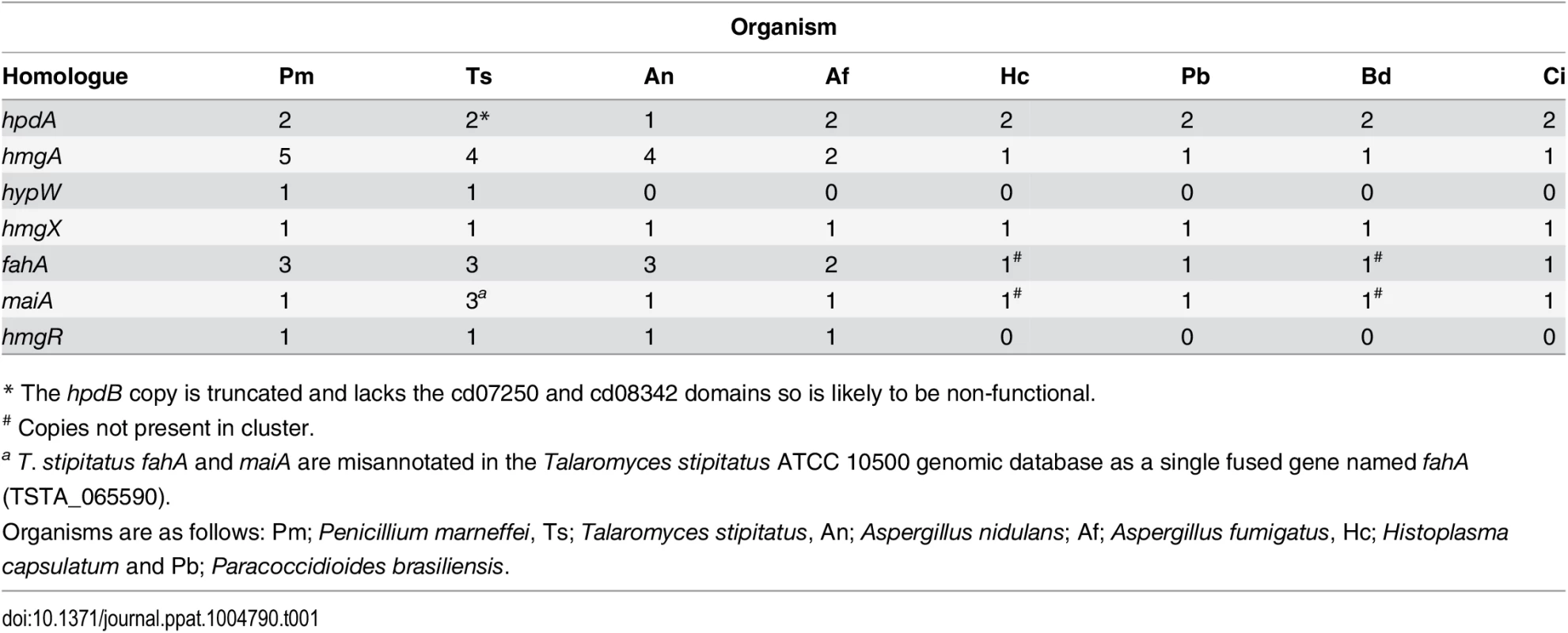
* The hpdB copy is truncated and lacks the cd07250 and cd08342 domains so is likely to be non-functional. To investigate if this cluster is conserved in other fungi, approximately 30Kb of the genomic region which encompasses the orthologue of hpdA, from P. marneffei’s closest sexual relative T. stipitatus, from the dimorphic fungal pathogens P. brasiliensis, Coccidioides immitis, H. capsulatum and Blastomyces dermatitidis, and the filamentous fungi A. nidulans and A. fumigatus was compared to that of P. marneffei (Fig. 2B). The homologues of hmgA, maiA, fahA and hmgX were within close proximity of hpdA in all species except H. capsulatum and B. dermatitidis in which the catabolic cluster was split into two and three, respectively, different genomic locations (Fig. 2B). T. stipitatus fahA and maiA are misannotated in the database as a single fused gene named fahA (TSTA_065590). The cluster in T. stipitatus also contains a homologue of hypW and mfpA (Fig. 2B). P. brasiliensis, C. immitis, H. capsulatum and B. dermatitidis also lacked a homologue of the Zn(II)2-Cys6 binuclear cluster transcription factor encoded by hmgR.
The expression of tyrosine catabolic cluster genes is regulated by nitrogen source via the HmgR and AreA transcription factors
To investigate the expression of genes in the tyrosine catabolic cluster, RNA was isolated from cells grown in liquid culture for 2 days at 25°C (hyphal cells) or 6 days at 37°C (yeast cells) then transferred into medium containing ammonium or tyrosine as the sole nitrogen source at 25°C or 37°C for 4 hours. At both 25°C and 37°C, expression of hmgA, hmgX, fahA and maiA was low in the presence of ammonium and high in tyrosine suggesting that expression is induced in the presence of tyrosine (Fig. 3). Although hpdA showed the same pattern of expression at 25°C, at 37°C expression in ammonium and tyrosine was almost equivalent suggesting that unlike the other genes in the cluster, hpdA expression at 37°C is not repressed in the presence of a preferred nitrogen source (Fig. 3). The hmgR gene was expressed in the presence of both ammonium and tyrosine at both 25°C and 37°C, with expression only slightly higher in tyrosine (Fig. 3). Interestingly, expression of mfpA, the gene within the tyrosine metabolic cluster only in P. marneffei and T. stipitatus which is not predicted to have a role in tyrosine catabolism, was induced on tyrosine at both 25°C and 37°C (Fig. 3). This suggests that the cluster may also be under a more global level of regulation such as that mediated by chromatin effects and any gene captured within this genomic region may consequently come under it’s regulation. The analysis of hypW expression using RT-PCR primers spanning the predicted intron showed an expression pattern similar to the other genes of the tyrosine catabolic cluster (S3A Fig). However, the size of the RT PCR product suggested the predicted intron is not spliced. Examination of RNAseq data for this region confirmed the lack of this predicted intron and indicated that the predicted start site is incorrect (H. Weerasinghe and A. Andrianopoulos, personal communication)(S3B Fig). The small number of the RNA seq reads were also restricted to the 5’ region which is in close proximity to hmgX (163bp) (S3B Fig). Overall, this suggests that this ORF is not expressed under the conditions tested and the expression observed by RT PCR is due to read through from hmgX. Expression of the paralogues of tyrosine catabolic genes could not be detected under these conditions.
Fig. 3. The tyrosine induced expression of genes of the catabolism cluster requires the HmgR transcription factor. 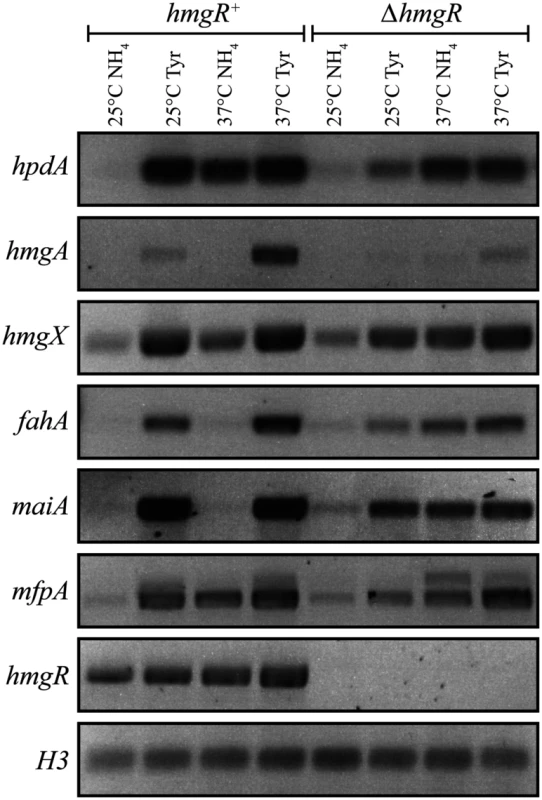
RNA was isolated from wildtype (hmgR+) and ΔhmgR strains grown in liquid culture for 2 days at 25°C or 6 days at 37°C and transferred into media containing ammonium (NH4) or tyrosine (Tyr) as the sole nitrogen source at 25°C or 37°C for 4 hours. Expression of hpdA (PMAA_031950), hmgA (PMAA_031960), hmgX (PMAA_031980), fahA (PMAA_031990), maiA (PMAA_032000), mfpA (PMAA_031010) and hmgR (PMAA_032020) was detected by RT PCR. To investigate if the tyrosine-induced expression of the genes in the cluster is a result of a general response to limiting nitrogen levels or a specific induction by tyrosine, hpdA, maiA and fahA expression was also assessed in alanine at 25°C and 37°C. Expression of these genes was higher in alanine compared to ammonium but lower than tyrosine at both temperatures. This suggests that the increased expression observed in tyrosine is not solely due to derepression of tyrosine catabolic genes in response to general limiting nitrogen levels but rather specific induction due to the presence of tyrosine (Fig. 4A and B).
Fig. 4. AreA negatively regulates expression of tyrosine catabolism genes. 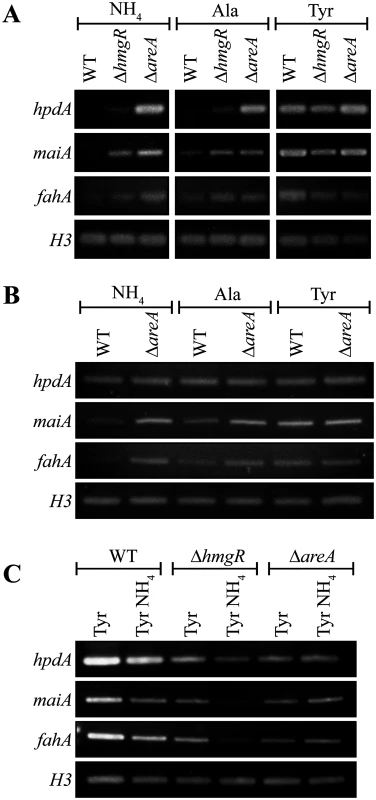
A. RNA was isolated from wildtype (WT), ΔhmgR and ΔareA strains grown in liquid culture for 2 days at 25°C (A and C) or 6 days at 37°C (B) and transferred into media containing ammonium (NH4), alanine (Ala) or tyrosine (Tyr) as the sole nitrogen source at 25°C (A) or at 37°C (B) for 4 hours, or media containing tyrosine (Tyr) or both tyrosine and ammonium (Tyr NH4) at 25°C for 4 hours (C). Expression of hpdA, maiA, fahA and a H3 loading control was detected by RT PCR. A gene encoding a Zn(II)2-Cys6 binuclear transcription factor, hmgR, is conserved in the tyrosine metabolic cluster across many fungi. This gene has been shown to be essential for tyrosine-induced expression of hppD (hpdA orthologue), hmgA, hmgX, fahA and maiA in A. fumigatus [5]. To investigate the role of hmgR in P. marneffei, the orthologous gene (PMAA_032020) was cloned and a deletion strain (ΔhmgR::pyrG+ (G825)) was generated by transforming P. marneffei strain G816 (ΔligD niaD1 pyrG1). To generate a complemented strain (ΔhmgR hmgR+ (G867)), a ΔhmgR pyrG- (G864) strain was transformed with a 5.1 kb hmgR fragment targeted to the P. marneffei pyrG locus.
In contrast to A. fumigatus, deletion of hmgR in P. marneffei resulted in only the partial loss of induction of hpdA, hmgA, hmgX, fahA and maiA in medium with tyrosine as the sole nitrogen source at both 25°C and 37°C (Fig. 3). This suggests that this transcription factor is required, but not essential, for tyrosine-induced expression in P. marneffei. Interestingly, deletion of hmgR resulted in increased expression of hpdA at 25°C and hmgA, hmgX, fahA and maiA at both 25°C and 37°C in medium containing ammonium as the sole nitrogen source (Fig. 3). Therefore, in addition to inducing the expression of tyrosine catabolism genes in the presence of tyrosine, this transcription factor is also required to repress their expression in the presence of a preferred nitrogen source.
Nitrogen metabolite repression is a regulatory mechanism utilized by microbes to allow the preferential use of readily assimilated (preferred) nitrogen sources. The areA gene encodes a positively-acting GATA-type transcription factor which regulates the global response to limiting nitrogen conditions in P. marneffei and other fungi [29, 30]. To investigate the contribution of AreA regulation to the tyrosine catabolic cluster, the expression of hpdA, maiA and fahA was examined in wildtype, ΔhmgR and ΔareA at 25°C and 37°C. RNA was isolated from cells grown in liquid culture for 2 days at 25°C or 6 days at 37°C and transferred into medium containing ammonium, alanine or tyrosine as the sole nitrogen source at 25°C or 37°C for 4 hours. In wildtype, expression of maiA and fahA was barely detectable in ammonium, increased to a low level in alanine and strongly induced in tyrosine at both 25°C and 37°C (Fig. 4A and B). Expression levels of hpdA were similarly dependent on the nitrogen source at 25°C but constitutive at 37°C (Fig. 4A and B). Compared to wildtype, the expression of hpdA, maiA and fahA in the ΔhmgR mutant at 25°C was partially derepressed in ammonium and alanine and full induction in tyrosine was not observed (Fig. 4A). Surprisingly unlike wildtype and the ΔhmgR mutant, the expression of hpdA, maiA and fahA on either ammonium, alanine or tyrosine was equivalent in the ΔareA mutant at 25°C (Fig. 4A). This was unexpected given that AreA usually acts as a positive regulator of gene expression under conditions of limiting nitrogen. Likewise at 37°C, the expression of maiA and fahA was equivalent on either ammonium, alanine or tyrosine in the ΔareA mutant (Fig. 4B). Deletion of areA did not affect the highly constitutive expression of hpdA at 37°C (Fig. 4B). These results suggest that AreA is not acting to positively regulate expression of the tyrosine catabolic cluster in the presence of tyrosine. In addition, the increased expression of these genes in the ΔareA mutant on ammonium and alanine suggests that AreA is negatively regulating expression of the cluster in the absence of tyrosine which is contradictory to the current model of AreA function.
To investigate if genes of the tyrosine catabolic cluster are under nitrogen metabolite repression mediated by AreA, the expression of hpdA, maiA and fahA was assessed on tyrosine and both tyrosine and ammonium at 25°C in both wildtype, ΔhmgR and ΔareA (Fig. 4C). In wildtype, the levels of hpdA, maiA and fahA are lower on tyrosine and ammonium medium compared to tyrosine alone suggesting that these genes are under nitrogen metabolite repression (Fig. 4C). Although the overall level of induction is reduced in the ΔhmgR mutant, the levels of hpdA, maiA and fahA are lower on tyrosine and ammonium medium compared to tyrosine alone suggesting that HmgR is not playing a role in nitrogen metabolite repression (Fig. 4C). In contrast, in the ΔareA mutant the levels of hpdA, maiA and fahA expression are equivalent between tyrosine and ammonium medium compared to tyrosine alone indicating that areA is required during nitrogen metabolite repression to repress expression of the cluster in the presence of ammonium (Fig. 4C).
Genes of the tyrosine catabolic cluster are required for hyphal growth on tyrosine as a nitrogen or carbon source at 25°C
To determine the role of genes present in the tyrosine catabolic cluster, hpdA, hmgA, hypW, hmgX, maiA and mpfA were cloned and deleted. Due to the fact that an intermediate in the tyrosine catabolism pathway, homogentisate, can be oxidized and polymerized to form the brown pigment pyomelanin, the wA gene required for the synthesis of DHN melanin was also cloned and deleted for comparison. As expected, the ΔwA strain appears white due to the absence of DHN melanin in asexual spores (conidia) (Fig. 5). To confirm the mutant phenotype was a result of the gene deletion events, deletions strains were complemented with the wildtype gene targeted to either pyrG or niaD.
Fig. 5. Genes of the tyrosine catabolic cluster are required for hyphal growth on tyrosine and phenylalanine as a nitrogen or carbon source at 25°C.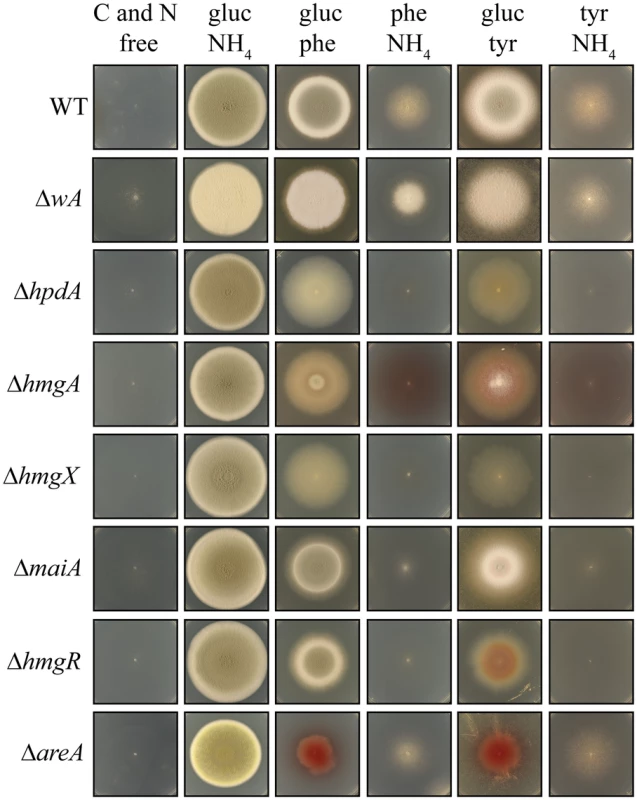 
Growth of the wildtype, ΔwA, ΔhpdA, ΔhmgA, ΔhmgX, ΔmaiA, ΔhmgR, and ΔareA strains on carbon and nitrogen free medium (C and N free), on ammonium as the sole nitrogen source (gluc NH4), on phenylalanine as the sole nitrogen source (gluc phe), on phenylalanine as the sole carbon source (phe NH4), on tyrosine as the sole nitrogen source (gluc tyr) or on tyrosine as the sole carbon source (tyr NH4) after 14 days at 25°C. Tyrosine can be utilized as both nitrogen and carbon sources for growth. In A. fumigatus, an L-amino oxidase is thought to catalyse the conversion of tyrosine to the α-keto acid, liberating ammonium as a nitrogen source. However, the P. marneffei genome lacks genes encoding L-amino oxidases so it is likely that a transamination reaction of tyrosine with α-ketoglutarate to produce glutamate as a nitrogen source is the first catabolic step. The enzyme that performs this reaction is currently unknown. The catabolism of tyrosine also produces fumarate and acetoacetate, which is further catabolised to acetyl-CoA, which feed into the TCA cycle to provide carbon (Fig. 2A). Phenylalanine is also catabolised via the tyrosine catabolism pathway to provide both nitrogen and carbon, although the enzymatic steps required to convert it to 4-hydroxyphenylpyruvate remain unclear. Growth of the wildtype, ΔwA, ΔhpdA, ΔhmgA, ΔhmgX and ΔmaiA strains was assessed at 25°C on a range of media in which the sole carbon source was either glucose, tyrosine or phenylalanine and the sole nitrogen source was either ammonium, tyrosine or phenylalanine or on medium lacking both a nitrogen and carbon source. Wildtype and the ΔwA strain grew well on tyrosine or phenylalanine as a sole nitrogen or carbon source (Fig. 5). Compared to wildtype and the ΔwA strain, the ΔhpdA, ΔhmgA, ΔhmgX and ΔmaiA strains showed reduced growth on tyrosine and phenylalanine as the sole nitrogen source which suggests a feedback loop regulates the nitrogen-liberating first step in the catabolism of tyrosine (Fig. 5). Unlike wildtype and the ΔwA strain, the ΔhpdA, ΔhmgA, ΔhmgX and ΔmaiA strains showed no growth on tyrosine or phenylalanine as the sole carbon source (Fig. 5). The medium became pigmented when the ΔhmgA mutant was grown on tyrosine or phenylalanine as the sole nitrogen or carbon source (Fig. 5). This suggests that accumulated homogentisate is being oxidized to produce pyomelanin, as has been observed in the Aspergilli [10, 11, 31]. Reintroduction of the wildtype gene to generate the ΔhmgA hmgA+, ΔhmgX hmgX+, ΔmaiA maiA+ complemented strains restored growth on tyrosine and phenylalanine as the sole nitrogen and carbon source at 25°C but to a level slightly below that of wildtype (S1A Fig). The ΔhpdA hpdA+ strain did not show the same extent of restoration of growth on tyrosine and phenylalanine as the sole carbon source, indicating either the complementation construct may lack some regulatory sequences required for full expression or the location of the gene in the cluster, as opposed to an ectopic site, is important for it’s regulation (S1A Fig). The growth phenotypes of the ΔhpdA and ΔhmgA strains suggests that there is no functional overlap between these genes and the paralogues located elsewhere in the genome with respect to this catabolic pathway. To confirm that these paralogues do not have a role in tyrosine catabolism, hpdB was cloned and deleted. This gene was selected as hpdA has only a single paralogue in the genome, whereas, hmgA has four paralogues. Unlike ΔhpdA, the ΔhpdB strain was indistinguishable from wildtype on tyrosine or phenylalanine as either the sole nitrogen or carbon source at 25°C and 37°C supporting the hypothesis that there is no functional overlap between hpdA and hpdB (S1B Fig and S4B Fig).
Accumulation of intermediates of tyrosine catabolism results in cellular toxicity due to the production of intermediary metabolites or their spontaneous degradation products (4-hydroxyphenlpyruvic acid in hpdA mutants, homogentistic acid in hmgA mutants and succinylacetone and succinylacetoacetate from fumarylacetoacetate in fahA mutants) [7, 8, 10, 32, 33]. To investigate if toxic metabolites are accumulating in the P. marneffei tyrosine catabolism mutants, the wildtype, ΔhpdA, ΔhmgA, ΔhmgX and ΔmaiA strains were grown on tyrosine medium also containing the non-repressive carbon sources sorbitol, lactose, acetate or proline. In contrast to wildtype, the ΔhpdA and ΔhmgA strains showed no growth under these conditions indicating that toxic intermediates are accumulating in these strains (S2 Fig). Growth of the ΔhmgX strain was less than wildtype but greater than the ΔhpdA and ΔhmgA mutants. This suggests that the levels of toxic intermediates are lower in this strain compared to ΔhpdA and ΔhmgA (S2 Fig). Growth of the ΔmaiA was unaffected on tyrosine medium also containing the non-repressive carbon sources suggesting that toxic intermediates are not accumulating in this strain (S2 Fig). These results are in contrast to A. nidulans in which deletion of hpdA and maiA, but not hmgA, results in cellular toxicity [10, 32, 33]. These results suggest putative differences between the metabolites derived from homogentisate and 4-malelyacetate in P. marneffei versus A. nidulans.
To assess if the genes within the cluster that are not conserved in other species also have a role in tyrosine catabolism in P. marneffei, growth of the ΔmfpA and ΔhypW strains was assessed at 25°C on glucose, tyrosine or phenylalanine as the sole carbon source, ammonium, tyrosine or phenylalanine as the sole nitrogen source or on medium lacking both a nitrogen and carbon source. As expected, the ΔmfpA strain was indistinguishable from wildtype under these conditions indicating that despite being transcriptionally regulated in response to the presence of tyrosine mfpA is not required for tyrosine catabolism (S1B Fig). However, unexpectedly, the ΔhypW strain showed reduced growth on tyrosine and phenylalanine as the sole nitrogen or carbon source compared to wildtype (S3C Fig). Due to the low transcriptional activity in this region and the ORF’s close proximity to hmgX, we hypothesized that deletion of this ORF may be affecting hmgX expression. To assess this, hmgX expression was evaluated in the ΔhypW strain at 25°C in ammonium and tyrosine as the sole nitrogen source. Deletion of hypW reduced expression of hmgX under both conditions suggesting that the reduced growth on tyrosine as the sole nitrogen source is a consequence of reduced hmgX expression rather than indicating a role for hypW in tyrosine catabolism (S3D Fig). In support of this hypothesis, the introduction of a construct containing both the hypW and hmgX wildtype genes complemented the phenotype of the ΔhypW strain at both 25°C and 37°C, whereas, introduction of a construct containing wildtype hypW and a truncated hypX lacking the start codon, did not complement this growth phenotype (S3C Fig). Therefore, hypW is not required for tyrosine catabolism
To further investigate the role of regulatory genes on the tyrosine catabolic cluster, the growth of the ΔhmgR and ΔareA strains was assessed at 25°C on glucose, tyrosine or phenylalanine as the sole carbon source, ammonium, tyrosine or phenylalanine as the sole nitrogen source or on medium lacking both a nitrogen and carbon source. Compared to wildtype, the ΔhmgR strain showed reduced growth on tyrosine and phenylalanine as the sole nitrogen or carbon source (Fig 5). Reintroduction of the wildtype gene in the ΔhmgR hmgR+ complemented strain restored growth on tyrosine and phenylalanine as the sole nitrogen and carbon source at 25°C (S2A Fig). This growth reduction correlates with the reduced expression of tyrosine catabolic genes on tyrosine as a sole nitrogen source in the ΔhmgR mutant (Fig. 3). To assess if the growth reduction of the ΔhmgR strain was specific to tyrosine and phenylalanine, the ΔhmgR strain was also grown on a variety of amino acids as the sole nitrogen source at 25°C and 37°C. No reduction in growth was observed on any other amino acids tested. The ΔareA strain showed reduced growth on tyrosine or phenylalanine as the sole nitrogen source but was unaffected on tyrosine and phenylalanine as a carbon source (Fig. 5). As AreA does not directly regulate expression of the tyrosine catabolic genes in the gene cluster (Fig. 4), this result suggests that the deamination step is AreA dependent and in it’s absence there can be no metabolites to flux through the pathway.
Yeast growth at 37°C on tyrosine as a nitrogen or carbon source requires genes of the tyrosine catabolic cluster
To assess if the tyrosine catabolic gene cluster is required for yeast-phase growth on tyrosine at 37°C, the wildtype, ΔwA, ΔhpdA, ΔhmgA, ΔhmgX and ΔmaiA strains were grown on glucose, tyrosine or phenylalanine as the sole carbon source and ammonium, tyrosine or phenylalanine as the sole nitrogen source at 37°C. Growth of wildtype on tyrosine or phenylalanine as a sole nitrogen or carbon source at 37°C is relatively robust when compared to growth on the preferred nitrogen and carbon sources of ammonium and glucose, respectively (Fig. 6). Compared to wildtype and the ΔwA, the ΔhpdA and ΔhmgX strains showed reduced growth on tyrosine and phenylalanine as the sole nitrogen source at 37°C (Fig. 6). However unlike 25°C, the ΔhmgA and ΔmaiA strains showed no growth reduction on tyrosine and phenylalanine as the sole nitrogen source at 37°C. This difference suggests that the feedback regulation of the pathway observed at 25°C is not operating at 37°C, possibly because of a higher demand for intermediate metabolites such as homogentisate for pyomelanin production (Fig. 6). In contrast to wildtype and the ΔwA, the ΔhpdA, ΔhmgA, ΔhmgX and ΔmaiA strains showed no growth on tyrosine or phenylalanine as the sole carbon source (Fig. 6). In the absence of growth on tyrosine as a carbon source, the un-utilized tyrosine in the medium crystalizes (Fig. 6). The ΔhmgA hmgA+, ΔhmgX hmgX+, ΔmaiA maiA+ complemented strains displayed growth almost to the same extent as wildtype on tyrosine and phenylalanine as the sole nitrogen and carbon source at 37°C (S4A Fig). Like at 25°C, the ΔhpdA hpdA+ strain did not show a complete restoration of wildtype growth supporting the hypothesis that the position of the gene in the cluster is important for it’s regulation (S4A Fig).
Fig. 6. Growth of yeast cells on tyrosine and phenylalanine as a nitrogen or carbon source at 37°C requires genes of the tyrosine catabolic cluster.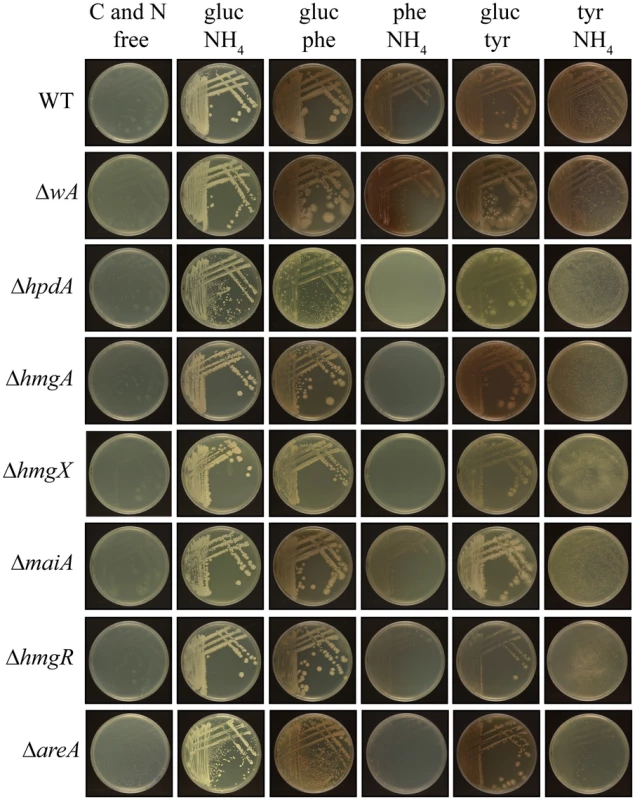 
Growth after 14 days at 37°C of the wildtype, ΔwA, ΔhpdA, ΔhmgA, ΔhmgX, ΔmaiA, ΔhmgR and ΔareA strains on carbon and nitrogen free medium (C and N free), on ammonium as the sole nitrogen source (gluc NH4), on phenylalanine as the sole nitrogen source (gluc phe), on phenylalanine as the sole carbon source (phe NH4), on tyrosine as the sole nitrogen source (gluc tyr) or on tyrosine as the sole carbon source (tyr NH4). To assess if mfpA has a role in tyrosine catabolism in P. marneffei at 37°C, growth of the ΔmfpA strain was assessed on glucose, tyrosine or phenylalanine as the sole carbon source and ammonium, tyrosine or phenylalanine as the sole nitrogen source. As expected, the ΔmfpA strain was indistinguishable from wildtype under these conditions indicating that it is not required for tyrosine catabolism (S4B Fig).
To investigate the role of regulatory genes on the tyrosine catabolic cluster at 37°C, the growth of the ΔhmgR and ΔareA strains was assessed at 37°C on glucose, tyrosine or phenylalanine as the sole carbon source and ammonium, tyrosine or phenylalanine as the sole nitrogen source. Compared to wildype, the ΔhmgR strain showed reduced growth on tyrosine and phenylalanine as the sole nitrogen or carbon source at 37°C and reintroduction of the wildtype gene restored growth (Fig. 6 and S4 Fig). This result is consistent with the reduced expression of tyrosine catabolic genes on tyrosine as a sole nitrogen source in the ΔhmgR mutant at 37°C (Fig. 3). The ΔareA strain showed reduced growth on tyrosine or phenylalanine as both the sole nitrogen and carbon source (Fig. 6).
Pyomelanin production requires genes of the tyrosine catabolic cluster
Tyrosine is an important precursor in the formation of two different types of melanin; DOPA melanin via the metabolism of tyrosine to form DOPA and pyomelanin through the oxidation and polymerization of homogentisate during tyrosine catabolism [10, 11]. To assess if P. marneffei can produce melanin from tyrosine, the wildtype strain was grown on medium containing ammonium or tyrosine as the sole nitrogen source, medium containing both ammonium plus tyrosine or tyrosine plus alanine as nitrogen sources or on L-DOPA at 37°C. P. marneffei was unpigmented on medium containing ammonium as a sole nitrogen source (Fig. 7A). In contrast, a brown pigment was evident on medium containing tyrosine as the sole nitrogen source, suggesting that this may be a melanin produced from tyrosine (Fig. 7A). This pigment was confirmed to be melanin by testing resistance to chemical degradation by boiling in acid (Materials and Methods)[23] (S5A Fig). If both ammonium and tyrosine are present the amount of melanization is greatly reduced suggesting that some of the genes required for production of this melanin are under nitrogen metabolite repression (Fig. 7A). The amount of melanization on tyrosine increases with the addition of another non-preferred nitrogen source such as alanine (Fig. 7C). Wildtype P. marneffei also becomes pigmented on L-DOPA medium indicating that P. marneffei is capable of producing DOPA melanin (Fig. 7B). To assess if deletion of genes of the tyrosine catabolic cluster affects the production of DOPA melanin, the wildtype, deletion and complementation strains were grown at 37°C on L-DOPA. The production of DOPA melanin was indistinguishable amongst all of the strains except ΔareA, which showed a decrease in DOPA melanin production (S5C Fig).
Fig. 7. Pyomelanin produced via the tyrosine catabolism pathway is under nitrogen metabolite repression.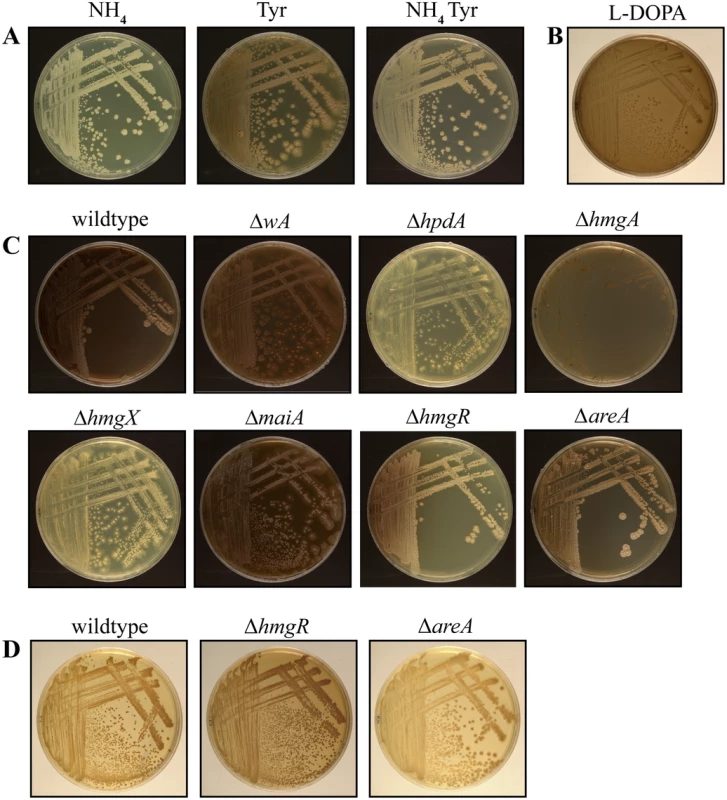 
A. Wildtype grown for 14 days at 37°C on ANM plus ammonium (NH4) or tyrosine (Tyr) as the sole nitrogen source or plus both ammonium and tyrosine (NH4 Tyr). Pyomelanin production via tyrosine catabolism is under nitrogen metabolite repression. B. Wildtype P. marneffei grown on L-DOPA medium for 14 days at 37°C. C. Pyomelanin formation on ANM plus alanine and tyrosine for 14 days at 37°C in the wildtype, ΔwA, ΔhpdA, ΔhmgA, ΔhmgX, ΔmaiA, ΔhmgR and ΔareA strains after 14 days growth at 37°C. D. Wildtype, ΔhmgR and ΔareA grown for 14 days at 37°C on ANM plus ammonium and tyrosine. The ΔhmgR mutant produces increased pyomelanin and the ΔareA mutant produces decreased pyomelanin under this growth condition. To investigate if pyomelanin production contributes to the melanization observed on tyrosine medium, the wildtype, ΔwA, ΔhpdA, ΔhmgA, ΔhmgX and ΔmaiA strains were examined after growth at 37°C on both tyrosine as a sole nitrogen source (Fig. 6) and tyrosine plus alanine as nitrogen sources (Fig. 7C). In contrast to wildtype and ΔwA, the ΔhpdA and ΔhmgX strains produced no visible melanin at 37°C when grown on tyrosine (Fig. 5 and 6C). This suggests that although capable of producing DOPA melanin from tyrosine, the major melanin produced by P. marneffei on medium containing tyrosine is the allomelanin pyomelanin produced during tyrosine catabolism. This was further confirmed by testing if melanin particles could be isolated from the ΔhpdA mutant grown on tyrosine as the sole nitrogen source for 14 days at 37°C (Materials and Methods). In contrast to wildtype under these growth conditions, no melanin particles were observed in the ΔhpdA mutant after boiling in acid (S5A Fig). This confirms that the melanin produced by P. marneffei on tyrosine is pyomelanin. The ΔmaiA strain showed a mild reduction in pyomelanin production compared to the wildtype (Fig. 6 and 7C). The ΔhmgA strain showed increased pyomelanin production on tyrosine as the sole nitrogen source (Fig. 6). The ΔhmgA strain failed to grow on medium containing both tyrosine and alanine (Fig. 7C), despite being able to grow on alanine or tyrosine when provided as sole nitrogen sources (Fig. 6). This suggests that like at 25°C toxic metabolites are accumulating in the ΔhmgA mutant, when grown with an additional non-repressing carbon source such as alanine. The ΔhpdA and ΔhmgX strains also showed reduced growth compared to wildtype on this medium suggesting some cellular toxicity, however this was to a lesser extent than at 25°C (Figs. 7C and S2 Fig). This suggests that fewer toxic metabolites accumulate in these mutants at 37°C possibly as a result of an increased demand for pyomelanin production via homogentisate. Reintroduction of the wildtype gene in the ΔhpdA hpdA+, ΔhmgA hmgA+, ΔhmgX hmgX+ and ΔmaiA maiA+ complemented strains restored pyomelanin production on tyrosine as a sole nitrogen source (S4A Fig).
To investigate if genes within the cluster which do not play a role in tyrosine catabolism and gene paralogues outside the cluster are required for pyomelanin production, the ΔmfpA and ΔhpdB strains were also grown on medium containing tyrosine as the sole nitrogen source and medium with both tyrosine and alanine as the nitrogen sources at 37°C. Consistent with no role in tyrosine catabolism, the ΔmfpA and ΔhpdB mutants were indistinguishable from wildtype under these conditions (S4B Fig).
Wildtype P. marneffei appears pigmented on the standard medium of brain heart infusion (BHI) from growth at 37°C. BHI is comprised of bovine brain and heart tissue that is postulated to be rich in phenolic compounds [23]. To assess if pyomelanin contributes to pigmentation on BHI, the wildtype, ΔwA, ΔhpdA, ΔhmgA, ΔhmgX, ΔmaiA and complemented control strains were grown at 37°C on BHI for 5 days. The ΔhpdA and ΔhmgX strains showed a large reduction in melanisation on BHI medium, whereas, the ΔhmgA strains showed an increase in melanisation (S5B Fig). The ΔmaiA strain displayed a minor decrease in melanisation and the complemented strains were indistinguishable from wildtype (S5 Fig). This suggests that pyomelanin contributes to some, but not all, of the melanisation observed when P. marneffei is grown on BHI medium. As expected, the ΔwA strain was also comparable with wildtype, suggesting that DHN melanin is not contributing to the melanisation observed on BHI medium (S5B Fig).
Deletion of hmgR results in reduced expression of the tyrosine catabolic cluster genes in the presence of tyrosine at both 25°C and 37°C (Figs. 3 and 4). To assess if this decreased expression translates into a visible decrease in pyomelanin production, the ΔhmgR and ΔhmgR hmgR+ strains were grown on medium containing both alanine and tyrosine at 37°C. Compared to wildtype and the ΔhmgR hmgR+ complemented strain, the ΔhmgR mutant showed decreased pyomelanin production suggesting that hmgR is required to positively regulate pyomelanin production in the presence of tyrosine (Fig. 7C). The ΔhmgR mutant also showed a reduction of melanization on BHI medium (S5 Fig). Deletion of hmgR also leads to partial derepression of tyrosine catabolic cluster genes in the presence of ammonium (Figs. 3 and 4A and C). To investigate if this derepression results in inappropriate pyomelanin production, the wildtype, ΔhmgR and ΔhmgR hmgR+ strains were also grown on ammonium plus tyrosine medium at 37°C. Compared to wildtype and the ΔhmgR hmgR+ complemented strain, the ΔhmgR mutant produced increased pyomelanin production on ammonium plus tyrosine medium at 37°C consistent with the partial derepression of gene expression (Fig. 7D).
Deletion of areA also leads to derepression of tyrosine catabolic cluster genes in the presence of ammonium at 25°C and 37°C and a complete loss of nitrogen metabolite repression in ammonium plus tyrosine medium at 25°C (Fig. 4A, B and C). To investigate if this derepression also results in inappropriate pyomelanin production in these strains, the wildtype and ΔareA strains were grown on ammonium plus tyrosine medium at 37°C. In contrast to the ΔhmgR mutant, no increase in pyomelanin production was observed in the ΔareA strain on ammonium plus tyrosine medium (Fig. 7D). Rather, the ΔareA strain produced less pyomelanin under this growth condition (Fig. 7D). The ΔareA strain also appeared slightly less melanised on BHI medium (S5 Fig). Similarly, deletion of areA lead to reduced pyomelanin production on alanine plus tyrosine medium (Fig. 7C). These results support the previous hypothesis that the deamination step is AreA dependent and in a ΔareA mutant there is reduced metabolic flux through the pathway.
Pyomelanin protects P. marneffei against oxidative stress
Both DHN - and DOPA-melanin have been shown to protect fungal cells from reactive oxygen species (ROS) produced by host innate immune cells [12–15, 20]. To investigate if pyomelanin plays a similar role, the wildtype, ΔwA, ΔhpdA, ΔhmgA, and complemented control strains were grown for 6 days at 37°C on medium containing tyrosine as the nitrogen source and varying concentrations of hydrogen peroxide (H2O2) (Materials and Methods). The ΔhpdA and ΔhmgA strains showed increased sensitivity to H2O2 compared to the wildtype, ΔhpdA hpdA+ and ΔhmgA hmgA+ complemented strains (Fig. 8). The wA gene, which encodes the polyketide synthase required for DHN melanin production in conidia, is required for resistance to H2O2 in A. fumigatus [15, 20]. However, only a very mild sensitivity to H2O2 was observed in the P. marneffei ΔwA strain and this was much lower than the ΔhpdA and ΔhmgA strains (Fig. 8).
Fig. 8. Pyomelanin is required for resistance to oxidative stress in P. marneffei.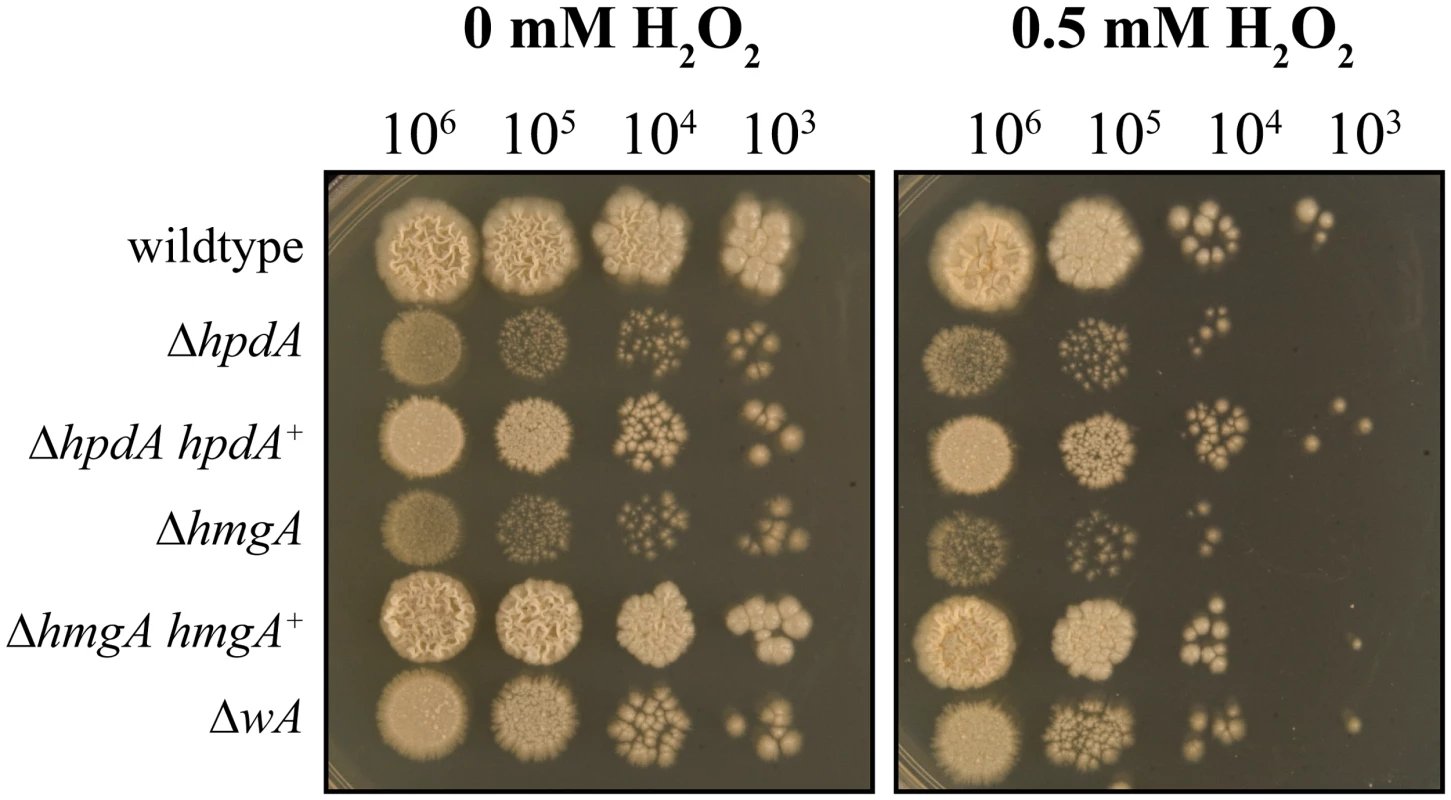 
Serial dilutions of conidial suspensions of the wildtype, ΔhpdA, ΔhpdA hpdA+, ΔhmgA, ΔhmgA hmgA+ and ΔwA strains dropped onto medium containing tyrosine and 0 or 0.5 mM H2O2 and incubated for 6 days at 37°C. Unlike the other genes of the tyrosine catabolic cluster hpdA is required for infectious growth in macrophages
The addition of the HpdA inhibitor NTBC prevented P. marneffei yeast cell production during macrophage infection suggesting hpdA is required for ex vivo growth (Fig. 1). To confirm the required for hpdA during ex vivo growth and to assess if the other genes of the tyrosine catabolic cluster are also required for ex vivo growth, conidia of the wildtype, ΔwA, ΔhpdA, ΔhmgA, ΔhypW, ΔhmgX, ΔmaiA, ΔmfpA and ΔhmgR strains were used to infect murine J774 macrophages and examined 24 hours post-infection. A control of wildtype conidia incubated in macrophage medium alone was also performed. After 24 hours, macrophages infected with wildtype conidia contain numerous yeast cells dividing by fission (Fig. 9A). In contrast, only 2.67 ± 0.70% of conidia had germinated in the macrophage medium control. This suggests that any growth of P. marneffei observed within macrophages is due to the derivation of nutrients from the host macrophage. In contrast to wildtype and consistent with the NTBC experiments, ungerminated conidia were predominately observed in macrophages infected with conidia of the ΔhpdA mutant 24 hours post-infection (Fig. 9A and B). In macrophages infected with the ΔhpdA strain, 82.2±6.78% of conidia remained ungerminated and only 17.4±6.57% germinated to produce yeast cells (compared to 10.6±4.65% ungerminated conidia and 89.5±4.65% yeast cells for wildtype) (Fig. 9B). This phenotype was complemented by reintroduction of hpdA+ in the ΔhpdA hpdA+ strain (10.9±0.41% ungerminated conidia and 88.4±0.66 yeast cells). No differences in germination rates were observed between the wildtype and ΔhpdA strain in vitro. Despite the ΔhpdA and ΔhmgX mutants having indistinguishable growth and pigmentation phenotypes on tyrosine, the ΔhmgX strain produced normal numbers of yeast cells ex vivo (Fig. 9A and B). The ΔhmgA strain showed a small increase in the number of ungerminated conidia (29.0±5.78%) and consequently a small decrease in the number of yeast cells (69.2±5.72%) produced at 24 hours post-infection (Fig. 9A and B). This phenotype was complemented by reintroduction of hmgA+ in the ΔhmgA hmgA+ strain (10.4±1.75% ungerminated conidia and 89.6±1.75% yeast). The production of yeast cells in macrophages infected with ΔwA, ΔhypW, ΔhmgX, ΔmaiA, ΔmfpA and ΔhmgR conidia was indistinguishable from wildtype, suggesting that tyrosine and phenylalanine catabolism as a source of carbon is not required for the initial stages of infection (Fig. 9B). Interestingly, the ΔwA strain produced numerous yeast cells at levels equivalent to wildtype after 24 hours infection (85.2±4.97% compared to 89.5±4.65% for wildtype) despite a previous report that suggested decreased expression of this gene using RNAi results in a decrease in virulence in a mouse model of infection (Fig. 9B) [16].
Fig. 9. ex vivo growth in macrophages. 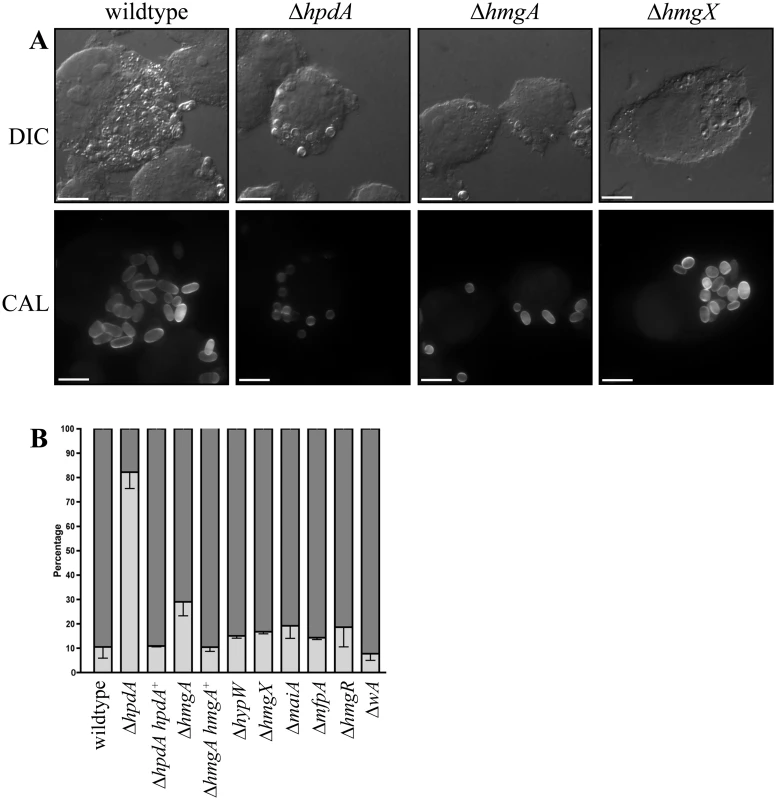
Conidia of the wildtype, ΔhpdA, ΔhpdA hpdA+, ΔhmgA, ΔhmgA hmgA+, ΔhypW, ΔhmgX, ΔmaiA, ΔmfpA, ΔhmgR and ΔwA strains were used to infect J774 murine macrophages and growth was assessed 24 hours post-infection. A. After 24 hours, macrophages infected with wildtype conidia contain numerous yeast cells dividing by fission. In contrast, ungerminated conidia were predominately observed in macrophages infected with conidia of the ΔhpdA mutant 24 hours post-infection. The ΔhmgA strain showed a small increase in the number of ungerminated conidia and a small decrease in the number of yeast cells. The ΔhmgX mutant was indistinguishable from wildtype. B. Quantitation of the percentage of ungerminated conidia and yeast cells in macrophages infected with wildtype, ΔhpdA, ΔhpdA hpdA+, ΔhmgA, ΔhmgA hmgA+, ΔhypW, ΔhmgX, ΔmaiA, ΔmfpA, ΔhmgR and ΔwA 24 hours post-infection. Dark grey indicates the percentage of yeast cells, whereas, light grey represents the percentage of ungerminated conidia. To observe if ΔhpdA conidia germinate with a longer period of incubation, conidia of the wildtype, ΔhpdA and ΔhpdA hpdA+ strains were used to infect murine J774 macrophages and examined 48 hours post-infection. After 48 hours, macrophages infected with wildtype and ΔhpdA hpdA+ conidia contain prolific numbers of yeast cells dividing by fission (wildtype 1.43±0.98% ungerminated conidia, 98.6±0.98% yeast cells and ΔhpdA hpdA+ 0.75±0.48 ungerminated conidia and 99.3±0.48% yeast cells). Conidia of the ΔhpdA strain have begun to germinate into yeast cells by 48 hours but show reduced cellular proliferation (38.5±5.71% ungerminated conidia, 61.3±5.63% yeast). Consistent with this observation, ΔhpdA conidia remain viable in macrophages. Plating the contents of lysed ΔhpdA infected macrophages at 37°C in vitro resulted in the growth of ΔhpdA colonies.
To assess if deletion of genes of the tyrosine catabolic cluster affects the phagocytosis of conidia by macrophages, conidia of the wildtype, ΔhpdA, ΔhmgA, ΔhmgX and ΔmaiA strains were used to infect murine J774 macrophages and the macrophages examined 2 hours post-infection. No differences in phagocytosis were observed (average number of conidia per 100 macrophages: wildtype 76.7±4.70, ΔhpdA 67.3±9.80, ΔhmgA 70.0±9.90, ΔhmgX 75.0±6.70 and ΔmaiA 79.3±8.20).
The wildtype, ΔhpdA, ΔhmgA and ΔhmgX strains were also used to infect human THP-1 macrophages. The ΔhpdA, ΔhmgA and ΔhmgX phenotypes in human macrophages were identical to those described for infection of murine macrophages (S6 Fig).
Discussion
The tyrosine catabolic cluster is subject to complex regulatory control
The expression of genes within the P. marneffei tyrosine catabolic gene cluster is subject to a complex network of regulatory control with four different levels of regulation; pathway specific regulation by the Zn(II)2-Cys6 binuclear transcription factor HmgR, cluster specific regulation by location, global regulation in response to nitrogen source by AreA and post-transcriptional feedback by pathway intermediates. The complex regulation of the tyrosine catabolism gene cluster reflects the multiple roles played by genes in this cluster, not only in adapting to the nutritional sources available but also the production of protective melanin. An additional, unique role for HpdA in P. marneffei, and probably other dimorphic fungi with an intracellular (host cell) phase, is also suggested.
Genes of the tyrosine catabolic cluster are regulated specifically in response to the presence of tyrosine by the Zn(II)2-Cys6 binuclear transcription factor HmgR encoded within the cluster. HmgR is required for full induction of these genes in the presence of tyrosine as the sole nitrogen source and deletion results in reduced and no growth on tyrosine as a sole nitrogen and carbon source, respectively. The orthologue in A. fumigatus, hmgR, is also required for tyrosine-induced expression of genes in the cluster, suggesting this protein plays a conserved role [5]. However, unlike A. fumigatus, the P. marneffei ΔhmgR mutation results in only a partial loss of tyrosine-induced expression, suggesting that an additional factor contributes to positively regulating expression of the tyrosine catabolic cluster in response to tyrosine. Zn(II)2-Cys6 binuclear transcription factors have been shown to positively regulate the expression of many nitrogen metabolic pathways in fungi [5, 34, 35, 36, 37]. Unexpectedly, deletion of hmgR in P. marneffei also resulted in partial derepression on ammonium suggesting that HmgR also has a negative role in regulating gene expression.
The genes of the tyrosine catabolic cluster are also under specific regulation by location. The expression of mfpA, the gene within the tyrosine metabolic cluster only in P. marneffei and T. stipitatus which does not appear to have a role in tyrosine catabolism, was induced on tyrosine at both 25°C and 37°C even though it does not share a promoter with another cluster gene. This suggests that any gene captured within this genomic region may consequently come under its regulation. Similar effects have been described for other gene clusters (for example spoCI in A. nidulans) [38]. In addition, the complementation strains which had the wildtype gene reintroduced at the pyrG or niaD loci, did not show complete restoration of wildtype growth on tyrosine and phenylalanine, also indicating that the position of the gene in the cluster is important for it’s regulation.
AreA is a positively-acting GATA-type transcription factor which regulates a large number of genes to effect nitrogen metabolite repression; the global response to limiting nitrogen conditions [30, 39]. We originally hypothesized that the additional factor contributing to positively regulating expression of the tyrosine catabolic cluster in response to tyrosine could be AreA. In A. nidulans, AreA remodels chromatin in cooperation with Zn(II)2-Cys6 binuclear transcription factors in response to specific nitrogen sources [36, 40]. However, deletion of areA in P. marneffei did not reduce induction of the tyrosine catabolic cluster genes in the presence of tyrosine but rather unexpectedly lead to a loss of repression in the presence of ammonium. This suggests that contrary to the paradigm, AreA is acting negatively on the tyrosine catabolic cluster genes under nitrogen repressing conditions. To our knowledge, only two examples exist of AreA acting as a negative regulator in fungi despite examples of a related human GATA factor acting as both an activator and repressor [41, 42, 43]. In A. nidulans, AreA negatively regulates the expression of nadA, an adenine deaminase encoding gene required for purine degradation, though unlike the P. marneffei tyrosine cluster genes, nadA is also expressed on ammonium [41]. There is only 4 bp between the unique UaY (positively acting Zn(II)2-Cys6 binuclear transcription factor) and AreA binding sites in the nadA promoter and AreA acts in part to negate UaY-mediated induction by directly competing with UaY binding [41]. However, the identification of potential AreA binding sites in the promoters of P. marneffei tyrosine catabolic cluster genes (HGATAR) shows that no predicted sites are in close proximity to the potential HmgR binding site predicted by RSAT analysis. AreA also acts as a repressor of genes required for arginine catabolism in A. nidulans [43]. Similar to the P. marneffei tyrosine gene cluster, genes required for arginine catabolism in A. nidulans are not expressed in ammonium, are induced by arginine and this induction is dependent on the ArcA Zn(II)2-Cys6 binuclear transcription factor [35]. Also similar to tyrosine, arginine can be utilized as both a nitrogen and carbon source. Under carbon repressing conditions (1% glucose) an areA loss of function mutant results in a loss of nitrogen metabolite repression of the arginine catabolism genes [43].
Tyrosine catabolism is also regulated by post-transcriptional feedback by pathway intermediates. Deletion of genes of the tyrosine catabolic cluster (hpdA, hmgA, hmgX and maiA) resulted in reduced growth on tyrosine as a sole nitrogen source. The proposed transamination reaction of tyrosine with α-ketoglutarate to produce glutamate as a nitrogen source occurs prior to the reactions catalysed by HpdA, HmgA and MaiA. This is indicative of a negative feedback loop regulating the catabolism of tyrosine in P. marneffei. This regulatory mechanism appears to be conserved as the hmgR and hmgX deletions in A. fumigatus also show reduced growth on tyrosine or phenylalanine as the sole nitrogen source [5].
The role of tyrosine catabolism and pyomelanin production during pathogenic growth of yeast cells
Dimorphism is intricately linked to pathogenicity and therefore differentially expressed genes, particularly those that display yeast-specific expression, are likely to play a role in the establishment and progression of infection. Based on this hypothesis, yeast-specific genes in P. marneffei were identified using a microarray-based expression profiling [6]. This profiling revealed that hpdA, encoding the enzyme 4-hydroxyphenylpyruvate dioxygenase (HpdA) required during the catabolism of tyrosine, was induced specifically in the pathogenic yeast cell type at 37°C [6]. Intriguingly, chemical inhibition of HpdA activity using NTBC or deletion of hpdA resulted in a severe defect in the production of yeast cells during macrophage infection, suggesting that HpdA is required for P. marneffei pathogenicity. Therefore the question arose as to whether this was a result of reduced nutrient acquisition, reduced melanin protection against oxidative stress or a combination of both. Interestingly, chemical inhibition of HpdA resulted in more significant delay in conidial germination compared to deletion of hpdA (Figs. 1 and 9). This is likely due to the inhibition of the host 4-hydroxyphenylpyruvate dioxygenase, as well as that of the fungus, thus leading to a further decrease in available nutrients for growth.
Deletion of hpdA, as well as other cluster genes encoding enzymes required for tyrosine catabolism (hmgA, hmgX and maiA) and the gene encoding the regulatory transcription factor (hmgR) resulted in reduced growth on tyrosine as the sole nitrogen or carbon source at both 25°C and 37°C, showing that these genes are required for tyrosine catabolism and that P. marneffei can utilize tyrosine as both a nitrogen and carbon source for growth. However, unlike ΔhpdA, the ΔhmgX, ΔmaiA and ΔhmgR strains did not show reduced yeast cell production in macrophages. The ΔhmgA mutant showed slightly reduced yeast cell production but this is likely to be attributed to reduced conidial germination due accumulation of homogentisate and consequent hypermelanization. This result suggests that nitrogen or carbon acquisition via tyrosine catabolism is not essential for the early stages of infection in macrophages and that the phenotype of the ΔhpdA mutant is not due to reduced nutrient acquisition. In addition, the lack of conidial germination in the ΔhpdA in macrophages is unlikely to be due to a toxic build-up of tyrosine catabolic intermediates, as the ΔhmgA mutant, which showed a greater reduction in growth than the ΔhpdA mutant due to toxicity in vitro, displayed only a mild reduction in yeast cell production ex vivo. Somewhat unexpectedly, ΔhmgX did not show the same phenotype as ΔhpdA during macrophage infection, despite the hypothesis that HmgX acts as an accessory protein to HpdA and the ΔhpdA and ΔhmgX strains showing an equivalent growth reduction on tyrosine as a sole nitrogen or carbon source at both 25°C and 37°C and an equivalent lack of pyomelanin production in vitro. It is possible that HmgX is only partially required for HpdA function under certain conditions. This hypothesis is supported by the reduced accumulation of toxic intermediates of the tyrosine catabolic pathway compared to ΔhpdA and ΔhmgA.
The ex vivo phenotype of the ΔhpdA mutant is unlikely to be a result of the inability to produce pyomelanin, as the ΔhmgX mutant, which like the ΔhpdA mutant cannot produce pyomelanin, produces wildtype levels of yeast cells in macrophages. It is possible that the ΔhmgX mutant can produce a colourless intermediate to the final pigmented pyomelanin which may be sufficient to afford protection in macrophages, however, the ΔmaiA and ΔhmgR mutants, which displayed reduced pyomelanin production at 37°C, also displayed ex vivo growth indistinguishable from wildtype. Instead these results may suggest that HpdA has an additional role to tyrosine catabolism and pyomelanin formation in P. marneffei and may also explain why hpdA is highly constitutively expressed at 37°C, in contrast to all of the other genes in the tyrosine catabolic cluster. Therefore the question arises as to what this additional novel role of hpdA during ex vivo growth could be. It is possible that tyrosine catabolism, and specifically the step catalysed by HpdA, is acting as a developmental signal. The enzyme tyrosine aminotransferase, which catalyses the first step in tyrosine catabolism and which an orthologue is lacking in P. marneffei, has been shown to promote apoptosis and prevent cell proliferation during cancer development in humans [44]. In addition, inhibition of hpd-1 by RNAi in the nematode Caenorhabditis elegans extends lifespan and results in a developmental arrest at the dauer larval stage, which prevents the completion of development when environmental conditions are unfavourable [45]. In C. elegans the link between tyrosine catabolism and development is proposed to occur via the insulin-like daf-2 signalling pathway [46]. Elevated tyrosine has inhibitory affects on DAF-2 signalling [47]. The DAF-2 receptor, orthologous to the insulin receptor in mammals, lies upstream of a phosphatidylinositol 3 signalling cascade which acts to phosphorylate the DAF-16 forkhead transcription factor preventing it’s entry into the nucleus [47]. Daf-16 positively regulates the expression of a number of genes regulating metabolism and lifespan including those encoding enzymes which protect against free radicals such as catalase-1 (ctl-1) and superoxide dismutase (sod-3) and 4-hydroxyphenylpyruvate dioxygenase (hpd-1) [45, 47]. The P. marneffei genome encodes a daf-16 homologue (PMAA_055700) and the hpdA promoter contains a putative DAF-16 binding site (ATGTTTGA) which is conserved in the Aspergilli (consensus WTGTTTVV), suggesting that this regulatory mechanism may be conserved in fungi.
Further elucidation of the role played by hpdA during ex vivo growth will clearly provide insight into the progression of conidia into yeast cells during intracellular pathogenic growth of P. marneffei and opens up an exciting new avenue of investigation.
Materials and Methods
Molecular techniques and plasmid construction
P. marneffei genomic DNA was isolated as previously described [48]. Southern and northern blotting was performed with Amersham Hybond N+ membrane using [a-32P]dATP labeled probe hybridization using standard methods [49].
Sequences of primers are provided in S1 Table. A PCR product encompassing P. marneffei hpdA (PMAA_031950) was generated with primers AA48 and AA49 and cloned into pGemT Easy to generate pAM7072. To make the deletion construct, pKB7717, an EcoRI/SalI 5’ PCR product generated with MM70 and MM71 was cloned into EcoRI/SalI pBluescript II SK+. A BamHI/SpeI 3’ PCR product generated with MM72 and MM73 was then cloned into the BamHI/SpeI sites and subsequently a BamHI/EcoRI pyrGb fragment from pAB4626 was cloned into the BamHI/EcoRI sites. The complementation construct, pKB7682, was generated by cloning a HindIII/SacI fragment from pAM7072 into pLS7804 (pyrG targeting pBluescript II SK+). hmgA (PMAA_031960) was PCR amplified using primers MM75 and MM76 and cloned into pGemT Easy to generate pKB7723. The deletion construct, pKB7724, was generated by replacing the BglII/StuI fragment from pKB7723 with BamHI/EcoRV pyrGb from pAB4626. The complementation construct, pKB7791, was generated cloning a BamHI/XhoI digested PCR product generated using OO78 and OO79 into BamHI/XhoI pLS7804. maiA (PMAA_032000) was cloned by PCR amplification with primers PP18 and PP19 and cloned into pGemT Easy to generate pKB7786. The deletion construct, pKB7805, was generated by replacing the BglII/XhoI fragment from pKB7786 with a BamHI/XhoI fragment from pAB4626. The maiA complementation construct, pKB858, was generated by cloning a PstI/EcoRV fragment from pKB7786 into PstII/EcoICRI pHB7615 (niaD targeting pBluescript SK+). hypW (PMAA_031870) was cloned by PCR amplification with primers KK37 and KK38 and cloned into pGemT Easy to generate pSM7504. The deletion construct, pSM7546, was generated by ligating an inverse PCR product generated using primers KK71 and KK72 to EcoRV/SmaI pyrGb from pAB4626. Likewise, hmgX (PMAA_031980) was cloned by PCR amplification with primers KK39 and KK40 and cloned into pGemT Easy to generate pSM7505. The deletion construct, pSM7651, was generated by ligating an inverse PCR product generated using primers KK73 and KK74 to EcoRV/SmaI pyrGb from pAB4626. The hypW and hmgX complementation construct, pKB7801, was generated by cloning a SpeI/StuI fragment from pSM7505 into XbaI/EcoICRI pLS7804. The complementation construct containing hypW and a truncated hmgX allele, pKB7940, was generated by cloning a PstI/HindIII fragment from pSM7505 into PstI/HindIII pHB7615. The regulatory gene hmgR (PMAA_032020) was cloned by ligating the PCR product generated using primers LL13 and LL14 into EcoRV digested pBluescript II SK+, to generate pSH7541. The deletion construct, pSH7578, was generated by cloning an EcoRV/SmaI pyrGb fragment from pAB4626 into an inverse PCR product of pSH7541 with primers LL56 and LL57. The complementation construct, pKB7683, was generated by cloning in the KpnI/SpeI fragment from pSH7541 into KpnI/XbaI pLS7804. mfpA (PMAA_032010) was PCR amplified using primers QQ50 and QQ52 and cloned into pGemT Easy to generate pLS7828. The deletion construct, pLS7852, was generated using a Gateway method as described in [50] using the primers QQ57 and QQ58. hpdB (PMAA_089170) was amplified using primers PP14 and PP15 and cloned into pGemT Easy to generate pLS7785. The deletion construct, pLS7845, was generated using the Gateway method using the primers QQ59 and QQ60. wA (PMAA_082120) was amplified using primers HH45 and HH46 and cloned into pGemT Easy to generate pSJ7351. The deletion construct, pHW7445, was generated by replacing the BglII/EcoRV fragment of pSJ7351 with the BamHI/EcoRV pyrGb fragment from pAB4626.
Fungal strains and media
Transformation was performed using the previously described protoplast method [48]. Strains used in this study are listed in Table 2. The ΔhpdA pyrG+ strain (G827) was generated by transformation of strain G832 (pkuA TK barA niaD1 pyrG1) with linearised pKB7717 which removes sequences from -11 to +1517 of hpdA and selecting for pyrG+. The ΔhmgA::pyrG+ (G823), ΔhypW::pyrG+ (G854), ΔhmgX::pyrG+ (G856), ΔmaiA::pyrG+ (G895), ΔmfpA::pyrG+ (G946) and ΔhpdB strains were generated by transforming strain G816 (ΔligD niaD1 pyrG1) with linearised deletion constructs pKB7724 (which removes from +138 to +717 of hmgA), pSM7546 (which removes from +5 to +1145 of hypW), pSM7651 (which removes from -53 to +929 of hmgX), pKB7805 (which removes from +42 to +249 of maiA) and pLS7852 (removes from -22 to +2741 of mfpA), pLS7845 (which removes from -39 to +1127 of hpdB) and pSH7578 (which removes from -278 to +2601 of hmgR), and selecting for pyrG+ or glufosinate resistance. The ΔwA::pyrG+ (G748) strain was generated by transforming strain G147 (niaD1 pyrG1) with pHW7445 and selecting for pyrG+. Deletion was confirmed by genomic Southern blot analysis. pyrG- strains (Table 2) were generated by plating on medium containing 1 mg mL-1 5-fluoroorotic acid (5-FOA) supplemented with 10 mM NH4SO4 and 5 mM uracil to select for the loss of the pyrG marker. These strains are unable to grow in the absence of 5 mM uracil. The deletion mutants were complemented by transformation of pyrG- strains with pKB7682 (hpdA), pKB7791 (hmgA), pKB7800 (maiA), pKB7801 (hypW+ hmgX+), pKB7940 (hypW+ hmgXtrun), pKB7801 (hmgX), pKB7683 (hmgR) and selecting for pyrG+ or transformation of G895 with pKB7858 (maiA) and selecting for niaD+. Integration of the complementation constructs at pyrG or niaD was confirmed by Southern blot analysis of genomic DNA.
Tab. 2. Strains used in this study. 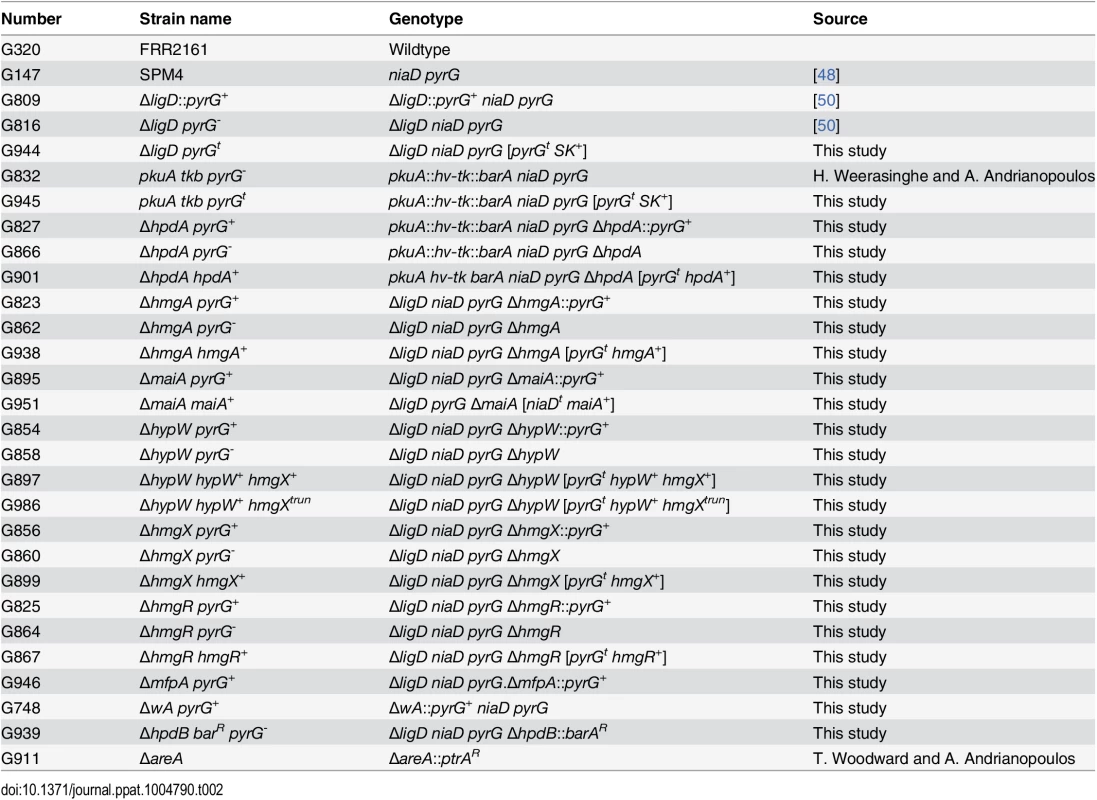
To test growth on tyrosine or phenylalanine as a sole nitrogen source, strains were grown at 25°C and 37°C on A. nidulans minimal medium (ANM) supplemented with 1% glucose and 10 mM ammonium sulphate ((NH4)2SO4), 10 mM phenylalanine, or 10 mM tyrosine [51,52]. To test growth on tyrosine or phenylalanine as a sole carbon source, strains were grown at 25°C and 37°C for 14 days on A. nidulans minimal medium (ANM) supplemented with 10 mM ammonium sulphate ((NH4)2SO4) and 50 mM phenylalanine or 10 mM tyrosine [51,52]. NTBC (2-(2-nitro-4-trifluoromethylbenzoyl)-cyclohexane-1, 3-dione) was added for HpdA inhibition experiments at a final concentration of 400μg mL-1. To assess pyomelanin production, strains were grown at 37°C for 14 days on 1% glucose A. nidulans minimal medium (ANM) supplemented with either 10 mM ammonium sulphate (NH4SO4), 10 mM tyrosine, both 10 mM NH4SO4 and 10 mM tyrosine or both 10 mM tyrosine and 10 mM alanine. Strains were also grown at 37°C on BHI for 5 days. The growth of the ΔhmgR strain was assessed after 14 days on 10 mM alanine, arginine, asparagine, cysteine, glutamate, glycine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, proline, serine, threonine, tryptophan, tyrosine or valine as the sole nitrogen source at both 25°C and 37°C. To test the effect of accumulating toxic tyrosine catabolism intermediates on growth, the wildtype (WT) and ΔhpdA, ΔhmgA and ΔhmgX strains were grown on carbon-free medium containing 10mM Gaba and either 10mM sorbitol, 1% lactose, 10mM NaAc or 10mM proline with or without 10mM tyrosine for 14 days at 25°C.
L-DOPA medium was prepared by making a 50 mL dH20 solution containing 0.5 g L-asparagine, 0.5 g glucose, 1.5 g KH2PO4, 0.125 g MgSO4-7H20 and 100 mg L-DOPA and adjusting the pH to 5.6. Subsequently, 0.5 mg thiamine-HCL and 0.5 mL of 1M biotin solution were added and the solution filter sterilized. This solution was added to 500 mL of sterilized 2% agar solution. Strains were grown at 37°C for 14 days.
Oxidative stress was tested by plating on ANM + Supps medium supplemented with 10 mM tyrosine and 0 mM, 0.25 mM, 0.5 mM, 0.75 mM and 1 mM of H2O2. Plates were inoculated with 10 μL drops of 10 fold serial dilutions of a 1x 107 conidia mL-1 suspension and were incubated at 37°C for 14 days.
Purification of melanin particles
The wildtype and ΔhpdA mutant were grown on ANM + Supps medium supplemented with 10 mM tyrosine for 14 days at 37°C. Cells were suspended in 0.1M sodium citrate and 1M sorbitol, pH 5.5 plus 10mg/mL lytic enzyme overnight at 30°C. Cells were pelleted and resuspended in 4M guanidine thiocyanate solution and incubated at room temperature overnight. Cells were then washed in PBS, resuspended in 6.6M HCL and boiled for 1.5 hours. The remaining particles (none were visible for the ΔhpdA mutant) were centrifuged and the pellets washed 3x in PBS and resuspended (in PBS).
Macrophage assay
J774 murine macrophages (1 x 105) or THP-1 human macrophages were seeded into each well of a 6 well microtitre tray containing one sterile coverslip and 2 mL of complete Dulbecco’s Modified Eagle Medium for J774 (complete DMEM: DMEM, 10% fetal bovine serum, 8 mM L-glutamine and penicillin-streptomycin) or 2 mL of RPMI (complete RPMI: RPMI, 10% fetal bovine serum, 2 mM L-glutamine and penicillin-streptomycin) for THP-1. Macrophages were incubated at 37°C for 24 hours before activation with 0.1 μg mL-1 lipopolysaccharide (LPS) from E. coli (Sigma) and THP-1 macrophages differentiated with phorbol 12-myristate 13-acetate. Macrophages were incubated a further 24 hours at 37°C, washed in phosphate buffered saline (PBS) and 2 mL of complete DMEM or RPMI medium containing 1 x 106 conidia was added. A control lacking conidia was also performed. Macrophages were incubated for 2 hours at 37°C (to allow conidia to be engulfed), washed once in PBS (to remove free conidia) and incubated a further 24 or 48 hours at 37°C. Macrophages were fixed in 4% paraformaldehyde and stained with 1 mg mL-1 fluorescent brightener 28 (calcofluor—CAL) to observe fungal cell walls. Mounted coverslips were examined using differential interference contrast (DIC) and epifluorescence optics for cell wall staining and viewed on a Reichart Jung Polyvar II microscope. Images were captured using a SPOT CCD camera (Diagnostic Instruments Inc) and processed in Adobe PhotoshopTM. The numbers of ungerminated conidia, germlings or yeast cells were recorded in a population of approximately 100 in three independent experiments. Mean and standard error of the mean values were calculated using GraphPad Prism3. 400μg mL-1 of NTBC was added at the time of infection for HpdA inhibition experiments.
Expression analysis
Liquid cultures of FRR2161 (wildtype), ΔhmgR and 2206areA strains grown for 2 days at 25°C in ANM plus 10 mM (NH4)2SO4 and 6 days at 37°C in BHI were used to inoculate ANM plus 10 mM (NH4)2SO4, 10 mM alanine or, 10 mM tyrosine as the sole nitrogen source at 25°C and 37°C and RNA was isolated after 4 hours. Liquid cultures of FRR2161 (wildtype), ΔhmgR and ΔareA strains grown for 2 days at 25°C in ANM plus 10 mM (NH4)2SO4 were used to inoculate ANM plus 10 mM tyrosine or 10 mM tyrosine and 10 mM (NH4)2SO4 at 25°C and RNA was isolated after 4 hours. 2 day 25°C liquid cultures of ΔhypW grown in ANM with 10 mM (NH4)2SO4, was used to inoculate ANM with 10 mM (NH4)2SO4 or 10 mM tyrosine as the sole nitrogen source at 25°C and RNA was isolated after 4 hours. RNA was extracted using TRIzol Reagent (Invitrogen) and a MP FastPrep-24 bead beater according to the manufacturer’s instructions. At least two biological repeats were performed. Illumina RNA sequencing was performed on RNA isolated from liquid cultures of FRR2161 (wildtype) grown for 6 days at 37°C in BHI. RNA was DNAase treated (Promega) prior to RT PCR analysis. For every gene in this study, 3 increasing cycle numbers were used on two biological repeats to ensure the product was not saturated and the result was representative. H3 (histone H3) was used as a loading control. Expression was determined by RT PCR using primers CC53 and CC54 (hpdA), DD13 and DD14 (hmgA), MM64 and MM65 (hmgX), MM32 and MM33 (hypW), DD19 and DD20 (fahA), DD17 and DD18 (maiA), II94 and II95 (mfpA), II88 and II89 (hmgR), and GG4 and GG5 (H3).
Supporting Information
Zdroje
1. Lorenz MC, Fink GR (2002) Life and death in a macrophage: role of the glyoxylate cycle in virulence. Eukaryot Cell 1 : 657–662. 12455685
2. Nunes LR, Costa de Oliveira R, Leite DB, Schmidt da Silva V, doa Reis Marques E, et al. (2005) Transcriptome analysis of Paracoccidioides brasiliensis cells undergoing mycelium-to-yeast transition. Eukaryot Cell 4 : 2115–2128. 16339729
3. Fan W, Kraus PR, Boily MJ, Heitman J (2005) Cryptococcus neoformans gene expression during murine macrophage infection. Eukaryot Cell 4 : 1420–1433. 16087747
4. Hwang L, Hocking-Maurray D, Bahramai AK, Andersson M, Rine J, et al. (2003) Identifying phase-specific genes in the fungal pathogen Histoplasma capsulatum using a genomic shotgun microarray. Mol Biol Cell 14 : 2314–2326. 12808032
5. Keller S, Macheleidt J, Scherlach K, Schmaler-Ripcke J, Jacobsen ID, et al. (2011) Pyomelanin formation in Aspergillus fumigatus requires HmgX and the transcriptional activator HmgR but is dispensable for virulence. PLoS One 6: e26604. doi: 10.1371/journal.pone.0026604 22046314
6. Pasricha S, Payne M, Canovas D, Pase L, Ngaosuwankul N, et al. (2013) Cell-type-specific transcriptional profiles of the dimorphic pathogen Penicillium marneffei reflect distinct reproductive, morphological, and environmental demands. G3 (Bethesda) 3 : 1997–2014.
7. Fernandez-Canon JM, Penalva MA (1995) Fungal metabolic model for human type I hereditary tyrosinaemia. PNAS 92 : 9132–9136. 7568087
8. Penalva MA (2001) A fungal perspective on human inborn errors of metabolism: alkaptonuria and beyond. Fungal Genet Biol 34 : 1–10. 11567547
9. Langfelder K, Streibel M, Jahn B, Haase G, Brakhage AA (2003) Biosynthesis of fungal melanins and their importance for human pathogenic fungi. Fungal Genet Biol 38 : 143–158. 12620252
10. Fernandez-Canon JM, Penalva MA (1995) Molecular characterization of a gene encoding a homogentisate dioxygenase from Aspergillus nidulans and identification of its human and plant homologues. J Biol Chem 270 : 21199–21205. 7673153
11. Schmaler-Ripcke J, Sugareva V, Gebhardt P, Winkler R, Kniemeyer O, et al. (2009) Production of pyomelanin, a second type of melanin, via the tyrosine degradation pathway in Aspergillus fumigatus. Appl Environ Microbiol 75 : 493–503. doi: 10.1128/AEM.02077-08 19028908
12. Kwon-Chung KJ, Polacheck I, Popkin TJ (1982) Melanin-lacking mutants of Cryptococcus neoformans and their virulence for mice. J Bact 150 : 1414–1421. 6804444
13. Wang Y, Casadevall A (1994) Susceptibility of melanized and nonmelanized Cryptococcus neoformans to nitrogen - and oxygen - derived oxidants. Infect Immun 62 : 3004–3007. 8005689
14. Dixon DM, Polak A, Szaniszlo PJ (1987) Pathogenicity and virulence of wild-type and melanin-deficient Wangiella dermatitidis. J Med Vet Mycol 25 : 97–106. 3598824
15. Jahn B, Koch A, Schmidt A, Wanner G, Gehringer H, et al. (1997) Isolation and characterization of a pigmentless-conidium mutant of Aspergillus fumigatus with altered connidial surface and reduced virulence. Infect Immun 65 : 5110–5117. 9393803
16. Woo PC, Tam EW, Chong KT, Cai JJ, Tung ET, et al. (2010) High diversity of polyketide synthase genes and the melanin biosynthesis gene cluster in Penicillium marneffei. Febs J 277 : 3750–3758. doi: 10.1111/j.1742-4658.2010.07776.x 20718860
17. Mednick AJ, Nosanchuk JD, Casadevall A (2005) Melanization of Cryptococcus neoformans affects lung inflammatory responses during cryptococcal infection. Infect Immun 73 : 2012–2019. 15784542
18. da Silva MB, Marques AF, Nosanchuk JD, Casadevall A, Travassos LR, et al. (2006) Melanin in the dimorphic fungal pathogen Paracoccidioides brasiliensis: effects on phagocytosis, intracellular resistance and drug susceptibility. Microbes Infect 8 : 197–205. 16213179
19. Romero-Martinez R, Wheeler M, Guerrero-Plata A, Rico G, Torres-Guerrero H (2000) Biosynthesis and functions of melanin in Sporothrix schenckii. Infect Immun 68 : 3696–3703. 10816530
20. Jahn B, Boukhallouk F, Lotz J, Langfelder K, Wanner G, et al. (2000) Interaction of human phagocytes with pigmentless Aspergillus conidia. Infect Immun 68 : 3736–3739. 10816538
21. Jahn B, Langfelder K, Schneider U, Schindel C, Brakhage AA (2002) PKSP-dependent reduction of phagolysosome fusion and intracellular kill of Aspergillus fumigatus conidia by human monocyte-derived macrophages. Cell Microbiol 4 : 793–803. 12464010
22. Huffnagle GB, Chen GH, Curtis JL, McDonald RA, Strieter RM, et al. (1995) Down-regulation of the afferent phase of T cell-mediated pulmonary inflammation and immunity by a high melanin-producing strain of Cryptococcus neoformans. J Immunol 155 : 3507–3516. 7561046
23. Youngchim S, Hay RJ, Hamilton AJ (2005) Melanization of Penicillium marneffei in vitro and in vivo. Micrbiology 151 : 291–299. 15632446
24. Taborda CP, da Silva MB, Nosanchuk JD, Travassos LR (2008) Melanin as a virulence factor of Paracoccidioides brasiliensis and other dimorphic pathogenic fungi: a minireview. Mycopathologia 165 : 331–339. 18777637
25. Gomez BL, Nosanchuk JD, Diez S, Youngchim S, Aisen P, et al. (2001) Detection of melanin-like pigments in the dimorphic fungal pathogen Paracoccidioides brasiliensis in vitro and during infection. Infect Immun 69 : 5760–5767. 11500453
26. Nosanchuk JD, Gomez BL, Youngchim S, Diez S, Aisen P, et al. (2002) Histoplasma capsulatum synthesizes melanin-like pigments in vitro and during mammalian infection. Infect Immun 70 : 5124–5131. 12183562
27. Keon J, Hargreaves J (1998) Isolation and heterologous expression of a gene encoding 4-hydroxyphenylpyruvate dioxygenase from the wheat leaf-spot pathogen, Mycosphaerella graminicola. FEMS Microbiol Lett 161 : 337–343. 9570125
28. Lock EA, Ellis MK, Gaskin P, Robinson M, Auton TR, et al. (1998) From toxicological problem to therapeutic use: the discovery of the mode of action of 2-(2-nitro-4-trifluoromethylbenzoyl)-1,3-cyclohexanedione (NTBC), its toxicology and development as a drug. J Inherit Metab Dis 21 : 498–506. 9728330
29. Marzluf GA (1997) Genetic regulation of nitrogen metabolism in the fungi. Microbiol Mol Biol Rev 61 : 17–32. 9106362
30. Bugeja HE, Hynes MJ, Andrianopoulos A (2012) AreA controls nitrogen source utilisation during both growth programs of the dimorphic fungus Penicillium marneffei. Fungal Biol 116 : 145–154. doi: 10.1016/j.funbio.2011.10.009 22208609
31. Rodriguez-Rojas A, Mena A, Martin S, Borrell N, Oliver A, et al. (2009) Inactivation of the hmgA gene of Pseudomonas aeruginosa leads to pyomelanin hyperproduction, stress resistance and increased persistence in chronic lung infection. Microbiology 155 : 1050–1057. doi: 10.1099/mic.0.024745-0 19332807
32. da Silva Ferreira ME, Savoldi M, Sueli Bonato P, Goldman MH, Goldman GH (2006) Fungal metabolic model for tyrosinemia type 3: molecular characterization of a gene encoding a 4-hydroxy-phenyl pyruvate dioxygenase from Aspergillus nidulans. Eukaryot Cell 5 : 1441–1445. 16896227
33. Fernandez-Canon JM, Penalva MA (1998) Characterization of a fungal maleylacetoacetate isomerase gene and identification of its human homologue. J Biol Chem 273 : 329–337. 9417084
34. Sharma KK, Arst HN Jr. (1985) The product of the regulatory gene of the proline catabolism gene cluster of Aspergillus nidulans is a positive-acting protein. Curr Genet 9 : 299–304. 3916725
35. Empel J, Sitkiewicz I, Andrukiewicz A, Lasocki K, Borsuk P, et al. (2001) arcA, the regulatory gene for the arginine catabolic pathway in Aspergillus nidulans. Mol Genet Genomics 266 : 591–597. 11810230
36. Berger H, Basheer A, Bock S, Reyes-Dominguez Y, Dalik T, et al. (2008) Dissecting individual steps of nitrogen transcription factor cooperation in the Aspergillus nidulans nitrate cluster. Mol Microbiol 69 : 1385–1398. doi: 10.1111/j.1365-2958.2008.06359.x 18673441
37. Gournas C, Oestreicher N, Amillis S, Diallinas G, Scazzocchio C (2011) Completing the purine utilisation pathway of Aspergillus nidulans. Fungal Genet Biol 48 : 840–848. doi: 10.1016/j.fgb.2011.03.004 21419234
38. Miller BL, Miller KY, Roberti KA, Timberlake WE (1987) Position-dependent and -independent mechanisms regulate cell-specific expression of the SpoC1 gene cluster of Aspergillus nidulans. Mol Cell Biol 7 : 427–434. 3550422
39. Hynes MJ (1975) Studies on the role of the areA gene in the regulation of nitrogen catabolism in Aspergillus nidulans. Aust J Biol Sci 28 : 301–313. 52352
40. Hawker KL, Montague P, Kinghorn JR (1992) Nitrate reductase and nitrite reductase transcript levels in various mutants of Aspergillus nidulans: confirmation of autogenous regulation. Mol Gen Genet 231 : 485–488. 1538701
41. Oestreicher N, Ribard C, Scazzocchio C (2008) The nadA gene of Aspergillus nidulans, encoding adenine deaminase, is subject to a unique regulatory pattern. Fungal Genet Biol 45 : 760–775. 18055231
42. Welch JJ, Watts JA, Vakoc CR, Yao Y, Wang H, et al. (2004) Global regulation of erythroid gene expression by transcription factor GATA-1. Blood 104 : 3136–3147. 15297311
43. Macios M, Caddick MX, Weglenski P, Scazzocchio C, Dzikowska A (2012) The GATA factors AREA and AREB together with the co-repressor NMRA, negatively regulate arginine catabolism in Aspergillus nidulans in response to nitrogen and carbon source. Fungal Genet Biol 49 : 189–198. doi: 10.1016/j.fgb.2012.01.004 22300944
44. Fu L, Dong SS, Xie YW, Tai LS, Chen L, Kong KL, Man K, Xie D, Li Y, Cheng Y, Tao Q, Guan XY (2010) Down-regulation of tyrosine aminotransferase at a frequently deleted region 16q22 contributes to the pathogenesis of hepatocellular carcinoma. Hepatology 51 : 1624–1634. doi: 10.1002/hep.23540 20209601
45. Lee SS, Kennedy S, Tolonen AC, Ruvkun G (2003) DAF-16 target genes that control C. elegans life-span and metabolism. Science 300 : 644–647. 12690206
46. Ferguson AA, Roy S, Kormanik KN, Kim Y, Dumas KJ, Ritov VB, Matern D, Hu PJ, Fisher AL (2013) TATN-1 mutations reveal a novel role for tyrosine as a metabolic signal that influences developmental decisions and longevity in Caenorhabditis elegans. PLoS Genet 9: e1004020. doi: 10.1371/journal.pgen.1004020 24385923
47. Finch CE, Ruvkun G (2001) The genetics of aging. Annu Rev Genomics Hum Genet 2 : 435–462. 11701657
48. Borneman AR, Hynes MJ, Andrianopoulos A (2000) The abaA homologue of Penicillium marneffei participates in two developmental programmes: conidiation and dimorphic growth. Mol Microbiol 38 : 1034–1047. 11123677
49. Sambrook J, Fritsch EF, Maniatis T (1989) Molecular Cloning, A Laboratory Manual. Cold Spring Harbor, USA: Cold Spring Harbor Laboratory Press.
50. Bugeja HE, Boyce KJ, Weerasinghe H, Beard S, Jeziorowski A, et al. (2012) Tools for high efficiency genetic manipulation of the human pathogen Penicillium marneffei. Fungal Genet Biol 49 : 772–778. doi: 10.1016/j.fgb.2012.08.003 22921264
51. Ausubel FM, Brent R, Kingston RE, Moore RE, Seidman JA, et al. (1994) Current protocols in molecular biology. New York: John Wiley and Sons, Inc.
52. Cove DJ, Quatrano RS, Hartmann E (1996) The alignment of the axis of asymmetry in regenerating protoplasts of the moss, Ceratodon purpureus, is determined independently of axis polarity. Development 122 : 371–379. 8565849
Štítky
Hygiena a epidemiologie Infekční lékařství Laboratoř
Článek A Phospholipase Is Involved in Disruption of the Liver Stage Parasitophorous Vacuole MembraneČlánek Host ESCRT Proteins Are Required for Bromovirus RNA Replication Compartment Assembly and FunctionČlánek Enhanced CD8 T Cell Responses through GITR-Mediated Costimulation Resolve Chronic Viral Infection
Článek vyšel v časopisePLOS Pathogens
Nejčtenější tento týden
2015 Číslo 3- Jak souvisí postcovidový syndrom s poškozením mozku?
- Měli bychom postcovidový syndrom léčit antidepresivy?
- Farmakovigilanční studie perorálních antivirotik indikovaných v léčbě COVID-19
- 10 bodů k očkování proti COVID-19: stanovisko České společnosti alergologie a klinické imunologie ČLS JEP
-
Všechny články tohoto čísla
- To Be or Not IIb: A Multi-Step Process for Epstein-Barr Virus Latency Establishment and Consequences for B Cell Tumorigenesis
- Is Antigenic Sin Always “Original?” Re-examining the Evidence Regarding Circulation of a Human H1 Influenza Virus Immediately Prior to the 1918 Spanish Flu
- The Great Escape: Pathogen Versus Host
- Coping with Stress and the Emergence of Multidrug Resistance in Fungi
- Catch Me If You Can: The Link between Autophagy and Viruses
- Bacterial Immune Evasion through Manipulation of Host Inhibitory Immune Signaling
- Evidence for Ubiquitin-Regulated Nuclear and Subnuclear Trafficking among Matrix Proteins
- BILBO1 Is a Scaffold Protein of the Flagellar Pocket Collar in the Pathogen
- Production of Anti-LPS IgM by B1a B Cells Depends on IL-1β and Is Protective against Lung Infection with LVS
- Virulence Regulation with Venus Flytrap Domains: Structure and Function of the Periplasmic Moiety of the Sensor-Kinase BvgS
- α-Hemolysin Counteracts the Anti-Virulence Innate Immune Response Triggered by the Rho GTPase Activating Toxin CNF1 during Bacteremia
- Induction of Interferon-Stimulated Genes by IRF3 Promotes Replication of
- Intracellular Growth Is Dependent on Tyrosine Catabolism in the Dimorphic Fungal Pathogen
- HCV Induces the Expression of Rubicon and UVRAG to Temporally Regulate the Maturation of Autophagosomes and Viral Replication
- Spatiotemporal Analysis of Hepatitis C Virus Infection
- Subgingival Microbial Communities in Leukocyte Adhesion Deficiency and Their Relationship with Local Immunopathology
- Interaction between the Type III Effector VopO and GEF-H1 Activates the RhoA-ROCK Pathway
- Attenuation of Tick-Borne Encephalitis Virus Using Large-Scale Random Codon Re-encoding
- Establishment of HSV1 Latency in Immunodeficient Mice Facilitates Efficient Reactivation
- XRN1 Stalling in the 5’ UTR of Hepatitis C Virus and Bovine Viral Diarrhea Virus Is Associated with Dysregulated Host mRNA Stability
- γδ T Cells Confer Protection against Murine Cytomegalovirus (MCMV)
- Rhadinovirus Host Entry by Co-operative Infection
- A Phospholipase Is Involved in Disruption of the Liver Stage Parasitophorous Vacuole Membrane
- Dermal Neutrophil, Macrophage and Dendritic Cell Responses to Transmitted by Fleas
- Elucidation of Sigma Factor-Associated Networks in Reveals a Modular Architecture with Limited and Function-Specific Crosstalk
- A Conserved NS3 Surface Patch Orchestrates NS2 Protease Stimulation, NS5A Hyperphosphorylation and HCV Genome Replication
- Host ESCRT Proteins Are Required for Bromovirus RNA Replication Compartment Assembly and Function
- Disruption of IL-21 Signaling Affects T Cell-B Cell Interactions and Abrogates Protective Humoral Immunity to Malaria
- Compartmentalized Replication of R5 T Cell-Tropic HIV-1 in the Central Nervous System Early in the Course of Infection
- Diminished Reovirus Capsid Stability Alters Disease Pathogenesis and Littermate Transmission
- Characterization of CD8 T Cell Differentiation following SIVΔnef Vaccination by Transcription Factor Expression Profiling
- Visualization of HIV-1 Interactions with Penile and Foreskin Epithelia: Clues for Female-to-Male HIV Transmission
- Sensing Cytosolic RpsL by Macrophages Induces Lysosomal Cell Death and Termination of Bacterial Infection
- PKCη/Rdx-driven Phosphorylation of PDK1: A Novel Mechanism Promoting Cancer Cell Survival and Permissiveness for Parvovirus-induced Lysis
- Metalloprotease NleC Suppresses Host NF-κB/Inflammatory Responses by Cleaving p65 and Interfering with the p65/RPS3 Interaction
- Immune Antibodies and Helminth Products Drive CXCR2-Dependent Macrophage-Myofibroblast Crosstalk to Promote Intestinal Repair
- Adenovirus Entry From the Apical Surface of Polarized Epithelia Is Facilitated by the Host Innate Immune Response
- The RNA Template Channel of the RNA-Dependent RNA Polymerase as a Target for Development of Antiviral Therapy of Multiple Genera within a Virus Family
- Neutrophils: Between Host Defence, Immune Modulation, and Tissue Injury
- CD169-Mediated Trafficking of HIV to Plasma Membrane Invaginations in Dendritic Cells Attenuates Efficacy of Anti-gp120 Broadly Neutralizing Antibodies
- Japanese Encephalitis Virus Nonstructural Protein NS5 Interacts with Mitochondrial Trifunctional Protein and Impairs Fatty Acid β-Oxidation
- Yip1A, a Novel Host Factor for the Activation of the IRE1 Pathway of the Unfolded Protein Response during Infection
- TRIM26 Negatively Regulates Interferon-β Production and Antiviral Response through Polyubiquitination and Degradation of Nuclear IRF3
- Parallel Epigenomic and Transcriptomic Responses to Viral Infection in Honey Bees ()
- A Crystal Structure of the Dengue Virus NS5 Protein Reveals a Novel Inter-domain Interface Essential for Protein Flexibility and Virus Replication
- Enhanced CD8 T Cell Responses through GITR-Mediated Costimulation Resolve Chronic Viral Infection
- Exome and Transcriptome Sequencing of Identifies a Locus That Confers Resistance to and Alters the Immune Response
- The Role of Misshapen NCK-related kinase (MINK), a Novel Ste20 Family Kinase, in the IRES-Mediated Protein Translation of Human Enterovirus 71
- Chitin Recognition via Chitotriosidase Promotes Pathologic Type-2 Helper T Cell Responses to Cryptococcal Infection
- Activates Both IL-1β and IL-1 Receptor Antagonist to Modulate Lung Inflammation during Pneumonic Plague
- Persistence of Transmitted HIV-1 Drug Resistance Mutations Associated with Fitness Costs and Viral Genetic Backgrounds
- An 18 kDa Scaffold Protein Is Critical for Biofilm Formation
- Early Virological and Immunological Events in Asymptomatic Epstein-Barr Virus Infection in African Children
- Human CD8 T-cells Recognizing Peptides from () Presented by HLA-E Have an Unorthodox Th2-like, Multifunctional, Inhibitory Phenotype and Represent a Novel Human T-cell Subset
- Decreased HIV-Specific T-Regulatory Responses Are Associated with Effective DC-Vaccine Induced Immunity
- RSV Vaccine-Enhanced Disease Is Orchestrated by the Combined Actions of Distinct CD4 T Cell Subsets
- Concerted Activity of IgG1 Antibodies and IL-4/IL-25-Dependent Effector Cells Trap Helminth Larvae in the Tissues following Vaccination with Defined Secreted Antigens, Providing Sterile Immunity to Challenge Infection
- Structure of the Low pH Conformation of Chandipura Virus G Reveals Important Features in the Evolution of the Vesiculovirus Glycoprotein
- PPM1A Regulates Antiviral Signaling by Antagonizing TBK1-Mediated STING Phosphorylation and Aggregation
- Lipidomic Analysis Links Mycobactin Synthase K to Iron Uptake and Virulence in .
- Roles and Programming of Arabidopsis ARGONAUTE Proteins during Infection
- Impact of Infection on Host Macrophage Nuclear Physiology and Nucleopore Complex Integrity
- The Impact of Host Diet on Titer in
- Antimicrobial-Induced DNA Damage and Genomic Instability in Microbial Pathogens
- Herpesviral G Protein-Coupled Receptors Activate NFAT to Induce Tumor Formation via Inhibiting the SERCA Calcium ATPase
- The Causes and Consequences of Changes in Virulence following Pathogen Host Shifts
- Small GTPase Rab21 Mediates Fibronectin Induced Actin Reorganization in : Implications in Pathogen Invasion
- Positive Role of Promyelocytic Leukemia Protein in Type I Interferon Response and Its Regulation by Human Cytomegalovirus
- NEDDylation Is Essential for Kaposi’s Sarcoma-Associated Herpesvirus Latency and Lytic Reactivation and Represents a Novel Anti-KSHV Target
- β-HPV 5 and 8 E6 Disrupt Homology Dependent Double Strand Break Repair by Attenuating BRCA1 and BRCA2 Expression and Foci Formation
- An O Antigen Capsule Modulates Bacterial Pathogenesis in
- Variable Processing and Cross-presentation of HIV by Dendritic Cells and Macrophages Shapes CTL Immunodominance and Immune Escape
- Probing the Metabolic Network in Bloodstream-Form Using Untargeted Metabolomics with Stable Isotope Labelled Glucose
- Adhesive Fiber Stratification in Uropathogenic Biofilms Unveils Oxygen-Mediated Control of Type 1 Pili
- Vaccinia Virus Protein Complex F12/E2 Interacts with Kinesin Light Chain Isoform 2 to Engage the Kinesin-1 Motor Complex
- Modulates Host Macrophage Mitochondrial Metabolism by Hijacking the SIRT1-AMPK Axis
- Human T-Cell Leukemia Virus Type 1 (HTLV-1) Tax Requires CADM1/TSLC1 for Inactivation of the NF-κB Inhibitor A20 and Constitutive NF-κB Signaling
- Suppression of RNAi by dsRNA-Degrading RNaseIII Enzymes of Viruses in Animals and Plants
- Spatiotemporal Regulation of a T4SS Substrate by the Metaeffector SidJ
- Antigenic Properties of the Human Immunodeficiency Virus Envelope Glycoprotein Gp120 on Virions Bound to Target Cells
- Dependence of Intracellular and Exosomal microRNAs on Viral Oncogene Expression in HPV-positive Tumor Cells
- Identification of a Peptide-Pheromone that Enhances Escape from Host Cell Vacuoles
- Impaired Systemic Tetrahydrobiopterin Bioavailability and Increased Dihydrobiopterin in Adult Falciparum Malaria: Association with Disease Severity, Impaired Microvascular Function and Increased Endothelial Activation
- Transgenic Expression of the Dicotyledonous Pattern Recognition Receptor EFR in Rice Leads to Ligand-Dependent Activation of Defense Responses
- Comprehensive Antigenic Map of a Cleaved Soluble HIV-1 Envelope Trimer
- Low Doses of Imatinib Induce Myelopoiesis and Enhance Host Anti-microbial Immunity
- Impaired Systemic Tetrahydrobiopterin Bioavailability and Increased Oxidized Biopterins in Pediatric Falciparum Malaria: Association with Disease Severity
- PLOS Pathogens
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Bacterial Immune Evasion through Manipulation of Host Inhibitory Immune Signaling
- BILBO1 Is a Scaffold Protein of the Flagellar Pocket Collar in the Pathogen
- Antimicrobial-Induced DNA Damage and Genomic Instability in Microbial Pathogens
- Attenuation of Tick-Borne Encephalitis Virus Using Large-Scale Random Codon Re-encoding
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



