-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaFGF Signaling Regulates the Number of Posterior Taste Papillae by Controlling Progenitor Field Size
The sense of taste is fundamental to our ability to ingest nutritious substances and to detect and avoid potentially toxic ones. Sensory taste buds are housed in papillae that develop from epithelial placodes. Three distinct types of gustatory papillae reside on the rodent tongue: small fungiform papillae are found in the anterior tongue, whereas the posterior tongue contains the larger foliate papillae and a single midline circumvallate papilla (CVP). Despite the great variation in the number of CVPs in mammals, its importance in taste function, and its status as the largest of the taste papillae, very little is known about the development of this structure. Here, we report that a balance between Sprouty (Spry) genes and Fgf10, which respectively antagonize and activate receptor tyrosine kinase (RTK) signaling, regulates the number of CVPs. Deletion of Spry2 alone resulted in duplication of the CVP as a result of an increase in the size of the placode progenitor field, and Spry1−/−;Spry2−/− embryos had multiple CVPs, demonstrating the redundancy of Sprouty genes in regulating the progenitor field size. By contrast, deletion of Fgf10 led to absence of the CVP, identifying FGF10 as the first inductive, mesenchyme-derived factor for taste papillae. Our results provide the first demonstration of the role of epithelial-mesenchymal FGF signaling in taste papilla development, indicate that regulation of the progenitor field size by FGF signaling is a critical determinant of papilla number, and suggest that the great variation in CVP number among mammalian species may be linked to levels of signaling by the FGF pathway.
Published in the journal: . PLoS Genet 7(6): e32767. doi:10.1371/journal.pgen.1002098
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002098Summary
The sense of taste is fundamental to our ability to ingest nutritious substances and to detect and avoid potentially toxic ones. Sensory taste buds are housed in papillae that develop from epithelial placodes. Three distinct types of gustatory papillae reside on the rodent tongue: small fungiform papillae are found in the anterior tongue, whereas the posterior tongue contains the larger foliate papillae and a single midline circumvallate papilla (CVP). Despite the great variation in the number of CVPs in mammals, its importance in taste function, and its status as the largest of the taste papillae, very little is known about the development of this structure. Here, we report that a balance between Sprouty (Spry) genes and Fgf10, which respectively antagonize and activate receptor tyrosine kinase (RTK) signaling, regulates the number of CVPs. Deletion of Spry2 alone resulted in duplication of the CVP as a result of an increase in the size of the placode progenitor field, and Spry1−/−;Spry2−/− embryos had multiple CVPs, demonstrating the redundancy of Sprouty genes in regulating the progenitor field size. By contrast, deletion of Fgf10 led to absence of the CVP, identifying FGF10 as the first inductive, mesenchyme-derived factor for taste papillae. Our results provide the first demonstration of the role of epithelial-mesenchymal FGF signaling in taste papilla development, indicate that regulation of the progenitor field size by FGF signaling is a critical determinant of papilla number, and suggest that the great variation in CVP number among mammalian species may be linked to levels of signaling by the FGF pathway.
Introduction
Taste sensory capability is mediated by aggregates of receptor cells, called taste buds, which reside within the oral and pharyngeal cavities. The majority of taste buds in mammals reside on the tongue in epithelial-mesenchymal specializations termed gustatory papillae. In the rodent tongue, the smaller fungiform papillae, each of which possesses a single taste bud, are found in a distributed array on the anterior tongue. By contrast, the larger bilateral foliate papillae and a single midline circumvallate papilla (CVP) each house hundreds of buds and reside on the posterior tongue (Figure 1A). Recently, there has been increasing recognition that anterior fungiform taste buds differ from those of the posterior CVP in terms of both gene expression and taste function [1]–[4].
Fig. 1. Deletion of Spry2 leads to a duplication of the CVP in the posterior tongue. 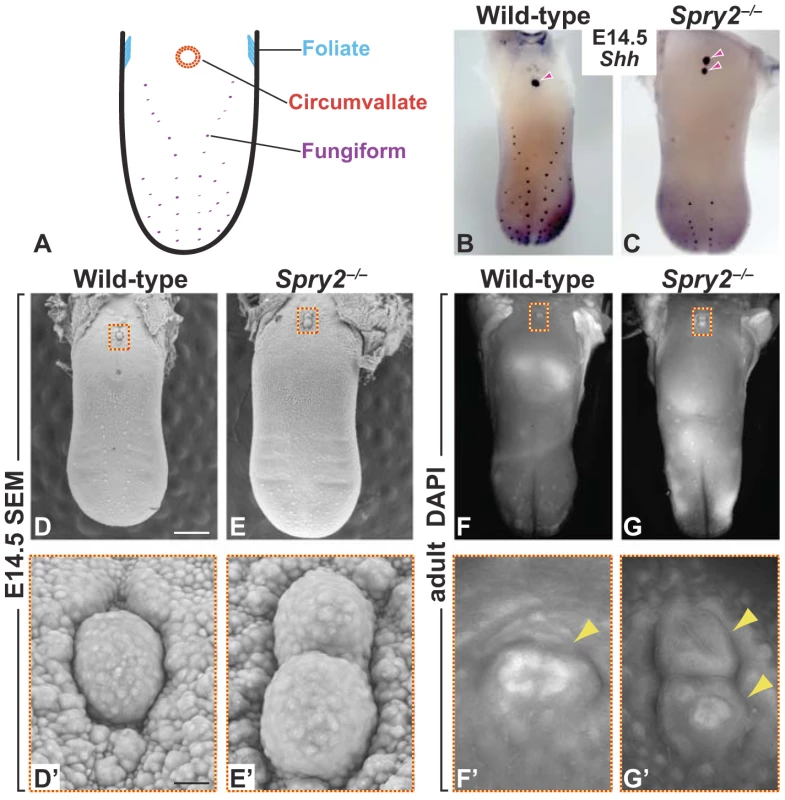
(A) Cartoon showing location of gustatory papillae in the rodent tongue. (B,C) In situ hybridization staining of Shh. Wild-type mice possess a single CVP in the posterior tongue, whereas the CVP is duplicated in Spry2−/− mice (arrowheads). (D,E) SEM images of the tongue at E14.5. Scale bar, 500 µm. (F,G) DAPI fluorescence staining shows that the two CVPs (arrowheads) persist into adulthood in Spry2−/− mice. (D'-G') Higher magnification images of boxed areas. (D',E') Scale bar, 25 µm. In recent years, significant progress has been made in defining the molecular regulation of fungiform development. Fungiform papillae initially form as placodes that subsequently undergo epithelial morphogenesis and acquire a mesenchymal core [5] in a process that is similar to morphogenesis of other vertebrate epithelial specializations, such as hair, teeth, and mammary glands [6], [7]. The development of these other organs requires signaling between epithelium and mesenchyme, suggesting that such epithelial-mesenchymal interactions are also involved in patterning and morphogenesis of taste placodes. However, expression of all of the key signaling factors implicated in taste placode development – including Sonic Hedgehog (SHH), Bone Morphogenetic Proteins (BMPs), Epidermal Growth Factor (EGF), and WNTs – is restricted to the epithelium [8]–[15]. Thus, to date, no inductive, mesenchyme-derived factor involved in taste development has been identified.
Despite its importance in taste function and its status as the largest of the taste papillae, very little is known about the genes involved in the development of the CVP. Like fungiform placodes, the CVP forms as an initial epithelial thickening that undergoes complex morphogenesis to form a large papilla. However, it appears that genes known to regulate fungiform development do not function similarly in development of the CVP. For example, inhibition of SHH results in more and larger fungiform placodes, but has no effect on the CVP [11]. In addition, BMP7 and its antagonist follistatin have significant functions in fungiform development, but the CVP appears unaffected by inactivation of either gene [16]. These differences may be ascribed to the distinct embryonic origins of the anterior tongue, which is thought to be derived from ectoderm, whereas the posterior tongue likely has endodermal origins [4].
Expression of several Fibroblast Growth Factors (FGFs) and their receptors has previously been detected in the developing tongue [17]. Therefore, we hypothesized that Sprouty (Spry) genes, which antagonize several receptor-tyrosine kinase (RTK) signaling pathways including those triggered by FGFs [18]–[20], may play a role in the development of taste papillae. Originally, spry was identified as a regulator of tracheal branching in Drosophila [21], and it was later found that three of the four mouse Sprouty genes (Spry1, Spry2, and Spry4) are expressed during embryonic development [22], [23].
Here, we used mouse genetic models to show that FGF signaling is required for CVP formation. We found that Spry1 and Spry2 antagonize signaling by FGF10 to restrict the size of the progenitor field of the circumvallate (CV) placode, such that loss of Sprouty function results in a dramatic expansion of the CV placode as it first forms. In adult Spry2 mutants, a striking and complete duplication of the CVP emerges, whereas embryos lacking both Spry1 and Spry2 have multiple CVPs. Our findings thus represent the first example of molecular genetic regulation of taste organs in the posterior tongue. We found that Fgf10, which is expressed exclusively in the mesenchyme underlying the CV placode, is required for formation of the CVP, and thus we provide the first example of an inductive, mesenchymal signal involved in specifying the epithelial domain of a developing taste papilla. Further, while FGF10 signaling drives CVP development and Spry1 and Spry2 repress this process, fungiform taste papillae are oppositely affected by the loss of these genes: Fgf10−/− tongues appear to possess more and larger fungiform papillae, and the loss of Sprouty genes results in fewer fungiform papillae. Thus, these results demonstrate that molecular mechanisms regulating development of anterior and posterior taste organs differ considerably. Finally, we postulate that the role of FGF signaling in defining the size of the CV progenitor field in mice may underlie the large variation in CVP number across mammalian taxa, and that changes in FGF signaling during evolution may have caused expansion of the initial progenitor field to allow formation of multiple CVPs in some species.
Results
Deletion of Spry2 leads to a duplication of the CVP
Wild-type mice possess a single CVP in the midline of the posterior tongue; foliate papillae reside on the lateral tongue and multiple fungiform papillae populate the anterior tongue (Figure 1A, 1B). In Spry2−/− mice, we observed a duplication of the CVP in the anterior-posterior orientation at embryonic day (E) 14.5 using Shh expression as a marker (Figure 1C). By contrast, fungiform placodes in the anterior tongue, which also express Shh, were significantly reduced in number (61±4.37 in wild-type versus 33±2.02 in Spry2−/− embryos; p = 0.0002; n = 6 embryos per genotype). Because alterations in fungiform papilla development had been reported in other mutants [9]–[15], we chose to pursue the novel CVP duplication phenotype. The presence of two CVPs in the embryonic tongue was confirmed using scanning electron microscopy (SEM; Figure 1D-1E'). To test whether the duplicated CVP persisted into adulthood, we used DAPI staining in adult tongues and again identified two discrete papillae (Figure 1F-1G'). Notably, both CVPs in Spry2−/− mice house fully differentiated taste buds containing the three types of taste receptor cells, and both showed positive staining for b3-tubulin, indicating that taste buds and the papillae are innervated (Figure S1). Taken together, these data indicate that the duplication in Spry2−/− embryos leads to two functional adult CVPs.
To reconstruct the development of the CVP, closely staged specimens between E11.5 to E14.5 were stained with E-cadherin and imaged in whole-mount (Figure 2A-2N), as well as analyzed by H&E staining of sections (Figure 2O-2X). In wild-type embryos, CVP development was initiated shortly after the tongue rudiment formed by fusion of the lateral lingual swellings, and it was detected as a more compact collection of epithelial cells at E11.5 (Figure 2A) [24]. At E12.5, a placodal condensation was readily observable on coronal sections (Figure 2O). Between E13 and E13.5, the wild-type placode underwent major morphological changes as the epithelial trenches characteristic of the CVP began to form (Figure 2P, 2Q). The growth of the trenches was coincident with the initiation of innervation at approximately E13 (Figure S2), and the trenches continued to grow into the mesenchyme at E14.5 (Figure 2G, 2S) [24], [25].
Fig. 2. Development of the CVP in wild-type and Spry2−/− mice. 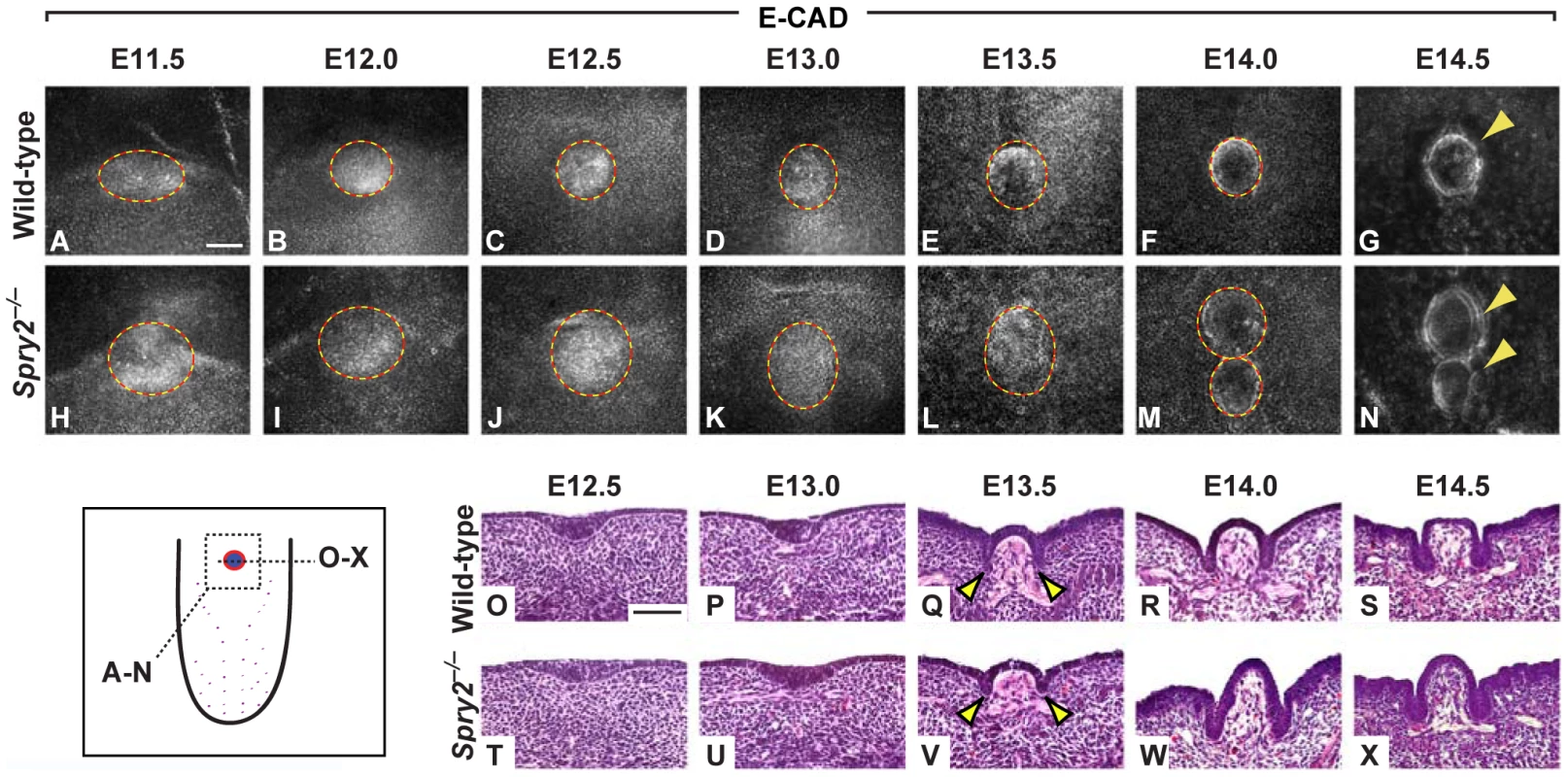
(Inset) Cartoon depicting the planes of observation. (A–N) E-cadherin (E-CAD) immunofluorescence staining of CV placodes and papillae in wild-type (A–G) and Spry2−/− (H-N) mice. (O–X) H&E stains of coronal sections of the single CV placode and papilla in wild-type and the proximal CV placode and papilla in Spry2−/− mice. The size of the papilla was quantified between E13 to E14 (D–F, K–M; ×10−3 um). Arrowheads indicate developing trenches. Scale bars, 50 µm. In contrast to the wild-type, the CV placode in Spry2−/− embryos was larger at E11.5 (compare Figure 2A, 2H). Relative to the wild-type, the mutant placode continued to grow larger both laterally and along the anterior-posterior axis between E11.5 and E13.5 (Figure 2B-2E, 2I-2L). By E14, two distinct structures were observed in Spry2−/− embryos, with the anterior of the two papillae usually appearing smaller than the posterior at E14 and E14.5 (Figure 2M, 2N). In addition, CVPs in Spry2−/− embryos showed a raised dome shape compared to the flatter shape of the wild-type CVP (Figure 2R, 2S, 2W, 2X).
Several possible mechanisms could account for the duplication of the CVP at ∼E14, including a timer that causes the organ to split at a certain point after its development starts or a threshold that causes splitting after a critical size limit is achieved. To address this issue, we quantified the size of the CV placode and CVP in wild-type and Spry2−/− tongues between E13 to E14, corresponding to the time points before, during and after CVP duplication (Figure S2J). Relative to wild-type mice, the mutant CV placode was already significantly larger at E13 and E13.5, but when CVP duplication was observed at E14, there was a further and dramatic increase in the total area of the papillae. Thus, our data suggest that there is a critical threshold size for the CV placode, beyond which the placode destabilizes, leading to CVP duplication.
The CV progenitor field is enlarged in Spry2−/− mice
Sox2 is one of the earliest markers of taste placodes and is required for taste bud development [26]. Sox2 expression was expanded in the placode domain in both anterior-posterior and lateral directions in Spry2−/− embryos compared to wild-type littermates at both E11.5 (data not shown) and E12.5 (Figure 3A-3B'). Next, we analyzed the expression of three additional factors that mark the early placode: Shh, Bmp7, and Wnt10b (Figure 3C-3H'). Shh has previously been shown to be expressed in the developing CVP [9], [27], and in fungiform placodes, Shh is expressed specifically by taste bud progenitors [28]. Bmp7 and Wnt10b have also been implicated in development of fungiform papillae [15], [16]. The expression domains of all these markers were expanded in the CV placode of Spry2−/− embryos relative to wild-type (Figure 3C-3H'). We quantified the differences in expression levels by qPCR, and interestingly, whereas Sox2 and Bmp7 levels were strongly upregulated in the mutants, consistent with the expanded expression domains seen in whole mount, Shh and Wnt10b expression levels were not dramatically different in the mutants (Figure S3A). This is consistent with the decreased intensity of Shh and Wnt10b staining despite an increased CV placodal domain size (Figure 3D, 3D', 3H, 3H').
Fig. 3. The CV progenitor field at E12.5 is expanded in Spry2−/− mice. 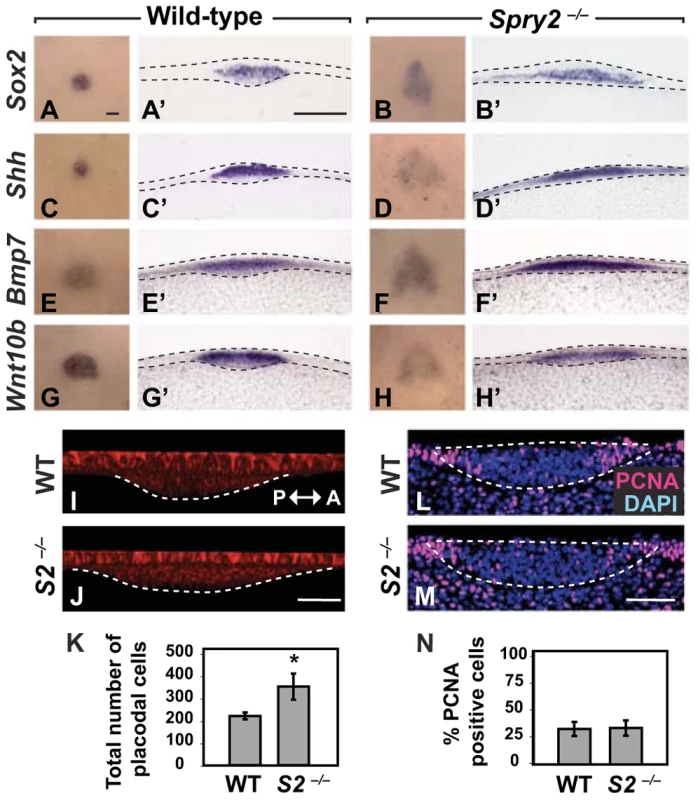
(A–H) Dorsal view of the expression of Sox2, Shh, Bmp7, and Wnt10b in wild-type and Spry2−/− littermates. Scale bar, 50 µm. (A'–H') Coronal sections of A–H. Scale bar, 50 µm. (I,J) Sagittal views of the CV placodes stained with E-cadherin show an expansion of the placode in the anterior(A)-posterior(P) orientation in Spry2−/− mice. Scale bar, 20 µm. (K) Quantification of the total number of cells in the CV placode. (L,M) Proliferating cells were detected by PCNA staining in coronal view. Scale bar, 20 µm. (N) The percentage of PCNA-positive cells in the CV placode at E12.5. * p<0.01. WT, wild-type; S2−/−, Spry2−/−. We next asked whether there was a difference in the number of cells in the placode between wild-type and mutant. We quantified the number of cells in the CV placode of Spry2−/− embryos relative to wild-type littermates at E12.5 from three-dimensional confocal images of E-cadherin-positive stained cells (Figure 3I-3K). We counted cells in 10 mm intervals beginning at the deepest tip of invaginated cells using ventral views; the borders of the placode were defined by the invagination. This analysis showed that already by the mid-placode stage, the mutant placode had almost twice as many cells as the wild-type. Because differences in proliferation or cell death could account for the larger placode and the morphogenetic abnormalities in the CVPs of Spry2 mutants, we assayed proliferation by PCNA staining (Figure 3L-3N), and cell death by TUNEL staining (Figure S3B-S3D). We detected no differences between wild-type and mutant CV placode at E12.5 for either assay, suggesting that a larger number of cells was recruited into the placode at the earliest stages of placodogenesis. Thus, our data indicate that inactivation of Spry2 leads to a larger progenitor field, as detected by multiple molecular markers, such that the duplication in the Spry2 mutant results from recruitment of more cells into the CV placode at the earliest stages of its development.
Spry1 and Spry2 are co-expressed in the CV placode, and hypersensitivity to FGF signaling is detected in Spry2 mutants
Because Sprouty genes are often co-expressed during development [23], [29], we assayed for expression of Spry1, Spry2, and Spry4 in CV placodes at E12.5 (Figure 4A-4C); Spry3 was not analyzed due to its lack of expression in embryonic craniofacial tissues [23]. We found that Spry1 and Spry2 were both expressed in the epithelium of the CV placode at E12.5, whereas no expression was observed for Spry4 (Figure 4A-4C).
Fig. 4. Spry1 and Spry2 are co-expressed in the CV placode and increased FGF signaling is detected in Spry2 mutants. 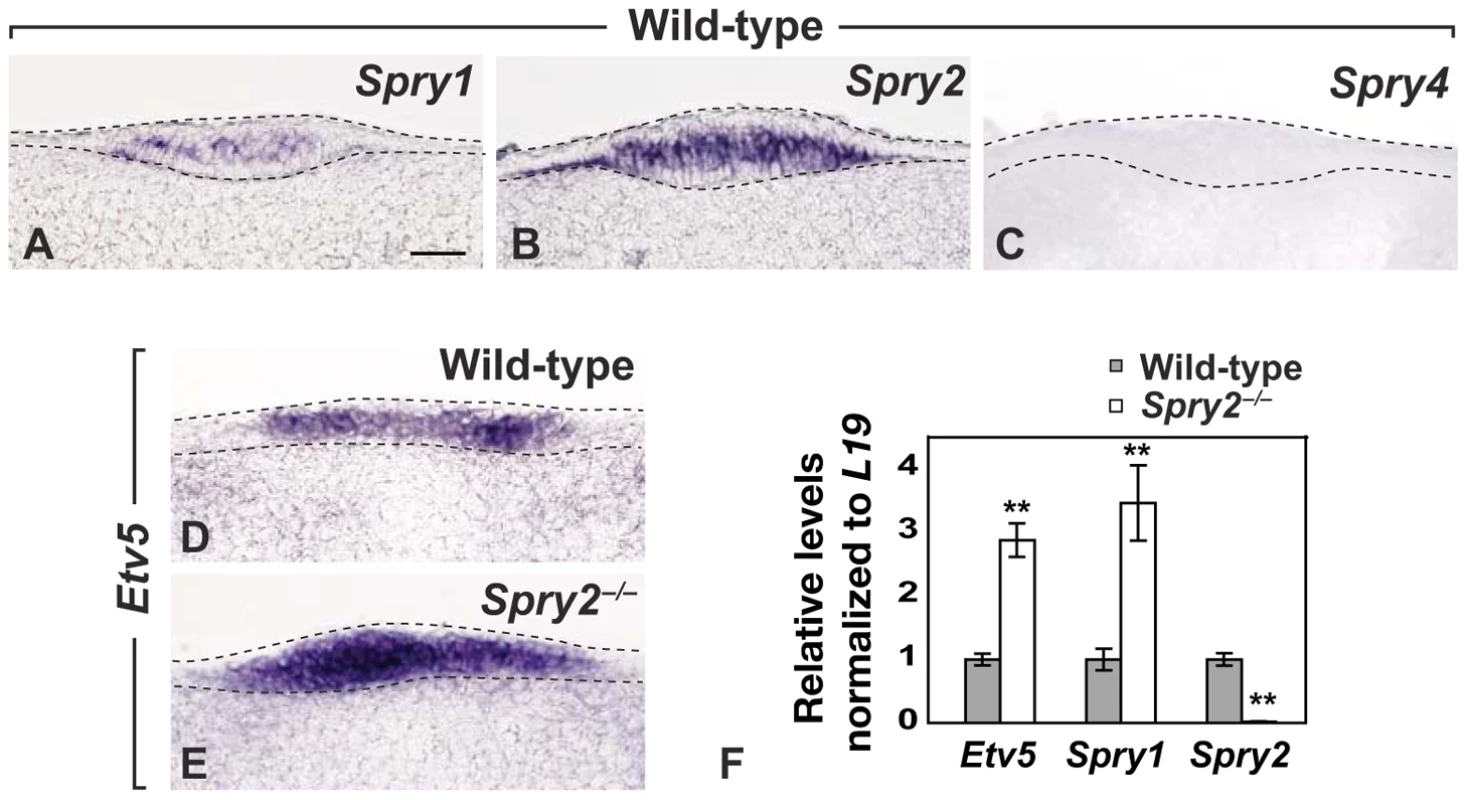
(A–C) Expression of Spry1, Spry2, and Spry4 in the CV placode of wild-type mice. (D,E) Expression of Etv5 in the CV placode of wild-type and Spry2−/− mice. (F) qPCR analysis of the expression profiles of Etv5 and Sprouty genes in the posterior tongue of wild-type and Spry2−/− mice at E12.5. Scale bar, 20 µm. **, p<0.001. The expression of Etv5 (Erm), which is a target of FGF signaling [30], was upregulated in Spry2−/− embryos (compare Figure 4D, 4E), consistent with increased FGF signaling observed with Sprouty loss-of-function in other developmental contexts [31]–[34]. To quantify expression levels, RNA was extracted from the area containing the CV placode at E12.5 and analyzed by qPCR, and a three-fold increase in Etv5 was observed (Figure 4F). As expected, essentially no Spry2 expression was detected in the Spry2 nulls, but interestingly, Spry1 expression was dramatically increased in Spry2−/− mice (Figure 4F). This upregulation is consistent with the known role of Sprouty genes as FGF targets [21], [23], and it suggested that increased expression of Spry1 may serve a compensatory role in Spry2−/− mice. qPCR showed that Spry4 was expressed at very low levels, with no detectable difference between wild-type and Spry2−/− littermates (data not shown).
Fgf10 is required for CVP development and is antagonized by Spry2
The epithelial expression of Spry1, Spry2, and Etv5, all of which are expressed in response to RTK activity [21], [23], [30], strongly suggested the presence of a mesenchymal RTK ligand that signals to the epithelium. To identify such a ligand, we surveyed the expression of a number of candidate RTKs and their cognate ligands by ISH, including Fgfr1-4, Egfr, Fgf7, Fgf8, Fgf9, and Fgf10 (data not shown). Of the ligands we examined, only Fgf10 was expressed in the mesenchyme subjacent to the epithelium along the midline of the posterior tongue at E11.5 and E12.5 (Figure 5A-5B'). Two FGF receptors, Fgfr2 and Fgfr3, were also expressed in the CV placode, whereas no expression of Fgfr1 or Fgfr4 was detected (Figure 5C-5F).
Fig. 5. Fgf10 is required for CVP development and is antagonized by Spry2. 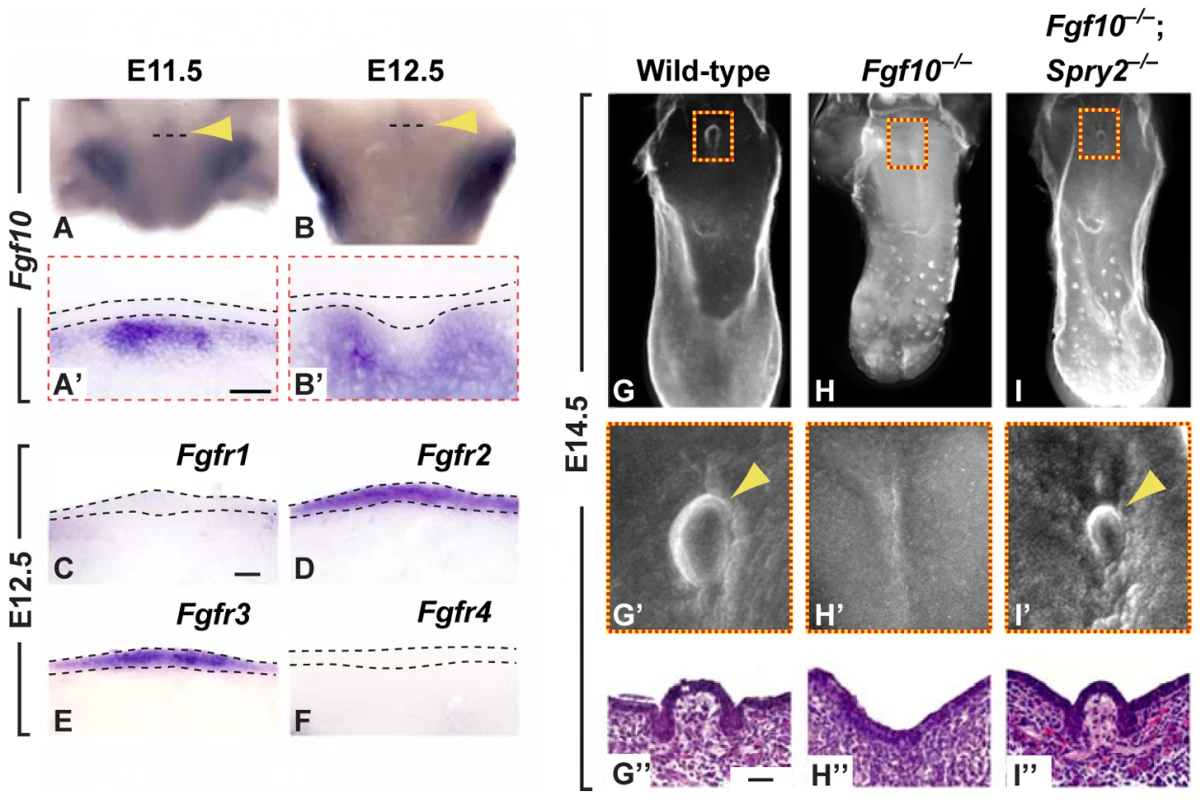
(A–B') Expression of Fgf10 in the CV placode of wild-type mice (arrowhead). Dashed line depicts the coronal plane of section for A' and B'. (C–F) Expression of FGF receptors (Fgfr1-4) in coronal sections. (G–I”) Genetic rescue of the loss of CVP phenotype in Fgf10−/− mice (H) by deletion of Spry2 (I) as visualized by E-cadherin immunofluorescence staining. (G'–I') Higher magnification views of boxed areas. (G”–I”) H&E staining of coronal sections through the CVP. Scale bars, 25 µm. To further test the role of FGF signaling in CVP development, we isolated tongues from wild-type embryos at E11.5 and E12.5 and grew them in vitro either in the absence or presence of SU5402 [35], a potent FGF receptor inhibitor (Figure S4). In the absence of SU5402, we found the formation of a single CVP. The inhibition of FGF signaling by SU5402 led to the absence of the CVP in tongues cultured at E11.5 and to either an absent or a malformed CVP in tongues cultured at E12.5. These observations demonstrate the requirement of FGF signaling during CVP formation at E11.5 and to a lesser extent at E12.5.
Because Fgf10 was expressed directly underneath the developing CV placode, we hypothesized that the ligand encoded by this gene may play a role in CV development, and therefore we examined the CVP in Fgf10−/− embryos. Strikingly, the deletion of Fgf10 led to the complete absence of the CVP (Figure 5H-5H”), indicating that this gene plays a critical inductive role in development of this structure. Next, we reasoned that the combined deletion of Fgf10 and Spry2 may balance each other and rescue CVP development, and indeed, we found a single CVP in Fgf10−/−;Spry2−/− mice (Figure 5I-5I”). This result also implies the presence of a yet-unknown secondary ligand that can fulfill the function of FGF10 if the epithelium is rendered hypersensitive to RTK signaling by inactivation of Spry2. Taken together, our data indicate that the FGF signaling pathway is critical for proper CVP development and that it determines the final number of CVPs.
Combined deletion of Spry1 and Spry2 leads to multiple CVPs
Based on the overlapping expression of Spry1 and Spry2 in the CV placode and the upregulation of Spry1 in Spry2−/− embryos (Figure 4F), we hypothesized that these two family members may be partially redundant. We therefore generated and analyzed Spry1−/− as well as Spry1−/−;Spry2−/− (DKO) embryos. No difference in the number or morphology of the CVP was detected in Spry1−/− embryos (data not shown), indicating that Spry1 is not required for CVP development if Spry2 is present. In contrast, CVPs from DKO embryos were dramatically abnormal relative to control Spry1+/−;Spry2+/− (DHet) littermates, whose tongues were indistinguishable from those of wild-types both as embryos and as adults (data not shown). At E12.5, the expression domains of Etv5 and Sox2 were expanded in the CV placodes of DKO mice relative to DHet littermates (Figure 6A, 6B, 6F, 6G). The expanded expression domains in the DKOs were greater than those seen in Spry2−/− mice (compare to Figure 4D, 4E and Figure 3A-3B'). At later stages, in contrast to the single duplication seen in Spry2−/− mice, the DKO embryos had multiple CVPs (Figure 6C, 6E, 6H, 6J). The multiple CVPs in DKO embryos were smaller than the single CVP in controls and showed pronounced raised domes (Figure 6D, 6I) similar to Spry2−/− embryos (Figure 2S, 2X). Immunofluorescence staining of β3-tubulin revealed innervation of the multiple CVPs (Figure S5).
Fig. 6. Formation of multiple CVPs in Spry1−/−;Spry2−/− (DKO) mice. 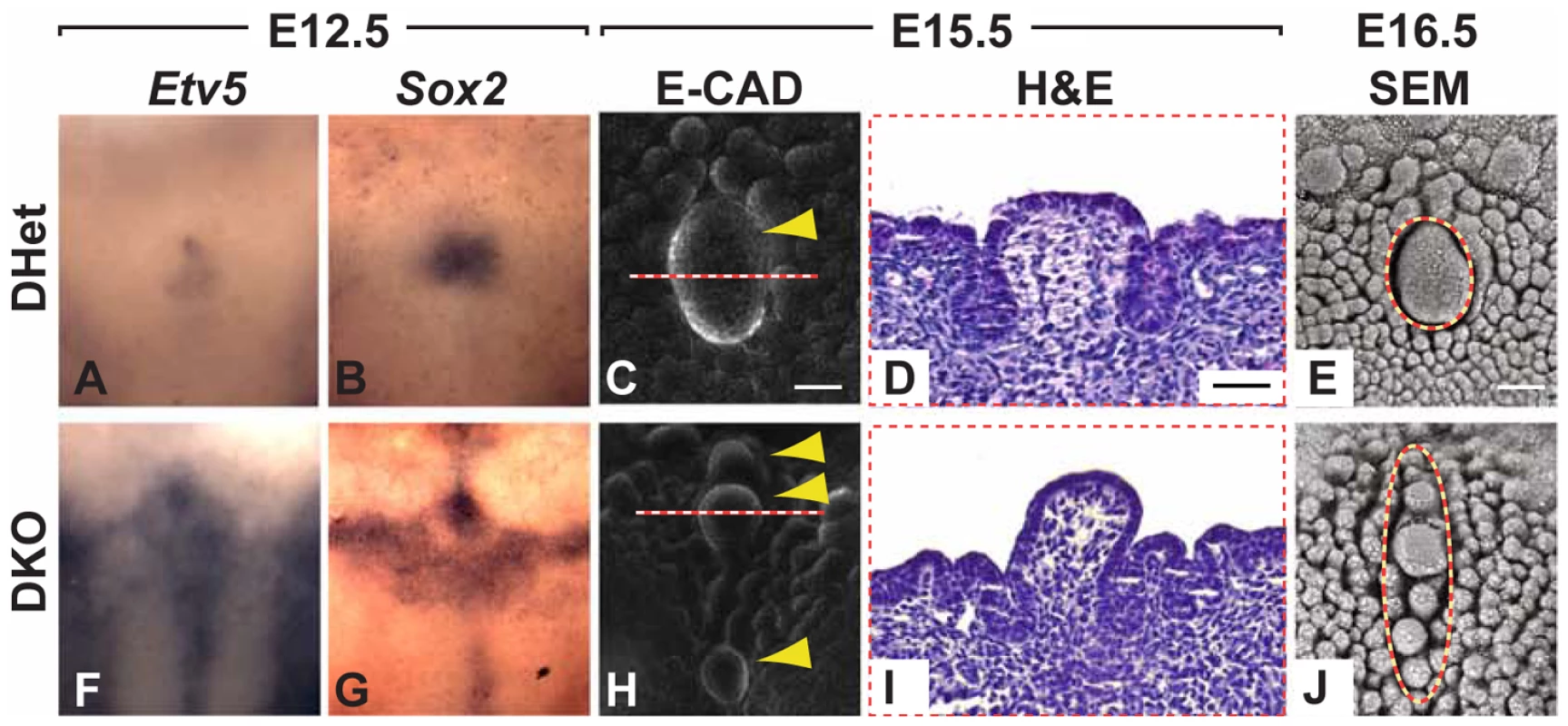
(A,B,F,G) Dorsal views of the expression patterns of Etv5 and Sox2 in Spry1+/−;Spry2+/− (DHet) and DKO mice. (C,H) Multiple CVPs in the anterio-posterior direction were observed in DKO mice visualized by E-cadherin immunofluorescence staining (arrowheads). Scale bar, 50 µm. (D,I) H&E staining of coronal sections through the CVP as indicated by dashed lines in C and H. Scale bar, 25 µm. (E,J) Multiple, presumptive CVPs visualized by SEM are indicated by dashed circles. Scale bar, 50 µm. Discussion
Of the numerous gustatory papillae, very little is known about the development of the CVP, despite the great variation in CVP number in mammals, its importance in taste function, and its status as the largest of the taste papillae. Most of the studies on gustatory organ and taste development have focused on fungiform papillae, and it is probably for this reason that significant genetic abnormalities of the CVP have not yet been reported in mice. Here, we report several findings: first, FGF signaling plays a key role in the regulation of taste papilla development; second, FGF10 is an inductive, mesenchyme-derived factor required for CVP development; third, the regulation of the progenitor field size by Sprouty genes is a critical determinant of CVP number. Finally, we postulate that the great variation in CVP number among mammalian species may be linked to alterations in the genetic dosage of agonists and antagonists in the FGF signaling pathway.
Antagonism between mesenchymal FGF10 and epithelial Sprouty genes controls CVP number
Antagonism of FGF signaling by Sprouty genes occurs during development of a number of organs [31]–[34]. Because FGFs and their receptors had previously been detected in the developing tongue [17], we hypothesized that Sprouty genes, which modulate several RTK signaling pathways including those triggered by FGFs [18]–[20], may play a role in taste papillae development. Whereas deletion of Spry2 led to a duplication of the CVP, Fgf10−/− mice showed a complete loss of the CVP, and importantly, CVP development was rescued in compound Spry2−/−; Fgf10−/− mutants. These results confirm the specific antagonism of FGF10 signaling by SPRY2 and establish the importance of the FGF signaling pathway in determining CVP number. Additionally, the results with the Spry2−/−;Fgf10−/− mutants indicate the presence of an unknown, secondary factor; such a factor could induce CVP formation in lieu of FGF10 when the epithelium is hypersensitive to RTK signaling due to the absence of Spry2.
Because of the overlapping expression profiles of Spry1 and Spry2, and the upregulation of Spry1 in the Spry2−/− CV placode, we generated Spry1−/−; Spry2−/− embryos. We observed multiple CVPs along the A-P axis in the compound mutant embryos, demonstrating the redundancy of Sprouty genes, for which there is precedent in other tissues [32], in regulating CVP development. Although a slightly dysplastic CVP was recently reported in ectodysplasin mutant mice [36], the phenotypes in Fgf10−/−, Spry2−/−, and Spry1−/−;Spry2−/− mice are the first genetic abnormalities reported involving the regulation of CVP number.
Despite indications that mesenchyme-derived factors are involved in lingual papillae development [5], [37], the only mesenchymal factor implicated in fungiform papillae formation to date is follistatin, an inhibitor of BMP signaling [16]. Therefore, FGF10 is the first inductive, mesenchyme-derived factor to be identified in taste papilla development.
Expansion of the CV progenitor field
The duplication of the CVP at E14 in Spry2−/− embryos was preceded by an increase in placode size. The CV placode in mutant mice was significantly larger at E13 and E13.5 relative to wild-type, and at E14 there was a dramatic increase in total CVP size as duplication occurred (Figure S2J). Thus, there appears to be a critical size threshold for the CV placode, beyond which the placode destabilizes, resulting in splitting and duplication. Recently, Munne et al. [38] reported that, during incisor tooth development, the impairment of BMP4 signaling or an increase in Activin concentration led to the destabilization of the large, single placode and the formation of multiple incisors. Future experiments will enable comparison of the relative roles of different signaling pathways in maintenance of epithelial placode integrity.
The increase in placode size at E13 and E13.5 was preceded at E11.5 and E12.5 by the expansion of the expression domains for various genes, such as Etv5, Sox2, Shh, Bmp7, and Wnt10b. By E12.5, the mutant CV placode already had more cells than the control, and we did not detect any apoptosis in the CV placode or any differences in the percentage of proliferating cells. Thus, the larger CV placode in the Spry2 mutant appears to result at least in part from an increase in the recruitment of CV placodal cells. At the earliest stages of placode formation, this could potentially occur either through specification of more placodal precursors or through migration of more cells into the placodal domain. Future studies will be needed to distinguish between these possibilities, but interestingly, several reports have pointed to a role for Sprouty genes in cell migration [39]–[41].
In the CV placode of Spry1−/−;Spry2−/− tongues, there were further increases in the expression domains of Etv5 and Sox2 beyond what was seen in the Spry2−/− alone, demonstrating the redundancy of Spry1 and Spry2 in this organ (Figure 6A, 6B, 6F, 6G). The compound deletion of Spry1 and Spry2 led to an even larger placode (Figure 6A, 6B, 6F, 6G), which resulted in more than two CVPs.
Comparision of fungiform and CV papillae
Whereas the deletion of Spry2 led to CVP duplication, it also resulted in a marked decrease in the number of fungiform papillae (Figure 1B, 1C). In contrast, in Fgf10−/−mice, there was an absence of the CVP, whereas the fungiform papillae appeared to be larger and more closely spaced (Figure 4G, 4H). Thus, Fgf10 and Sprouty genes differentially affect the anterior versus posterior taste fields. Together with the previous studies showing effects of SHH and BMP7 on anterior but not posterior taste fields, these results provide further evidence for important developmental differences between these fields [9], [10], [12], [16]. This observation is consistent with the notion that fungiform papillae in the anterior tongue are derived from ectoderm, whereas the CVP is likely derived from endoderm [4]. The diverse responses of fungiform versus CV papillae to developmental factors have only recently been appreciated and will require further efforts to tease apart.
Evolution of CVP number in mammals
Although most rodents, including the mouse, possess a single midline CVP in the posterior tongue, there is great variation in mammalian CVP number (Figure S6B, Figure S7). For example, humans possess anywhere from 3 to 14 CVPs in an inverted V or Y orientation [42]. Other mammals such as the hyrax [43], [44] and hippopotamus [45] possess no CVP, whereas siamang, chimpanzee, gorilla [46] and lemur [47] possess at least two CVPs along the midline, in addition to varying numbers of lateral CVPs (Figure S6A). We have shown that modifications in FGF signaling can lead to increased or decreased numbers of CVPs. Thus, we speculate that changes in levels of signaling through this pathway provide an attractive candidate for producing variation in CVP number, and in particular, for generating CVPs that are multiplied along the midline or anterior-posterior axis. It is interesting that both Fgf10 and Etv5, an indicator of FGF signaling, were expressed along the midline, which correlates with anterior-posterior CVP multiplication in Spry2−/− and Spry1−/−;Spry2−/− mice.
The effect of CVP number on taste is currently unclear. Correlation between taste sensitivity and the number of taste buds within the CVP has been previously reported [48]. However, because there is variation in the number of taste buds per CVP, as well as in the number of taste cells within each taste bud [49], there is no clear indication of what the number of CVPs reveals about taste preferences. Whether the variation in mammalian CVP number provides evolutionary advantages in terms of identification of nutritious substances and detection and avoidance of potentially toxic ones remains to be elucidated.
In 1950, Spuhler [42] postulated that the variation in CVP number observed in humans was likely due to genetic factors involving at least 5 multiple alleles. Our studies indicate, for the first time, that the perturbation of a single gene such as Fgf10 or Spry2 may be sufficient to confer the vast genetic variation in mammalian CVP number.
Materials and Methods
Ethics statement
This study was carried out in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. The protocol was approved by the UCSF Institutional Animal Care and Use Committee (Protocol Number: AN084146). All efforts were made to minimize animal suffering.
Animals
Mouse lines carrying null or floxed alleles of Spry1 [31], Spry2 [34], and Fgf10 [50] or β-actin Cre transgenes [51] were maintained and genotyped as described. Spry1−/−;Spry2−/− mutant embryos were generated by crossing β-actin Cre;Spry1+/−;Spry2+/− males with Spry1flox/flox;Spry2flox/flox females (double mutants produced at an expected Mendelian frequency of 1∶4). Tongues and taste papillae from double heterozygous embryos (Spry1+/−;Spry2+/−) were indistinguishable from wild-type CD-1 embryos and served as controls. Mice were mated overnight, and the presence of a vaginal plug indicated embryonic day (E) 0.5.
Histology
Embryonic and adult tongues were fixed overnight in 4% paraformaldehyde at 4°C. For sections, tongues were dehydrated, embedded in paraffin wax, and serially sectioned at 7 µm. Histological sections were stained with haematoxylin and eosin (H&E).
In situ hybridization (ISH)
Whole-mount ISH using digoxigenin-labeled RNA probes was performed according to standard protocols. RNA probes were generated from plasmids containing fragments of Shh, Spry1, Spry2, Spry4, Fgf10, and Sox2 or from PCR-amplified fragments of Wnt10b and Bmp7. A 10 minute 2% H2O2 treatment was performed on tissues E14.5 and older. Stained specimens were incubated overnight in 30% sucrose, embedded in tissue freezing media (Triangle Biomedical Sciences, Durham, N.C.), and cryosectioned using a Leica CM1900 at 18 µm intervals.
Immunohistochemistry (IHC)
Whole-mount IHC was performed according to a published protocol [52]. Rat anti-E-cadherin (1∶1000; Invitrogen Cat. 1300) or mouse anti-β3 tubulin (1∶250; R&D Systems) were applied followed by incubation in goat anti-rat AlexaFluor 555 secondary antibody (1∶250, Invitrogen). For β3-tubulin staining, the Mouse on Mouse kit (Vectastain) was used. Specimens were counterstained with DAPI. For IHC on sectioned embryonic specimens the same procedure was used after a rehydration step. For IHC on adult tongues, tissue was processed according to a published protocol [28] for markers of three types of taste receptor cells: type 1, rabbit anti-NTPdase2 (Nucleoside triphosphate diphosphohydrolase-2, 1∶1000) is the ecto-ATPase of type I cells in taste buds [53]; type II, monoclonal anti-IP3R3 (Receptor for inositol 1,4,5-trisphosphate, 1∶1000; BD Transduction) is a second messenger that mediates the release of intracellular calcium [54]; and type III, rabbit anti-NCAM (Neural cell adhesion molecule, 1∶1000) [55]. Secondary antibody was goat anti-rabbit Alexa-488 (1∶500; Invitrogen); this was applied for 2 hours and counterstained with propidium iodide.
Analyses of placode size, apoptosis, proliferation, and total number of cells
For quantification of placode size, wild-type and Spry2−/− tongues between E13 and E14 were stained with anti-E-Cadherin antibody and the area measured using ImageJ software. Apoptosis in the CV placode at E12.5 was measured using the In situ Cell Death Detection kit (Roche) following manufacturer's protocol. Proliferating cells were identified by anti-PCNA immunofluorescence staining or injection of 1 mg BrdU for 2 hours followed by staining with anti-BrdU antibody (Invitrogen). Anti-PCNA stained and total (i.e. DAPI-stained) number of cells were counted using ImageJ software and presented as a percentage of proliferating cells. The total number of cells in the CV placode was quantified from confocal images of E-cadherin stained sections using Volocity5 software (Improvision).
qPCR
qPCR reactions were performed using the GoTaq qPCR Master Mix (Promega) in a Mastercycler Realplex (Eppendorf). All primers were designed using PerlPrimer3 software [56]; sequences are available upon request. qPCR conditions were as follows: 95°C, 2 minutes; 40 cycles at 95°C,15 seconds; 58°C,15 seconds; 68°C, 20 seconds; followed by a melting curve gradient. Expression levels for the genes of interest were normalized to levels of L19 and are presented as levels relative to wild-type.
Organ culture
Developing tongues at E11.5 and E12.5 were isolated and cultured on 0.45 µm Millicell-HA membranes (Millipore) in F12/DMEM medium (GIBCO/Invitrogen) containing 1% FBS, 2% B27 culture supplement (GIBCO/Invitrogen), and antibiotics. FGF signaling was inhibited by addition of 25 µM SU5402 suspended in DMSO (Calbiochem), and an equivalent volume of DMSO was added to control wells. After 3 days in culture, the tongues were fixed in 4% paraformaldehyde for 2 h and analyzed.
Microscopy
Fluorescent and bright field images were taken using a Leica DM5000B with a Leica DFC500 camera. For confocal images, a Leica SP5 Confocal was used.
Sample size, penetrance, and statistical analysis
Unless otherwise noted, all experiments were performed independently in triplicate on at least three different specimens (N≥3), and when applicable, presented as an average ± standard deviation. Unpaired Student t-test was used to determine p-values and p<0.01 was deemed to be significant. The CVP phenotype observed in Spry2 null mice was 100% penetrant (n>12); the loss of CVP in Fgf10 null mice was observed in 100% of the mice (n = 6), however, there were indications of small trenches or invaginations (although absence of innervation) in 66% of the embryos; the rescue of the single CVP in the double Fgf10;Spry2 null mice was 100% penetrant (n = 4); the presence of multiple (i.e. ≥3) CVPs in Spry1;Spry2 null mice was 86% penetrant (n = 6). The Fisher exact probability test was used to determine the p-values of the tongue culture experiments.
Supporting Information
Zdroje
1. NguyenHMBarlowLA 2010 Differential expression of a BMP4 reporter allele in anterior fungiform versus posterior circumvallate taste buds of mice. BMC Neurosci 11 129
2. TizzanoMDvoryanchikovGBarrowsJKKimSChaudhariN 2008 Expression of Galpha14 in sweet-transducing taste cells of the posterior tongue. BMC Neurosci 9 110
3. KimMRKusakabeYMiuraHShindoYNinomiyaY 2003 Regional expression patterns of taste receptors and gustducin in the mouse tongue. Biochem Biophys Res Commun 312 500 506
4. ZhangCOakleyB 1996 The distribution and origin of keratin 20-containing taste buds in rat and human. Differentiation 61 121 127
5. MistrettaCMLiuHX 2006 Development of fungiform papillae: patterned lingual gustatory organs. Arch Histol Cytol 69 199 208
6. ChuongCMChodankarRWidelitzRBJiangTX 2000 Evo-devo of feathers and scales: building complex epithelial appendages. Curr Opin Genet Dev 10 449 456
7. ChuongCMPatelNLinJJungHSWidelitzRB 2000 Sonic hedgehog signaling pathway in vertebrate epithelial appendage morphogenesis: perspectives in development and evolution. Cell Mol Life Sci 57 1672 1681
8. JungHSOropezaVThesleffI 1999 Shh, Bmp-2, Bmp-4 and Fgf-8 are associated with initiation and patterning of mouse tongue papillae. Mech Dev 81 179 182
9. HallJMBellMLFingerTE 2003 Disruption of sonic hedgehog signaling alters growth and patterning of lingual taste papillae. Dev Biol 255 263 277
10. LiuHXMaccallumDKEdwardsCGaffieldWMistrettaCM 2004 Sonic hedgehog exerts distinct, stage-specific effects on tongue and taste papilla development. Dev Biol 276 280 300
11. MistrettaCMLiuHXGaffieldWMacCallumDK 2003 Cyclopamine and jervine in embryonic rat tongue cultures demonstrate a role for Shh signaling in taste papilla development and patterning: fungiform papillae double in number and form in novel locations in dorsal lingual epithelium. Dev Biol 254 1 18
12. ZhouYLiuHXMistrettaCM 2006 Bone morphogenetic proteins and noggin: inhibiting and inducing fungiform taste papilla development. Dev Biol 297 198 213
13. LiuHXHensonBSZhouYD'SilvaNJMistrettaCM 2008 Fungiform papilla pattern: EGF regulates inter-papilla lingual epithelium and decreases papilla number by means of PI3K/Akt, MEK/ERK, and p38 MAPK signaling. Dev Dyn 237 2378 2393
14. IwatsukiKLiuHXGronderASingerMALaneTF 2007 Wnt signaling interacts with Shh to regulate taste papilla development. Proc Natl Acad Sci U S A 104 2253 2258
15. LiuFThirumangalathuSGallantNMYangSHStoick-CooperCL 2007 Wnt-beta-catenin signaling initiates taste papilla development. Nat Genet 39 106 112
16. BeitesCLHollenbeckPLKimJLovell-BadgeRLanderAD 2009 Follistatin modulates a BMP autoregulatory loop to control the size and patterning of sensory domains in the developing tongue. Development 136 2187 2197
17. NieX 2005 Apoptosis, proliferation and gene expression patterns in mouse developing tongue. Anat Embryol (Berl) 210 125 132
18. DikicIGiordanoS 2003 Negative receptor signalling. Curr Opin Cell Biol 15 128 135
19. GuyGRWongESYusoffPChandramouliSLoTL 2003 Sprouty: how does the branch manager work? J Cell Sci 116 3061 3068
20. KimHJBar-SagiD 2004 Modulation of signalling by Sprouty: a developing story. Nat Rev Mol Cell Biol 5 441 450
21. HacohenNKramerSSutherlandDHiromiYKrasnowMA 1998 sprouty encodes a novel antagonist of FGF signaling that patterns apical branching of the Drosophila airways. Cell 92 253 263
22. de MaximyAANakatakeYMoncadaSItohNThieryJP 1999 Cloning and expression pattern of a mouse homologue of drosophila sprouty in the mouse embryo. Mech Dev 81 213 216
23. MinowadaGJarvisLAChiCLNeubuserASunX 1999 Vertebrate Sprouty genes are induced by FGF signaling and can cause chondrodysplasia when overexpressed. Development 126 4465 4475
24. JitpukdeebodintraSChaiYSneadML 2002 Developmental patterning of the circumvallate papilla. Int J Dev Biol 46 755 763
25. AhnSKChungJLeeSHLeeWS 1996 Prominent pigmented fungiform papillae of the tongue. Cutis 58 410 412
26. OkuboTPevnyLHHoganBL 2006 Sox2 is required for development of taste bud sensory cells. Genes Dev 20 2654 2659
27. KimJYLeeMJChoKWLeeJMKimYJ 2009 Shh and ROCK1 modulate the dynamic epithelial morphogenesis in circumvallate papilla development. Dev Biol 325 273 280
28. ThirumangalathuSHarlowDEDriskellALKrimmRFBarlowLA 2009 Fate mapping of mammalian embryonic taste bud progenitors. Development 136 1519 1528
29. ZhangSLinYItarantaPYagiAVainioS 2001 Expression of Sprouty genes 1, 2 and 4 during mouse organogenesis. Mech Dev 109 367 370
30. RoehlHNusslein-VolhardC 2001 Zebrafish pea3 and erm are general targets of FGF8 signaling. Curr Biol 11 503 507
31. BassonMAAkbulutSWatson-JohnsonJSimonRCarrollTJ 2005 Sprouty1 is a critical regulator of GDNF/RET-mediated kidney induction. Dev Cell 8 229 239
32. KleinODLyonsDBBaloochGMarshallGWBassonMA 2008 An FGF signaling loop sustains the generation of differentiated progeny from stem cells in mouse incisors. Development 135 377 385
33. KleinODMinowadaGPeterkovaRKangasAYuBD 2006 Sprouty genes control diastema tooth development via bidirectional antagonism of epithelial-mesenchymal FGF signaling. Dev Cell 11 181 190
34. ShimKMinowadaGColingDEMartinGR 2005 Sprouty2, a mouse deafness gene, regulates cell fate decisions in the auditory sensory epithelium by antagonizing FGF signaling. Dev Cell 8 553 564
35. MohammadiMMcMahonGSunLTangCHirthP 1997 Structures of the tyrosine kinase domain of fibroblast growth factor receptor in complex with inhibitors. Science 276 955 960
36. WellsKLMouCHeadonDJTuckerAS 2011 Defects and rescue of the minor salivary glands in Eda pathway mutants. Dev Biol 349 137 146
37. BarlowLA 2003 Toward a unified model of vertebrate taste bud development. J Comp Neurol 457 107 110
38. MunnePMFelszeghySJussilaMSuomalainenMThesleffI 2010 Splitting placodes: effects of bone morphogenetic protein and Activin on the patterning and identity of mouse incisors. Evol Dev 12 383 392
39. LeeCCPutnamAJMirantiCKGustafsonMWangLM 2004 Overexpression of sprouty 2 inhibits HGF/SF-mediated cell growth, invasion, migration, and cytokinesis. Oncogene 23 5193 5202
40. PoppletonHMEdwinFJaggarLRayRJohnsonLR 2004 Sprouty regulates cell migration by inhibiting the activation of Rac1 GTPase. Biochem Biophys Res Commun 323 98 103
41. YigzawYCartinLPierreSScholichKPatelTB 2001 The C terminus of sprouty is important for modulation of cellular migration and proliferation. J Biol Chem 276 22742 22747
42. SpuhlerJN 1950 Genetics of three normal morphological variations: Patterns of superficial veins of the anterior thorax, peroneus tertius muscle, and number of vallate papillae. Cold Spring Harb Symp Quant Biol 15 175 189
43. EmuraSOkumuraTChenH 2008 Morphology of the lingual papillae and their connective tissue cores in the cape hyrax 29 34
44. YoshimuraKHamaNShindoJKobayashiKKageyamaI 2008 Light and scanning electron microscopic study on the lingual papillae and their connective tissue cores of the Cape hyrax Procavia capensis. Journal of Anatomy 573 582
45. SharmaRVidyadaranMZulkifliIAzlanJSumitaS 1999 Ecomorphological implications of the microstructures on the tongue of the fawn roundleaf bat, Hipposideros cervinus (Chiroptera: Hipposideridae). Australian Journal of Zoology 405 409
46. SonntagCF 1921 The comparative anatomy of the tongues of the Mammalia. II. Family 1. Simiidae. Proc Zool Soc 1 1 29
47. SonntagCF 1921 The comparative anatomy of the tongues of the Mammalia. V. Lemuroidea and Tarsioidea. Proc Zool Soc 1 741 755
48. MillerIJJrWhitneyG 1989 Sucrose octaacetate-taster mice have more vallate taste buds than non-tasters. Neurosci Lett 100 271 275
49. HosleyMAOakleyB 1987 Postnatal development of the vallate papilla and taste buds in rats. Anat Rec 218 216 222
50. MinHDanilenkoDMScullySABolonBRingBD 1998 Fgf-10 is required for both limb and lung development and exhibits striking functional similarity to Drosophila branchless. Genes Dev 12 3156 3161
51. LewandoskiMMartinGR 1997 Cre-mediated chromosome loss in mice. Nat Genet 17 223 225
52. MetzgerRJKleinODMartinGRKrasnowMA 2008 The branching programme of mouse lung development. Nature 453 745 750
53. BartelDLSullivanSLLavoieEGSevignyJFingerTE 2006 Nucleoside triphosphate diphosphohydrolase-2 is the ecto-ATPase of type I cells in taste buds. J Comp Neurol 497 1 12
54. MiuraHNakayamaAShindoYKusakabeYTomonariH 2007 Expression of gustducin overlaps with that of type III IP3 receptor in taste buds of the rat soft palate. Chem Senses 32 689 696
55. YeeCLYangRBottgerBFingerTEKinnamonJC 2001 "Type III" cells of rat taste buds: immunohistochemical and ultrastructural studies of neuron-specific enolase, protein gene product 9.5, and serotonin. J Comp Neurol 440 97 108
56. RozenSSkaletskyH 2000 Primer3 on the WWW for general users and for biologist programmers. KrawetzSMisenerS Bioinformatics Methods and Protocols: Methods in Molecular Biology Totowa, NJ Humana Press 365 386
Štítky
Genetika Reprodukční medicína
Článek Local Absence of Secondary Structure Permits Translation of mRNAs that Lack Ribosome-Binding SitesČlánek Independent Chromatin Binding of ARGONAUTE4 and SPT5L/KTF1 Mediates Transcriptional Gene SilencingČlánek Trade-Off between Bile Resistance and Nutritional Competence Drives Diversification in the Mouse GutČlánek Mammalian BTBD12 (SLX4) Protects against Genomic Instability during Mammalian SpermatogenesisČlánek Identification of Nine Novel Loci Associated with White Blood Cell Subtypes in a Japanese PopulationČlánek Differential Effects of and Risk Variants on Association with Diabetic ESRD in African AmericansČlánek Dynamic Chromatin Localization of Sirt6 Shapes Stress- and Aging-Related Transcriptional Networks
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2011 Číslo 6
-
Všechny články tohoto čísla
- Local Absence of Secondary Structure Permits Translation of mRNAs that Lack Ribosome-Binding Sites
- Statistical Inference on the Mechanisms of Genome Evolution
- Revisiting Heterochromatin in Embryonic Stem Cells
- A Two-Stage Meta-Analysis Identifies Several New Loci for Parkinson's Disease
- Identification of a Sudden Cardiac Death Susceptibility Locus at 2q24.2 through Genome-Wide Association in European Ancestry Individuals
- Genomic Prevalence of Heterochromatic H3K9me2 and Transcription Do Not Discriminate Pluripotent from Terminally Differentiated Cells
- Epistasis between Beneficial Mutations and the Phenotype-to-Fitness Map for a ssDNA Virus
- Recurrent Chromosome 16p13.1 Duplications Are a Risk Factor for Aortic Dissections
- Telomere DNA Deficiency Is Associated with Development of Human Embryonic Aneuploidy
- Genome-Wide Association Study of White Blood Cell Count in 16,388 African Americans: the Continental Origins and Genetic Epidemiology Network (COGENT)
- Unexpected Role for DNA Polymerase I As a Source of Genetic Variability
- Transportin-SR Is Required for Proper Splicing of Genes and Plant Immunity
- How Chromatin Is Remodelled during DNA Repair of UV-Induced DNA Damage in
- Independent Chromatin Binding of ARGONAUTE4 and SPT5L/KTF1 Mediates Transcriptional Gene Silencing
- Two Evolutionary Histories in the Genome of Rice: the Roles of Domestication Genes
- Natural Allelic Variation Defines a Role for : Trichome Cell Fate Determination
- Multiple Common Susceptibility Variants near BMP Pathway Loci , , and Explain Part of the Missing Heritability of Colorectal Cancer
- Pathogenic Mechanism of the FIG4 Mutation Responsible for Charcot-Marie-Tooth Disease CMT4J
- A Functional Variant in Promoter Modulates Its Expression and Confers Disease Risk for Systemic Lupus Erythematosus
- Drift and Genome Complexity Revisited
- Chromosomal Macrodomains and Associated Proteins: Implications for DNA Organization and Replication in Gram Negative Bacteria
- Trade-Off between Bile Resistance and Nutritional Competence Drives Diversification in the Mouse Gut
- Pathways of Distinction Analysis: A New Technique for Multi–SNP Analysis of GWAS Data
- Web-Based Genome-Wide Association Study Identifies Two Novel Loci and a Substantial Genetic Component for Parkinson's Disease
- Chk2 and p53 Are Haploinsufficient with Dependent and Independent Functions to Eliminate Cells after Telomere Loss
- Exome Sequencing Identifies Mutations in High Myopia
- Distinct Functional Constraints Partition Sequence Conservation in a -Regulatory Element
- CorE from Is a Copper-Dependent RNA Polymerase Sigma Factor
- A Single Sex Pheromone Receptor Determines Chemical Response Specificity of Sexual Behavior in the Silkmoth
- FGF Signaling Regulates the Number of Posterior Taste Papillae by Controlling Progenitor Field Size
- Maps of Open Chromatin Guide the Functional Follow-Up of Genome-Wide Association Signals: Application to Hematological Traits
- Increased Susceptibility to Cortical Spreading Depression in the Mouse Model of Familial Hemiplegic Migraine Type 2
- Differential Gene Expression and Epiregulation of Alpha Zein Gene Copies in Maize Haplotypes
- Parallel Adaptive Divergence among Geographically Diverse Human Populations
- Genetic Analysis of Genome-Scale Recombination Rate Evolution in House Mice
- Mechanisms for the Evolution of a Derived Function in the Ancestral Glucocorticoid Receptor
- Mammalian BTBD12 (SLX4) Protects against Genomic Instability during Mammalian Spermatogenesis
- Interferon Regulatory Factor 8 Regulates Pathways for Antigen Presentation in Myeloid Cells and during Tuberculosis
- High-Resolution Analysis of Parent-of-Origin Allelic Expression in the Arabidopsis Endosperm
- Specific SKN-1/Nrf Stress Responses to Perturbations in Translation Elongation and Proteasome Activity
- Graded Nodal/Activin Signaling Titrates Conversion of Quantitative Phospho-Smad2 Levels into Qualitative Embryonic Stem Cell Fate Decisions
- Genome-Wide Analysis Reveals PADI4 Cooperates with Elk-1 to Activate Expression in Breast Cancer Cells
- Trait Variation in Yeast Is Defined by Population History
- Meiosis-Specific Loading of the Centromere-Specific Histone CENH3 in
- A Genome-Wide Survey of Imprinted Genes in Rice Seeds Reveals Imprinting Primarily Occurs in the Endosperm
- Multiple Regulatory Mechanisms to Inhibit Untimely Initiation of DNA Replication Are Important for Stable Genome Maintenance
- SIRT1 Promotes N-Myc Oncogenesis through a Positive Feedback Loop Involving the Effects of MKP3 and ERK on N-Myc Protein Stability
- Bacteriophage Crosstalk: Coordination of Prophage Induction by Trans-Acting Antirepressors
- Role of the Single-Stranded DNA–Binding Protein SsbB in Pneumococcal Transformation: Maintenance of a Reservoir for Genetic Plasticity
- Genomic Convergence among ERRα, PROX1, and BMAL1 in the Control of Metabolic Clock Outputs
- Genome-Wide Association of Bipolar Disorder Suggests an Enrichment of Replicable Associations in Regions near Genes
- Identification of Nine Novel Loci Associated with White Blood Cell Subtypes in a Japanese Population
- DNA Ligase III Promotes Alternative Nonhomologous End-Joining during Chromosomal Translocation Formation
- Differential Effects of and Risk Variants on Association with Diabetic ESRD in African Americans
- Finished Genome of the Fungal Wheat Pathogen Reveals Dispensome Structure, Chromosome Plasticity, and Stealth Pathogenesis
- Dynamic Chromatin Localization of Sirt6 Shapes Stress- and Aging-Related Transcriptional Networks
- Extracellular Matrix Dynamics in Hepatocarcinogenesis: a Comparative Proteomics Study of Transgenic and Null Mouse Models
- Integrating 5-Hydroxymethylcytosine into the Epigenomic Landscape of Human Embryonic Stem Cells
- Vive La Différence: An Interview with Catherine Dulac
- Multiple Loci Are Associated with White Blood Cell Phenotypes
- Nuclear Accumulation of Stress Response mRNAs Contributes to the Neurodegeneration Caused by Fragile X Premutation rCGG Repeats
- A New Mutation Affecting FRQ-Less Rhythms in the Circadian System of
- Cryptic Transcription Mediates Repression of Subtelomeric Metal Homeostasis Genes
- A New Isoform of the Histone Demethylase JMJD2A/KDM4A Is Required for Skeletal Muscle Differentiation
- Genetic Determinants of Lipid Traits in Diverse Populations from the Population Architecture using Genomics and Epidemiology (PAGE) Study
- A Genome-Wide RNAi Screen for Factors Involved in Neuronal Specification in
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Recurrent Chromosome 16p13.1 Duplications Are a Risk Factor for Aortic Dissections
- Statistical Inference on the Mechanisms of Genome Evolution
- Genome-Wide Association Study of White Blood Cell Count in 16,388 African Americans: the Continental Origins and Genetic Epidemiology Network (COGENT)
- Chromosomal Macrodomains and Associated Proteins: Implications for DNA Organization and Replication in Gram Negative Bacteria
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



