-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaA Single Element Maintains Repression of the Key Developmental Regulator
In development, lineage-restricted transcription factors simultaneously promote differentiation while repressing alternative fates. Molecular dissection of this process has been challenging as transcription factor loci are regulated by many trans-acting factors functioning through dispersed cis elements. It is not understood whether these elements function collectively to confer transcriptional regulation, or individually to control specific aspects of activation or repression, such as initiation versus maintenance. Here, we have analyzed cis element regulation of the critical hematopoietic factor Gata2, which is expressed in early precursors and repressed as GATA-1 levels rise during terminal differentiation. We engineered mice lacking a single cis element −1.8 kb upstream of the Gata2 transcriptional start site. Although Gata2 is normally repressed in late-stage erythroblasts, the −1.8 kb mutation unexpectedly resulted in reactivated Gata2 transcription, blocked differentiation, and an aberrant lineage-specific gene expression pattern. Our findings demonstrate that the −1.8 kb site selectively maintains repression, confers a specific histone modification pattern and expels RNA Polymerase II from the locus. These studies reveal how an individual cis element establishes a normal developmental program via regulating specific steps in the mechanism by which a critical transcription factor is repressed.
Published in the journal: . PLoS Genet 6(9): e32767. doi:10.1371/journal.pgen.1001103
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1001103Summary
In development, lineage-restricted transcription factors simultaneously promote differentiation while repressing alternative fates. Molecular dissection of this process has been challenging as transcription factor loci are regulated by many trans-acting factors functioning through dispersed cis elements. It is not understood whether these elements function collectively to confer transcriptional regulation, or individually to control specific aspects of activation or repression, such as initiation versus maintenance. Here, we have analyzed cis element regulation of the critical hematopoietic factor Gata2, which is expressed in early precursors and repressed as GATA-1 levels rise during terminal differentiation. We engineered mice lacking a single cis element −1.8 kb upstream of the Gata2 transcriptional start site. Although Gata2 is normally repressed in late-stage erythroblasts, the −1.8 kb mutation unexpectedly resulted in reactivated Gata2 transcription, blocked differentiation, and an aberrant lineage-specific gene expression pattern. Our findings demonstrate that the −1.8 kb site selectively maintains repression, confers a specific histone modification pattern and expels RNA Polymerase II from the locus. These studies reveal how an individual cis element establishes a normal developmental program via regulating specific steps in the mechanism by which a critical transcription factor is repressed.
Introduction
Metazoan development is characterized by complex transcriptional programs specified by gene regulatory networks [1], [2]. Transcription factors in these networks occupy specific cis elements at target gene loci where they modulate chromatin remodeling and modification, and thereby transcription. The covalent modification of histones to yield specific histone marks promotes either the activation or repression of transcription [3]. Models of gene regulation have led to an attractive paradigm in which repression occurs in sequential stages of increasing stability [4]. While transcription factors bind and recruit chromatin-modifying and remodeling proteins, the relative contribution of individual cis elements residing within clusters of cis elements to the transcriptional control of endogenous loci is incompletely understood.
GATA factor cross-regulation represents an instructive model system for investigating the contribution of individual cis elements to the initiation and maintenance of transcriptional repression. The GATA family of transcription factors plays diverse roles in multiple developmental contexts [5]. GATA factors are often expressed in an overlapping but reciprocal pattern, such that expression of one GATA factor increases as expression of another decreases. For example, GATA-1 directly represses Gata2 transcription via displacing GATA-2 from chromatin sites at its own locus, a process termed a “GATA Switch” [6], [7].
GATA factor function has been extensively studied in the context of hematopoiesis, where GATA-1, GATA-2, and GATA-3 are key regulators. GATA-2 has a broad role in hematopoietic development, as demonstrated by impaired hematopoiesis in Gata2 knock-out mice resulting in lethality during midgestation [8], [9]. GATA-1 is critical for the production of red blood cells and platelets [10], and GATA-3 is required for specification of T cells [11]. Forced expression of GATA-2 blocks erythroid development [12], [13], [14], leading to a model in which GATA-1-mediated repression of Gata2 through specific cis elements is required for differentiation. Genome-wide studies revealed GATA-1 occupancy at only a small subset of cis elements in the genome [15]. These cis elements exist as single or more complex GATA motifs, although the functionality of different permutations of GATA motifs at endogenous loci has not been investigated.
The role of individual GATA-binding sites in gene regulation has been investigated extensively at the Gata2 locus, where several conserved GATA motif-containing regions span approximately 100 kb of the locus [16]. To test whether GATA switch sites function collectively or independently to regulate Gata2 expression, and to investigate the underlying mechanisms, we generated mice lacking one of these regulatory regions residing −1.8 kb upstream of the Gata2 promoter. We find that while this site is not essential for Gata2 expression in hematopoietic progenitors or initiation of Gata2 repression during erythropoiesis, it maintains Gata2 repression in erythroblasts. Molecular analyses demonstrate that loss of the −1.8 kb site reduces GATA-1 binding, allows for increased RNA Polymerase II (Pol II) occupancy at the locus, and results in changes in select histone marks. Further, elimination of the −1.8 kb site dysregulates Gata2 transcriptional control, disrupts the GATA-2-dependent genetic network, and interferes with red blood cell maturation. These results highlight the qualitatively distinct activities of individual cis elements in specific aspects of gene repression during development.
Results
Targeted deletion of the Gata2 −1.8 kb cis element
Previous studies in erythroid cell lines [17]–[22] and transgenic mouse models [23]–[25] have identified five GATA-binding regions upstream and in an intron of the Gata2 locus (Figure 1A). It remains unknown whether these regions function collectively to confer Gata2 transcriptional regulation, or if individual regions function uniquely at specific developmental stages and/or in select cell types. The site at −1.8 kb is of considerable interest, since it possesses strong GATA-2 binding activity that is lost upon repression [17]. Thus, we reasoned that removal of this site would phenocopy GATA-2-deficiency. As definitive analysis of cis element function requires genetic ablation of endogenous loci, we generated a mouse strain lacking the palindromic GATA-binding site 1.8 kb upstream of the Gata2 transcriptional start site (Δ-1.8 allele) (Figures 1B, S1). Mice homozygous for the Δ-1.8 allele were born at expected Mendelian ratios, as assessed by PCR genotyping (Figure 1C.), implying that embryonic development was largely unaffected. Morphologically, E12.5 wild-type and mutant embryos were similar (Figure S2A), and adult mutant mice lacked gross abnormalities (data not shown).
Fig. 1. Generation of mice lacking the −1.8 kb site of the Gata2 locus. 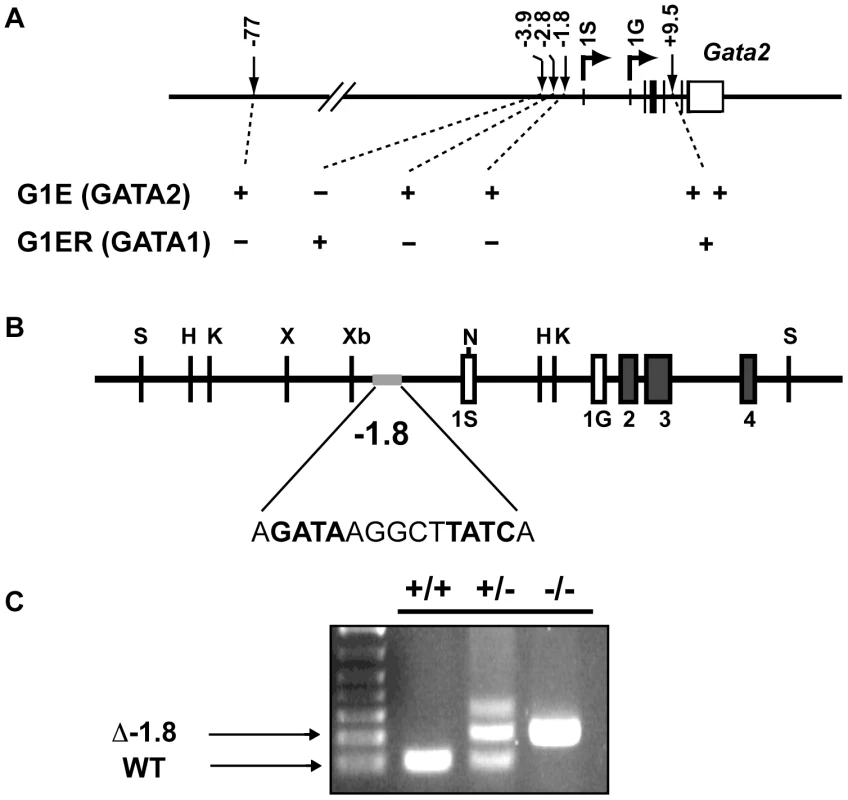
Diagram of GATA binding sites involved in the regulation of hematopoietic expression of Gata2 from its two alternate first exons, 1S and 1G, and their functional categories as defined by GATA-factor binding in G1E cells expressing only GATA2 and G1ER cells expressing only GATA1 (A). The region of the Gata2 locus targeted in generation of mouse strain deficient for a palindromic GATA binding site 1.8 kb upstream of the Gata2 transcriptional start site, referred to as the Δ-1.8 allele (B). PCR genotyping of mice deficient for the −1.8 GATA-binding site and littermate controls (C). We analyzed fetal liver erythropoiesis in Δ-1.8 mice for alterations in Gata2 expression. Using fluorescence-activated cell sorting with the erythroid markers CD71 and Ter119 [26], we isolated cells from Stages I, II, III, and IV, corresponding to CD71loTer119− (committed erythroid progenitors, Stage I), CD71hiTer119− (proerythroblasts, Stage II), CD71hiTer119+ (basophilic erythroblasts, Stage III), and CD71loTer119+ (late erythroblasts, Stage IV) (Figure 2A). In wild-type mice, Gata2 was most highly expressed in Stage I progenitors, after which it was repressed in Stages II, III, and IV (Figure 2B). Gata2 expression was modestly increased in Stage IV, to about one fifth of that observed in Stage I. In Δ-1.8 mice, Gata2 expression was normal in Stage I, and decreased normally in Stage II and III, indicating that the −1.8 kb site is not required for initiation of GATA-1-mediated repression. However, Gata2 expression was significantly elevated in Stage IV cells from the Δ-1.8 versus wild-type mice (p≤0.05) (Figure 2B). Thus, the −1.8 kb site is selectively required to maintain Gata2 repression in Stage IV erythroblasts.
Fig. 2. Alterations in Gata2 expression in late erythroid development in Δ-1.8 mice. 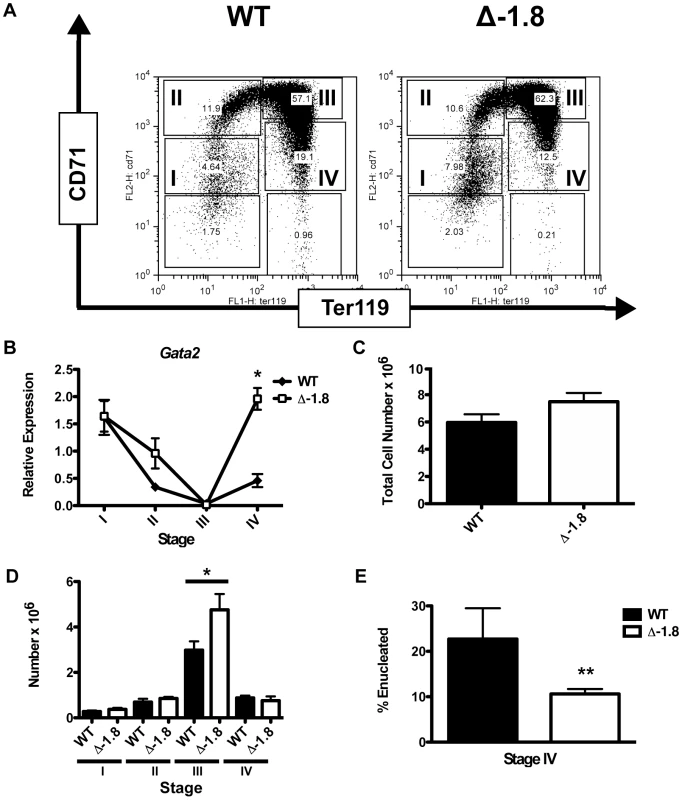
Representative FACS plots of E12.5 fetal liver cells stained for CD71 and Ter119 from wild-type and Δ-1.8 embryos (A). Gata2 expression in Stage I through Stage IV sorted erythroid cells from wild-type and Δ-1.8 embryos normalized to β-actin expression (B). Total fetal liver cellularity in wild-type and Δ-1.8 embryos (C). Absolute number of cells in Stage I through Stage IV in fetal liver cells from wild-type (n = 4) and Δ-1.8 embryos (n = 3) (D). Percentage of Stage IV cells that are enucleated (negative for Draq5 staining) from wild-type (n = 8) and Δ-1.8 (n = 8) embryos (E). Impaired erythroid development in Δ-1.8 mutant mice
To determine if GATA-2 derepression has functional consequences in erythropoiesis, we analyzed erythroid cells in E12.5 fetal livers from wild-type and Δ-1.8 mice. Total cell numbers from wild-type and mutant fetal livers were similar (Figure 2C). Cytospins of peripheral blood and fetal liver cells from wild-type and mutant E12.5 embryos had similar appearance upon May-Gruenwald-Giemsa staining (Figure S2B). At this stage in development, most of the embryonic blood is comprised of primitive erythroid cells. However, some enucleated definitive cells were detected in both wild-type and Δ-1.8 embryos (Figure S2B). Hematopoietic colony assays from wild-type and Δ-1.8 E14.5 fetal livers revealed that the total number of colonies and lineage distribution of colony types (representing multipotential and lineage-restricted progenitors) were similar (Figure S2C). Examination of cells spanning different stages of erythroid development revealed no difference in the absolute number of Stage I, Stage II or Stage IV erythroid progenitors. However, the absolute number of Stage III erythroid progenitors was increased significantly (p≤0.05) in the Δ-1.8 mice (Figure 2D). These results demonstrate that at E12.5, Stage III progenitors from Δ-1.8 mice expand relative to both their precursors and progeny, implying a block in the Stage III to Stage IV transition. The increased number of Stage III progenitors is in accordance with other models of ineffective erythropoiesis, in which impairment of erythroid cell maturation is accompanied by a compensatory increase in earlier red blood cell precursors [27]. The timing of this block corresponds to the stage at which Gata2 is reactivated (Figure 2B), indicating that Gata2 dysregulation perturbs erythroid development. To examine this further, we utilized red blood cell enucleation as a cellular read-out of erythroid differentiation. Enucleation was measured using Draq5 to quantitate DNA content in Stage IV cells from wild-type and mutant embryos (a representative FACS plot is shown in Figure S2D). Stage IV cells in Δ-1.8 embryos contained a significantly reduced (>2-fold, p≤0.05) proportion of enucleated cells compared to those from wild-type, demonstrating that mutant cells fail to differentiate efficiently upon reactivation of Gata2 expression (Figure 2E).
Gata2 reactivation in Δ-1.8 mice dysregulates GATA factor target genes
We reasoned that aberrant expression of GATA-2 target genes in Δ-1.8 mice might underlie the block in the transition from early to late erythroblasts. Increased Gata2 expression could reactivate GATA-2 target genes expressed in early erythropoiesis, including those associated with proliferation, at a stage in which cells should exit the cell cycle. Alternatively, increased Gata2 expression could aberrantly repress late erythroid genes necessary for efficient differentiation. Finally, abnormal reactivation of Gata2 expression in cells expressing GATA-1 and other transcription factors involved in specifying alternate lineage programs could lead to the aberrant transcription of non-erythroid genes. To distinguish among these possibilities, we quantified gene expression in fetal liver erythroid cells from E12.5 mice. Several gene expression changes were apparent in Stage IV erythroblasts (Figure 3A). Expression of Gata1 and Eraf, a globin chain stabilizing protein, were reduced by ∼40% (p≤0.01) and ∼50% (p≤0.05), respectively, in the late erythroblasts of Δ-1.8 versus wild-type mice. In contrast to Gata1 and Eraf, most late erythroid genes examined, including the transcription factors Scl, Eklf, and the heme synthesis enzyme Alas2, were expressed at similar levels, indicating that erythroid genes are differentially sensitive to Gata2 reactivation. Whereas expression of β-like globin genes (Hbb-y, Hbb-bh1, Hbb-b1) was normal (Figure 3B), expression of α-globin (Hba-a1) was reduced by 50% (p≤0.05) and ζ-globin (Hba-x) was increased by 2-fold (p≤0.05) (Figure 3B). We also examined two genes expressed early in erythropoiesis. Both cMyb and the established GATA-2 target cKit were upregulated 4-fold (p≤0.05) (Figure 3C). In mast cells and megakaryocytes, GATA-2 is expressed in combination with other transcription factors including SCL and GATA-1. As GATA-2 is aberrantly coexpressed with these factors in the Δ-1.8 erythroblasts, we examined select GATA-2 target genes from the mast cell and megakaryocyte lineages in our wild-type and mutant erythroblasts. Cpa3, active in mast cells, and cMpl, expressed in megakaryocytes, were upregulated 4 - and 2-fold, respectively, in mutant versus wild-type erythroblasts (p≤0.05) (Figure 3D). These results indicate that Gata2 reactivation is coupled with aberrant GATA-2 target gene expression. Given the dysregulation of genes associated with early progenitor proliferation, erythroid maturation, and alternate lineage fate, it is likely that these factors contribute in aggregate to the block in erythroid development.
Fig. 3. Disrupted gene expression in Δ-1.8 erythroblasts. 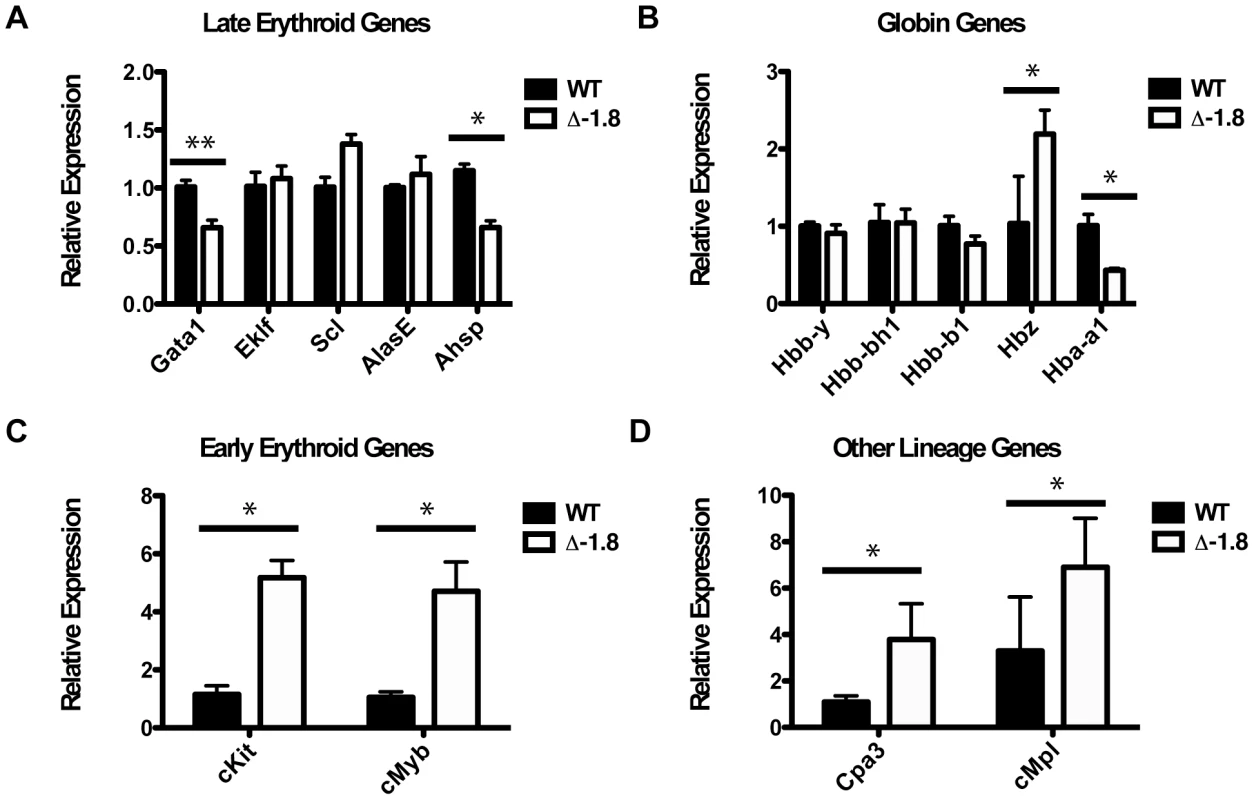
Relative expression of genes in Stage IV sorted erythroid cells from wild-type and Δ-1.8 embryos; genes expressed in late erythropoiesis (A), globin genes (B), genes expressed in early erythropoiesis (C), and genes expressed in other lineages (D). Defective stress erythropoiesis in Δ-1.8 mice
In contrast to the E12.5 fetal liver, erythroid progenitors isolated from the bone marrow of adult wild-type and Δ-1.8 mutant mice (Stage II–IV) did not reveal differences in Gata2 expression (data not shown), indicating that Gata2 transcription is differentially regulated during fetal and adult erythropoiesis. Adult erythropoiesis has several unique attributes relative to the fetal process, including differences in proliferative capacity and rate of transit through the differentiation program [28], [29]. Such differences might explain the ontogenic specificity of Gata2 reactivation. We reasoned that stress erythropoiesis in the adult, which resembles fetal liver erythropoiesis [28]–[32], might shift the regulation of Gata2 expression to a state mimicking that in the fetus. To establish stress erythropoietic conditions, peripheral anemia was induced through phenylhydrazine-mediated red blood cell lysis. Examination of erythropoietic recovery in Δ-1.8 mice revealed no differences in hematocrit, implying that there is no deficiency in recovery from acute anemia in these mice (data not shown). However, analysis of erythropoietic progenitor production in the bone marrow during recovery revealed that the absolute number of Stage III erythroid progenitors was significantly increased in Δ-1.8 mice (p≤0.05) (Figure 4A), while the number of Stage IV erythroid progenitors was similar, again indicating a block in the transition from Stage III to Stage IV. The increased number of Stage III cells is likely an indirect effect due to the increased sensitivity of Δ-1.8 mice to stress-induced ineffective erythropoiesis [27]. Expression analysis of sorted populations from the bone marrow of these mice showed that Gata2 transcription is increased significantly (p≤0.05) in Stage IV cells from Δ-1.8 mice (Figure 4B). These results mimic those obtained with E12.5 fetal liver (Figure 2B,D), indicating that the −1.8 kb site controls Gata2 expression in both stress and fetal erythropoiesis.
Fig. 4. Gata2 dysregulation during stress erythropoiesis in Δ-1.8 mice. 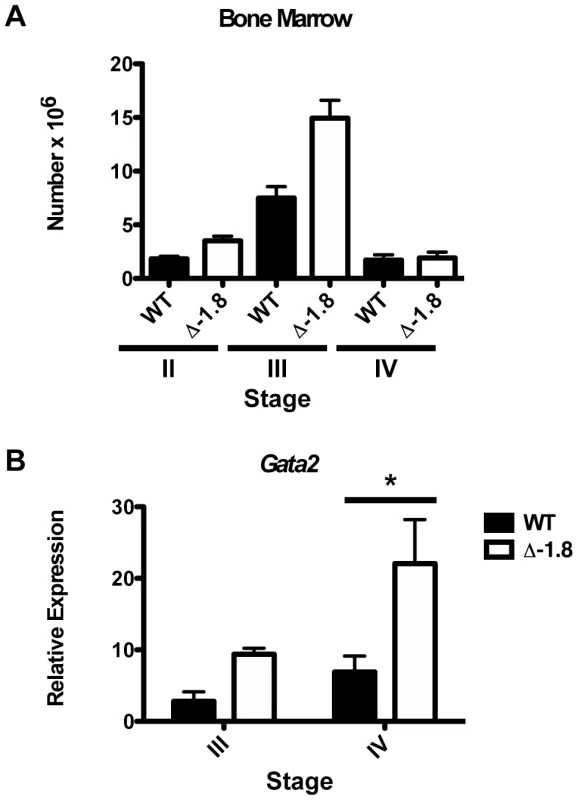
Total number of erythroid progenitors in the bone marrow of wild-type (n = 4) and Δ-1.8 (n = 6) mice 4 days post-phenylhydrazine injection (A). Gata2 expression in Stage III and Stage IV erythroid cells sorted from the bone marrow of phenylhydrazine-treated wild-type and Δ-1.8 mice (B). Δ-1.8 cells possess altered nucleoprotein architecture of the Gata2 locus
Gata2 is transcribed from two alternate promoters, termed 1S and 1G, leading to two transcripts with different first exons [33]. To determine whether the loss of Gata2 repression in Δ-1.8 erythroid cells (Figure 5A) reflects increased transcripts derived from one or both of the promoters, we used primers specific for mature forms of the 1S and 1G transcripts. The majority of Gata2 transcripts expressed in Stage I were derived from the 1G promoter and were repressed in Stage II–IV similarly in wild-type and Δ-1.8 cells (Figure 5B). mRNA expression from the 1S promoter was increased nearly 8-fold (p≤0.05) in Δ-1.8 Stage IV cells relative to Stage I cells. While wild-type cells exhibited increased 1S-derived mRNA at Stage IV relative to Stage I, this increase was significantly smaller (Figure 5C). Quantitation of primary, unspliced transcripts derived from the 1S promoter revealed an even more striking increase in 1S-derived transcript from Δ-1.8 Stage IV cells (∼10-fold relative to Stage I) compared to wild-type cells from the same stage, which did not demonstrate any appreciable increase (p≤0.05) (Figure 5D). Together, these results demonstrate that loss of the −1.8 kb site selectively reactivates transcription from the 1S promoter in erythroid cells.
Fig. 5. Promoter usage in Δ-1.8 Gata2 dysregulation. 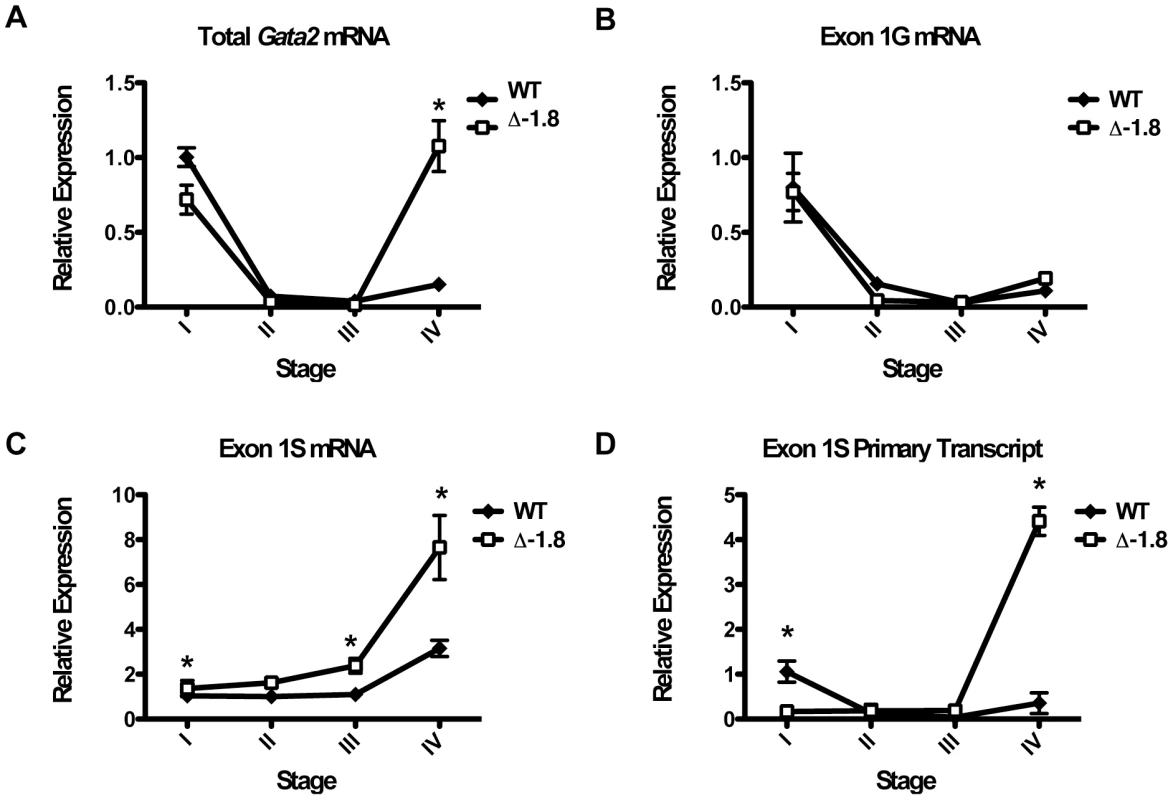
Total Gata2 mRNA (A), mRNA selectively arising from the 1G (B) or the 1S (C) promoter, and primary, unspliced transcript arising from the 1S promoter (D) in Stage I through Stage IV sorted erythroid cells from wild-type and Δ-1.8 embryos at E14.5. As expected, quantitative chromatin immunoprecipitation (ChIP) analysis of E14.5 fetal liver cells demonstrated reduction of GATA-1 occupancy at the −1.75 kb (used as a surrogate for measuring occupancy at the deleted −1.8 kb site) and the −2.8 kb sites (p = 0.057 and 0.058 respectively) and the proximal GATA-binding regions at −3.9 kb, (p≤0.01); occupancy was not significantly altered at the distal −77 and +9.5 kb sites (Figure 6A). ChIP analysis of Pol II demonstrated significantly increased occupancy at all sites examined upon mutation of the −1.8 kb site, with notable increases at the −77 kb enhancer (p≤0.01), the −1.75 kb site (p≤0.01) and the 1G promoter (p≤0.01) (Figure 6B). Importantly, Pol II occupancy of a distant gene (RPII215) did not change upon loss of the −1.8 kb site (Figure 6B), providing evidence for locus specificity. ChIP analysis of GATA-2 occupancy yielded signals near background levels, consistent with GATA-2 expression being below the limit of detection in this assay (data not shown). Average preimmune values for the wild-type and Δ-1.8 cells were 0.0018±0.00027 and 0.0041±0.0017, respectively.
Fig. 6. Loss of the −1.8 kb site leads to increased RNA Pol II occupancy of the Gata2 locus. 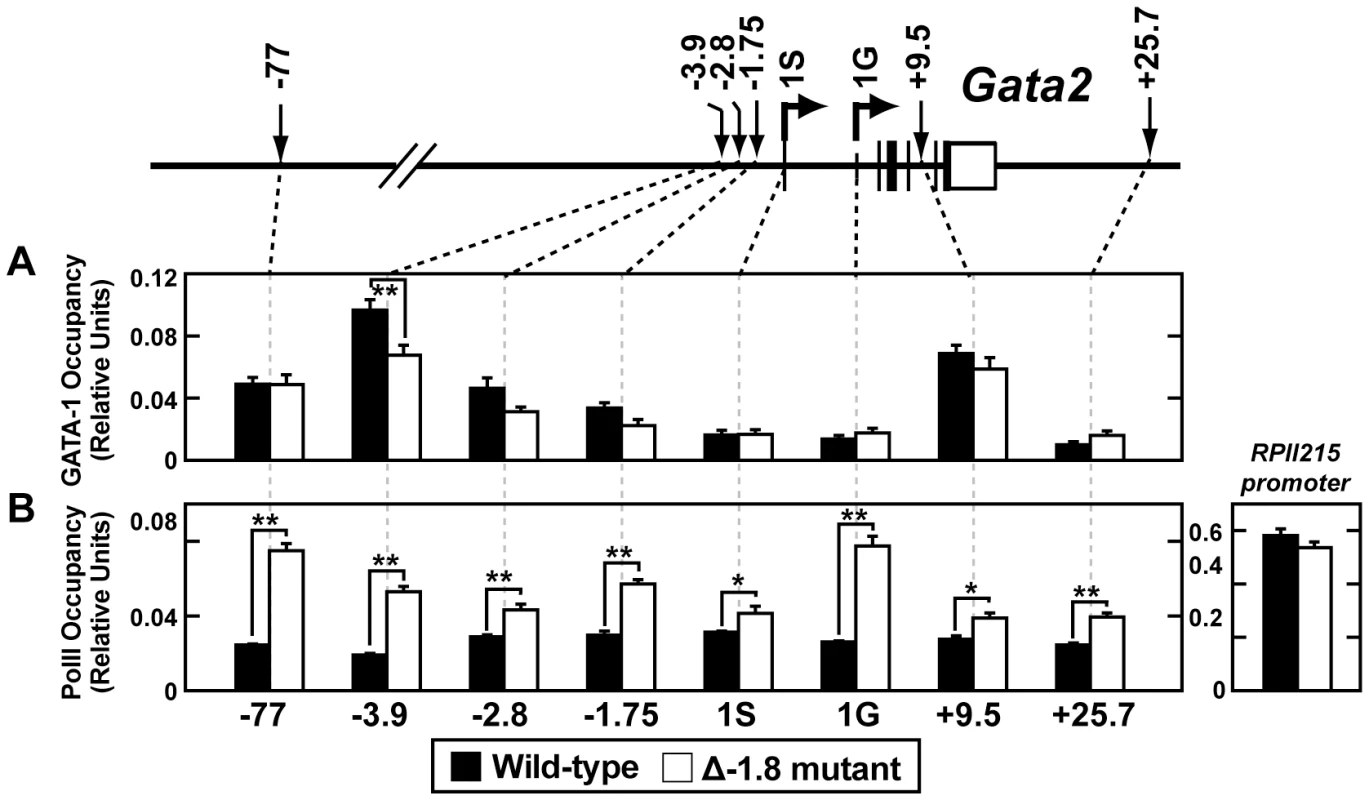
Quantitative ChIP analysis across the Gata2 locus using antibodies to GATA-1 (A) and RNA Pol II (B) in whole fetal liver cells from wild-type and Δ-1.8 embryos at E14.5. Calculations were derived using percentage of input and were normalized using relative units which were determined by defining 9% input sample as 1.0. To analyze histone modifications within the erythroid populations in which we observed an altered phenotype on removal of the −1.8 kb site (Figure 2), we performed quantitative ChIP on sorted fetal liver Stage III and IV cells. We quantitated dimeH3K4 and trimeH3K27, two marks shown to be associated with repression at the Gata2 locus [17], [21], [34]. dimeH3K4 was significantly reduced at the −1.75 kb site, neighboring proximal regulatory regions, and the 1S promoter in both Stage III and Stage IV Δ-1.8 cells (p≤0.05) (Figure 7A). The repressive mark trimeH3K27 was decreased to a small extent at the promoters in Stage III Δ-1.8 cells (p≤0.05) (Figure 7B). Preimmune values were similar between wild-type and −1.8 samples (Figure 7C). These results in primary erythroid progenitors provide direct evidence that the −1.8 kb cis element contributes to the maintenance of the dimeH3K4 mark in erythroid cells.
Fig. 7. Loss of the −1.8 kb site alters dimeH3K4 and trimeH3K27 marks in Stage III and IV erythroblasts. 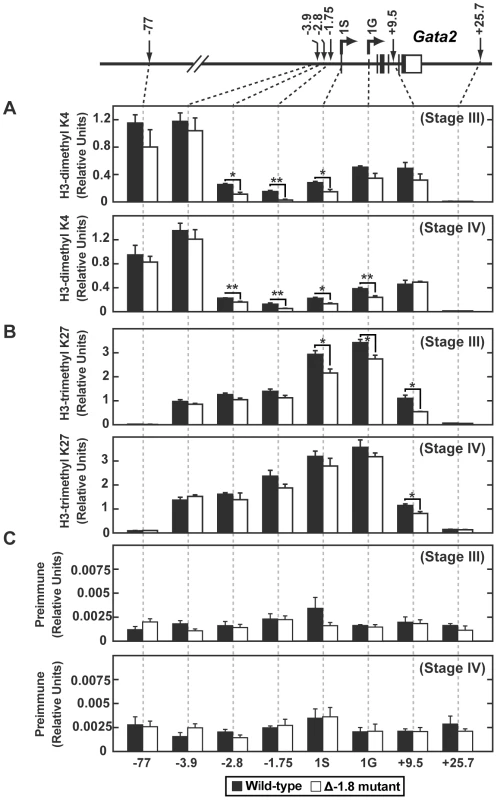
Quantitative ChIP analysis across the Gata2 locus using antibodies to dimeH3K4 (A) and trimeH3K27 (B) and preimmune (C) in Stage III and Stage IV cells sorted from wild-type and Δ-1.8 embryos at E14.5. Calculations were derived using percentage of input and were normalized using relative units which were determined by defining 9% input sample as 1.0. Contribution of the dimethylH3K4 modification to transcriptional regulation is incompletely understood [35]–[37]. By contrast, the trimethylH3K4 mark is thought to play a critical role in promoting gene activation [38], [39]. Also, recent attention has been focused on the monomethylH3K4 mark as an important regulator of enhancer elements [38], [40]. We reasoned that loss of dimethylH3K4 might play an indirect role by providing a substrate for increases in the mono - or trimethyl forms of H3K4. However, ChIP using E14.5 whole fetal liver cells revealed that the levels of trimeH3K4 were unchanged at all sites examined (Figure S3A). Even more strikingly, the levels of monomeH3K4 were reduced at the −2.8 (p≤0.05) and −1.75 kb sites (p≤0.01), as well as the 1G promoter (p≤0.05) (Figure S3B), similar to the reduction in dimeH3K4 observed in whole fetal liver cells (data not shown) and in sorted cells (Figure 7). Total H3 and preimmune values for ChIP using whole fetal liver cells were similar between wild-type and Δ-1.8 samples (data not shown).
These data indicate that loss of GATA-1 binding at the deleted −1.8 kb cis element leads to decreased GATA-1 occupancy at sites up to several kilobases away, reductions in dimeH3K4 and monomeH4K4 marks in the regulatory regions, and increased RNA Pol II occupancy. We propose a model in which this altered nucleoprotein structure favors a transcriptionally active locus, thereby permitting Gata2 reactivation.
CpG island methylation of the Gata2 1S promoter is independent of the −1.8 site
The Gata2 locus contains four CpG islands [17] located at the −2.8 kb GATA-binding region, both the 1S and 1G promoters, and an unclassified region between these promoters (Figure 8A). Stable repression at loci characterized by CpG-rich promoters is thought to depend, in part, upon methylation of these promoters [4], [41]. In addition, tissue specific gene silencing of Gata2 has been correlated with promoter methylation in some tissues [42], [43]. Thus, we tested whether methylation of the 1S promoter is important for stable repression in erythroid cells and whether the −1.8 kb cis element maintains repression through such a mechanism. Bisulfite sequencing was utilized to quantitate promoter methylation of a 3′ section of the 1S CpG island within sorted populations of Stage II–IV erythroid progenitors from wild-type and mutant mice. In wild-type mice, the CpG island located at the Gata2 1S promoter was largely unmethylated in Stage II, Stage III, and Stage IV progenitors, with an average methylation of 5.2%, 8.9%, and 7.1%, respectively (Figure 8B). As no specific residues were hypermethylated (Figure S3C), these data imply that methylation of the 1S CpG island is not important for maintenance of repression in these cells. In Δ-1.8 mice, the 1S CpG island displayed similar levels of methylation in Stage II, Stage III, and Stage IV progenitors (5.9%, 8.2%, and 7.1%, respectively, Figure 8B). Thus, the stable repression of Gata2 does not require DNA methylation of the 1S CpG island, and the −1.8 kb site maintains repression in Stage IV erythroblasts through other mechanisms including regulation of transcription factor occupancy, histone modifications, and Pol II access.
Fig. 8. Gata2 promoter methylation does not require the −1.8 kb site. 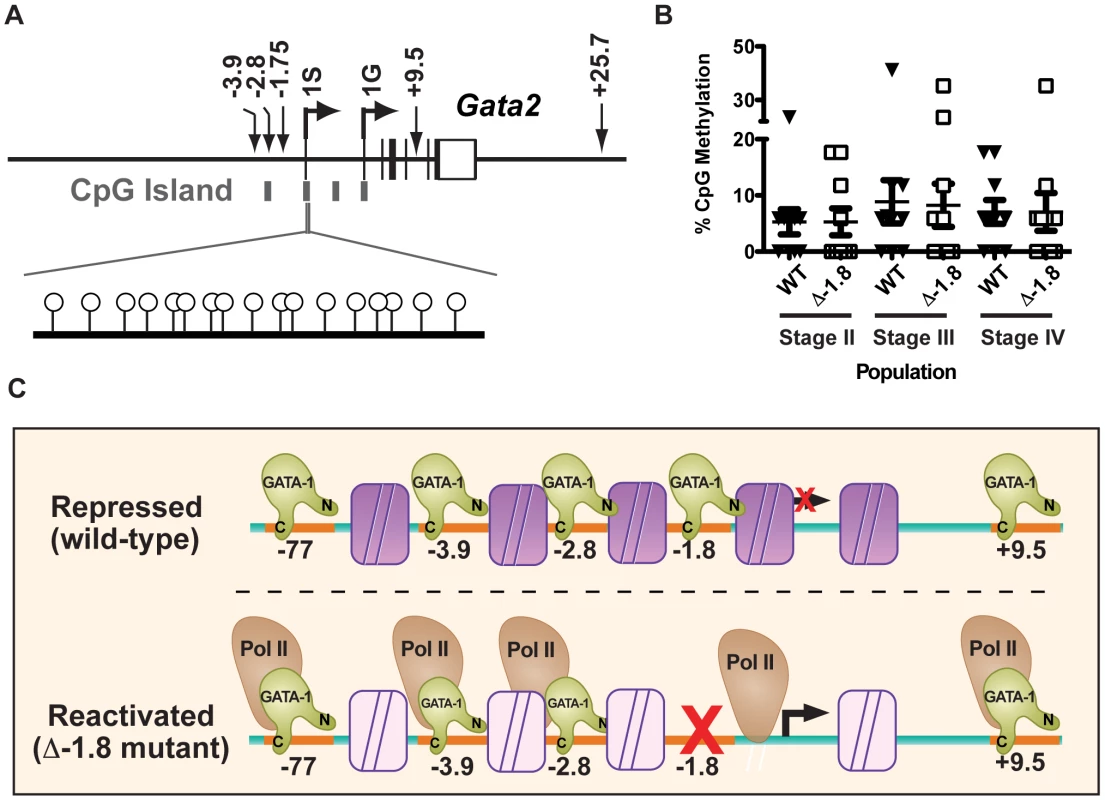
CpG island locations within the Gata2 locus (A). Percentage of CpG methylation within the 3′ region of the 1S promoter in Stage II, III and IV erythroid cells from E14.5 wild-type and Δ-1.8 fetal liver (B). A model of the maintenance of Gata2 locus repression, mediated through maximal GATA-1 binding, repressive histone marks (purple), and complete Pol II expulsion. In the absence of the −1.8 kb site, GATA-1 binding is diminished, leading to a reduction in repressive histone marks (transparent) and allowing Pol II occupancy (C). Discussion
We have described a loss of function strategy in mice to establish definitively whether one of the cis elements previously implicated in the control of Gata2 [17], [19], [21] functions independently or collectively with other cis elements to regulate Gata2 transcription in vivo. Unexpectedly, the endogenous −1.8 kb site is dispensable for activation of Gata2 and the initiation of repression, but instead selectively maintains Gata2 repression in terminally differentiating cells. Deletion of the −1.8 kb site reactivates Gata2 expression resulting in an erythroid maturation block, likely due to improper regulation of the reciprocally controlled Gata1, genes involved in globin synthesis, genes expressed earlier in erythropoiesis, and genes associated with other hematopoietic lineages.
While one or more additional GATA-binding sites in the locus must contribute to the initiation of repression, we propose that maintenance of repression is mediated through GATA-1 binding at the −1.8 kb site of the Gata2 1S promoter. In a wild-type setting, transcription factor binding and histone modifications lead to Pol II expulsion in a locus-wide manner to establish stable repression (Figure 8C). In the absence of the −1.8 kb cis element, GATA-1 occupancy is lost at this site. Our results demonstrate that locus-wide Pol II expulsion requires maximal GATA-binding at the 5′ proximal regulatory regions, highlighting a critical role for the −1.8 kb site in regulating Pol II occupancy. The loss of GATA-1 occupancy in the absence of the −1.8 kb site results in a reduction in one of the marks associated with repression at this locus, dimeH3K4, while having minimal effects on another repressive mark, trimeH3K27. Intriguingly, dimeH3K4 decreases in a manner consistent with expectations from studies in cultured cells [17], [21]. While this mark is commonly associated with activation in most contexts, recent genome-wide analysis studies have implicated this mark in both activation and repression [38], [39], and therefore our understanding of the functional consequences of this mark seems incomplete [35]–[37]. Two possibilities may account for the similarity of the dimethylH3K4 level in Δ-1.8 cells between repressed (III) and reactivated (IV) stages. First, dimethylH3K4 may not be the critical modification mediating maintenance of repression. Alternatively, other stage-specific factors in the nuclear milieu may lead to differential sensitivities to dimethylH3K4 between the repressed (III) and reactivated (IV) stages.
Substantial reduction in both dimeH3K4 and monomethylH3K4 were observed upon loss of the −1.8 kb site without a concomitant increase in trimethylH3K4. These findings suggest that the methylation states of H3K4 are regulated independently and locally through complexes recruited to the −1.8 kb GATA-binding site. These observations are in accordance with the finding that dimeH3K4 positive, trimethylH3K4 negative, marks are present at a subset of developmentally regulated hematopoietic genes [44]. Thus, our data highlight a potential role for these H3K4 marks in regulating transcription. It is interesting to note that the trimethylK27 mark, associated with GATA-1-mediated repression of the Gata2 locus [34], is not affected by the −1.8 kb GATA-binding site. In addition, reduction of H3K27 trimethylation, widely accepted as a repressive mark [45], does not appear to be required to reactivate gene expression at the Gata2 locus, perhaps indicating that it is involved selectively in the initiation of repression. Recent genome-wide analysis has also shown that H3K27 methylation is not merely present or absent, but rather increases quantitatively as the activity of the gene decreases [38], [40], suggesting that the level of transcriptional reactivation observed is within the range allowed by the H3K27 methylation level at this locus. Finally, in many cases, CpG rich promoters require hypermethylation of associated CpG islands for stable repression [4], [37]. We find that the CpG island at the Gata2 1S promoter lacks high levels of methylation during stable repression, and that loss of the −1.8 site does not affect methylation levels. This data further supports a model in which −1.8 kb site-dependent histone marks maintain stable repression.
We propose therefore that loss of GATA-1 binding and key repressive marks, including dimethyl - and monomethyl-H3K4, result in a locus permissive for Pol II occupancy and reactivation of transcription. This model predicts that a specific protein or proteins are recruited by GATA-1 to the −1.8 kb site to maintain repressive chromatin structure. GATA-1 is known to interact with CBP [46], HDACs 1 and 2 [47], [48], LSD1 [49], BRG1 [50], and polycomb repressive complex 2 (PRC2) [34]. As no GATA-1-interacting proteins have been reported that possess the requisite methyltransferase activity to establish the dimeH3K4 histone mark that is lost in the −1.8 kb mutant, novel GATA-1-containing complexes may be required to maintain the −1.8 kb site-dependent histone marks. Ongoing genetic ablation studies examining the contribution of the other known GATA-binding regions to Gata2 regulation and local chromatin architecture will be important for understanding the control of this complex locus.
Studies in multiple systems have led to a model of sequential gene repression during development [4], separable into distinct phases. Reversible repression is replaced by epigenetic mechanisms that alter the chromatin structure at the locus though modifications of histones, and in some cases DNA, to maintain stable repression. The results described herein support such a model and characterize molecular mechanisms associated with the selective maintenance of repression of an endogenous target gene by an individual cis element to confer normal developmental control.
Materials and Methods
Ethics statement
All animals were handled in strict accordance with good animal practice as defined by the relevant national and/or local animal welfare bodies, and the appropriate committee approved all animal work.
Generation of mice containing Gata2 Δ-1.8 knock-in allele
Briefly, to generate the −1.8 kb knock-in allele, we replaced the palindromic GATA sites (AGATAAGGCTTATCA) with two SalI sites in order to clone in a Neo resistance cassette flanked by loxP sites. Once the neo cassette is removed, the locus contains a single loxP site flanked by SalI sites. The new sequence does not contain any binding motifs known to be involved in hematopoietic development. In more detail, we first inserted a HpaI site into pBSK between NotI and SacI with an oligo. Then, we cloned a −7.2 kb to intron 1S fragment of the Gata2 locus into pBSK with KpnI and HpaI. We then replaced the two palindromic WGATAR sites with a Sal I site via PCR and replaced the wild-type XbaI to NdeI fragment with this mutated version. Then, we cloned HSV-TK cloned into the SacII site of pBSK. Following this, we cloned a second SalI site into the XbaI site of pflox21 with an oligo and used the flanking SalI sites to clone this loxP-PGKneo-loxP cassette into the SalI site of the Gata2-containing pBSK (Figure S1A). We screened targeted CJ7 ES clones by PCR and confirmed correct targeting by Southern blotting. We used standard blastocyst injection techniques to generate chimeric mice and screened F1 pups for germline transmission using Southern blotting (Figure S1B,C). In some mice, the loxP-neomycin resistance gene was deleted by crossing with Gata1-Cre mice, which were of CD1/Swiss-Webster background. We confirmed Cre-mediated excision of neo from these mice using PCR and all further genotyping was performed by PCR (Figure 1C). Mice were backcrossed onto a C57/Bl6 background for a minimum of 6 generations and were housed in a specific pathogen-free animal facility.
Fetal liver and bone marrow sampling
Fetal liver cells were obtained from embryos at E12.5 and E14.5 after timed matings. Mouse bone marrow cells were obtained from 8 - to 12-week-old animals by crushing femurs and tibias with either Iscove modified Dulbecco medium (IMDM) or Phosphate Buffered Saline (PBS) supplemented with 2% fetal calf serum (Mediatech, Herndon, VA). Single cell suspensions of fetal livers and spleens were made by passage through 70 micron nylon mesh (Sefar America, Kansas City, MO) in PBS supplemented 2% fetal calf serum (Mediatech, Herndon, VA). Cells were kept on ice until use and counts were performed using a Beckman Coulter AcT10 hematological analyzer.
Real-time reverse-transcriptase PCR
RNA was prepared from the described populations using the Trizol Kit (Invitrogen, San Diego, CA), DNAseI treated by RQ1 RNase-Free DNase (Promega, Madison, WI) and quantified. cDNA was synthesized using 1 µg of RNA with the iScript cDNA Synthesis Kit (Biorad, Hercules, CA). Typically, 1 µl of cDNA was then used as a template for quantitative PCR using the iQ SYBR Green Supermix (Biorad, Hercules, CA) in an iCycler thermocycler (Biorad, Hercules, CA). Primer sequences can be found in Text S1. Triplicate data sets were generated and results were normalized to β-actin reactions run in parallel.
Complete blood count
Whole PB was analyzed on a Beckman Coulter AcT10 hematological analyzer. White blood cell and progenitor subsets were analyzed from peripheral blood by staining with Gr-1 and Mac1 or CD3 and B220 after red blood cell lysis using ACK (NH4Cl) lysis buffer.
Flow-cytometric analysis and cell sorting
All antibodies for FACS were obtained from Pharmingen (San Diego, CA) or eBiosciences (San Diego, CA), and the following clones were used; Ly-76 (Ter-119), CD71 (C2), CD117 (2B8). Antibodies to surface markers of interest were used at 1∶60 dilution and after 30–60 minutes unbound antibody was washed away. In the case of biotinylated antibodies, streptavidin conjugated to various fluorochromes was added for the last 15–30 minutes of antibody incubation at 1∶100 dilution. For cell sorting experiments of erythroid progenitor subsets, fetal liver cells were stained with antibodies to CD71 and Ter119, and 7AAD was added to allow for exclusion of dead cells during sorting. For examination of enucleation, cells were stained with CD71 and Ter119 as above and incubated with Draq5 (Biostatus Limited, Leicestershire,United Kingdom) as per manufacturers instructions before analysis.
Quantitative chromatin immunoprecipitation (ChIP) assay
Rabbit anti-GATA-1 and anti-GATA-2 polyclonal antibodies have been described previously [16], [21], [51]. Rabbit anti-Pol II (N-20, sc-899) was from Santa Cruz Biotech. Rabbit anti-acetyl-histone H3 (#06-599), anti-trimethyl-histone H3 (Lys 9) (#07-442), anti-trimethyl-histone H3 (Lys 27) (#07-449) and anti-dimethyl-histone H3 (Lys 4) (#07-030) were from Millipore. Real-time-PCR-based quantitative chromatin immunoprecipitation (ChIP) analysis was conducted as described [52]. Single-cell suspensions were isolated from E14.5 wild-type and Δ-1.8 fetal liver cells, respectively, and crosslinked with 1% formaldehyde. Samples were analyzed by real-time PCR (ABI Prism 7000) using primers designed by PrimerExpress™ 1.0 software (PE Applied Biosystems) to amplify regions of 75–150 bp that overlap with the appropriate motif. Product was measured by SYBR Green fluorescence in 20 µl reactions, and the amount of product was determined relative to a standard curve generated from titration of input chromatin. Calculations were derived using percentage of input and were normalized using relative units which were determined by defining 9% input sample as 1.0. Analysis of dissociation-curves post-amplification showed that primer pairs generated single products.
Bisulfite sequencing
Bisulfite treatment of genomic DNA was performed as previously described using the Qiagen EpiTect Bisulfite Kit as per the manufacter's instructions. Sequence-specific PCR of the bisulfite-treated DNA was performed using primers specific to the murine Gata2 1S promoter (outside primers: F, 5′ - TTGTGTGGTGAGGGTGTAG-3′, R, 5′ - CAAATTTCTTTCCCTATTTTCT-3′; inside primers: F, 5′ - TAGGTGGGGGAGAGTGTAG -3′, R, 5′ - CAAATTTCTTTCCCTATTTTCT -3′. The PCR fragments were sub-cloned into the pCR®2.1-TOPO® vector (Invitrogen) and transformed into DH5α E. coli cells. Miniprep plasmid DNA was verified by EcoRI digestion and positive clones were sequenced using M13 forward (−20) or reverse primers.
Statistical analysis
Data are presented as mean ± SEM. Statistical significance was assessed by two-sided Student's t-test.
Supporting Information
Zdroje
1. ArnoneMI
DavidsonEH
1997 The hardwiring of development: organization and function of genomic regulatory systems. Development 124 1851 1864
2. SonejiS
HuangS
LooseM
DonaldsonIJ
PatientR
2007 Inference, validation, and dynamic modeling of transcription networks in multipotent hematopoietic cells. Ann N Y Acad Sci 1106 30 40
3. StrahlBD
AllisCD
2000 The language of covalent histone modifications. Nature 403 41 45
4. MohnF
SchubelerD
2009 Genetics and epigenetics: stability and plasticity during cellular differentiation. Trends Genet 25 129 136
5. BurchJB
2005 Regulation of GATA gene expression during vertebrate development. Semin Cell Dev Biol 16 71 81
6. BresnickEH
MartowiczML
PalS
JohnsonKD
2005 Developmental control via GATA factor interplay at chromatin domains. J Cell Physiol 205 1 9
7. KanekoH
ShimizuR
YamamotoM.
GATA factor switching during erythroid differentiation. Curr Opin Hematol 17 163 168
8. TsaiFY
OrkinSH
1997 Transcription factor GATA-2 is required for proliferation/survival of early hematopoietic cells and mast cell formation, but not for erythroid and myeloid terminal differentiation. Blood 89 3636 3643
9. TsaiFY
KellerG
KuoFC
WeissM
ChenJ
1994 An early haematopoietic defect in mice lacking the transcription factor GATA-2. Nature 371 221 226
10. FujiwaraY
BrowneCP
CunniffK
GoffSC
OrkinSH
1996 Arrested development of embryonic red cell precursors in mouse embryos lacking transcription factor GATA-1. Proc Natl Acad Sci U S A 93 12355 12358
11. TingCN
OlsonMC
BartonKP
LeidenJM
1996 Transcription factor GATA-3 is required for development of the T-cell lineage. Nature 384 474 478
12. BriegelK
LimKC
PlankC
BeugH
EngelJD
1993 Ectopic expression of a conditional GATA-2/estrogen receptor chimera arrests erythroid differentiation in a hormone-dependent manner. Genes Dev 7 1097 1109
13. PersonsDA
AllayJA
AllayER
AshmunRA
OrlicD
1999 Enforced expression of the GATA-2 transcription factor blocks normal hematopoiesis. Blood 93 488 499
14. HeyworthC
GaleK
DexterM
MayG
EnverT
1999 A GATA-2/estrogen receptor chimera functions as a ligand-dependent negative regulator of self-renewal. Genes Dev 13 1847 1860
15. FujiwaraT
O'GeenH
KelesS
BlahnikK
LinnemannAK
2009 Discovering hematopoietic mechanisms through genome-wide analysis of GATA factor chromatin occupancy. Mol Cell 36 667 681
16. WozniakRJ
BresnickEH
2008 Epigenetic control of complex loci during erythropoiesis. Curr Top Dev Biol 82 55 83
17. GrassJA
BoyerME
PalS
WuJ
WeissMJ
2003 GATA-1-dependent transcriptional repression of GATA-2 via disruption of positive autoregulation and domain-wide chromatin remodeling. Proc Natl Acad Sci U S A 100 8811 8816
18. PalS
CantorAB
JohnsonKD
MoranTB
BoyerME
2004 Coregulator-dependent facilitation of chromatin occupancy by GATA-1. Proc Natl Acad Sci U S A 101 980 985
19. MartowiczML
GrassJA
BoyerME
GuendH
BresnickEH
2005 Dynamic GATA factor interplay at a multicomponent regulatory region of the GATA-2 locus. J Biol Chem 280 1724 1732
20. MartowiczML
GrassJA
BresnickEH
2006 GATA-1-mediated transcriptional repression yields persistent transcription factor IIB-chromatin complexes. J Biol Chem 281 37345 37352
21. GrassJA
JingH
KimSI
MartowiczML
PalS
2006 Distinct functions of dispersed GATA factor complexes at an endogenous gene locus. Mol Cell Biol 26 7056 7067
22. WangH
ZhangY
ChengY
ZhouY
KingDC
2006 Experimental validation of predicted mammalian erythroid cis-regulatory modules. Genome Res 16 1480 1492
23. Kobayashi-OsakiM
OhnedaO
SuzukiN
MinegishiN
YokomizoT
2005 GATA motifs regulate early hematopoietic lineage-specific expression of the Gata2 gene. Mol Cell Biol 25 7005 7020
24. PimandaJE
OttersbachK
KnezevicK
KinstonS
ChanWY
2007 Gata2, Fli1, and Scl form a recursively wired gene-regulatory circuit during early hematopoietic development. Proc Natl Acad Sci U S A 104 17692 17697
25. WozniakRJ
BoyerME
GrassJA
LeeY
BresnickEH
2007 Context-dependent GATA factor function: combinatorial requirements for transcriptional control in hematopoietic and endothelial cells. J Biol Chem 282 14665 14674
26. ZhangJ
SocolovskyM
GrossAW
LodishHF
2003 Role of Ras signaling in erythroid differentiation of mouse fetal liver cells: functional analysis by a flow cytometry-based novel culture system. Blood 102 3938 3946
27. SocolovskyM
NamH
FlemingMD
HaaseVH
BrugnaraC
2001 Ineffective erythropoiesis in Stat5a(−/−)5b(−/−) mice due to decreased survival of early erythroblasts. Blood 98 3261 3273
28. PalisJ
2008 Ontogeny of erythropoiesis. Curr Opin Hematol 15 155 161
29. McGrathK
PalisJ
2008 Ontogeny of erythropoiesis in the mammalian embryo. Curr Top Dev Biol 82 1 22
30. LenoxLE
PerryJM
PaulsonRF
2005 BMP4 and Madh5 regulate the erythroid response to acute anemia. Blood 105 2741 2748
31. PerryJM
HarandiOF
PaulsonRF
2007 BMP4, SCF, and hypoxia cooperatively regulate the expansion of murine stress erythroid progenitors. Blood 109 4494 4502
32. PorayetteP
PaulsonRF
2008 BMP4/Smad5 dependent stress erythropoiesis is required for the expansion of erythroid progenitors during fetal development. Dev Biol 317 24 35
33. MinegishiN
OhtaJ
SuwabeN
NakauchiH
IshiharaH
1998 Alternative promoters regulate transcription of the mouse GATA-2 gene. J Biol Chem 273 3625 3634
34. YuM
RivaL
XieH
SchindlerY
MoranTB
2009 Insights into GATA-1-mediated gene activation versus repression via genome-wide chromatin occupancy analysis. Mol Cell 36 682 695
35. ShilatifardA
2008 Molecular implementation and physiological roles for histone H3 lysine 4 (H3K4) methylation. Curr Opin Cell Biol 20 341 348
36. PinskayaM
MorillonA
2009 Histone H3 lysine 4 di-methylation: a novel mark for transcriptional fidelity? Epigenetics 4 302 306
37. HublitzP
AlbertM
PetersAH
2009 Mechanisms of transcriptional repression by histone lysine methylation. Int J Dev Biol 53 335 354
38. BarskiA
CuddapahS
CuiK
RohTY
SchonesDE
2007 High-resolution profiling of histone methylations in the human genome. Cell 129 823 837
39. ZhaoXD
HanX
ChewJL
LiuJ
ChiuKP
2007 Whole-genome mapping of histone H3 Lys4 and 27 trimethylations reveals distinct genomic compartments in human embryonic stem cells. Cell Stem Cell 1 286 298
40. CuiK
ZangC
RohTY
SchonesDE
ChildsRW
2009 Chromatin signatures in multipotent human hematopoietic stem cells indicate the fate of bivalent genes during differentiation. Cell Stem Cell 4 80 93
41. IllingworthRS
BirdAP
2009 CpG islands–‘a rough guide’. FEBS Lett 583 1713 1720
42. SongF
SmithJF
KimuraMT
MorrowAD
MatsuyamaT
2005 Association of tissue-specific differentially methylated regions (TDMs) with differential gene expression. Proc Natl Acad Sci U S A 102 3336 3341
43. IrizarryRA
Ladd-AcostaC
WenB
WuZ
MontanoC
2009 The human colon cancer methylome shows similar hypo - and hypermethylation at conserved tissue-specific CpG island shores. Nat Genet 41 178 186
44. OrfordK
KharchenkoP
LaiW
DaoMC
WorhunskyDJ
2008 Differential H3K4 methylation identifies developmentally poised hematopoietic genes. Dev Cell 14 798 809
45. SchuettengruberB
ChourroutD
VervoortM
LeblancB
CavalliG
2007 Genome regulation by polycomb and trithorax proteins. Cell 128 735 745
46. BlobelGA
NakajimaT
EcknerR
MontminyM
OrkinSH
1998 CREB-binding protein cooperates with transcription factor GATA-1 and is required for erythroid differentiation. Proc Natl Acad Sci U S A 95 2061 2066
47. HongW
NakazawaM
ChenYY
KoriR
VakocCR
2005 FOG-1 recruits the NuRD repressor complex to mediate transcriptional repression by GATA-1. Embo J 24 2367 2378
48. RodriguezP
BonteE
KrijgsveldJ
KolodziejKE
GuyotB
2005 GATA-1 forms distinct activating and repressive complexes in erythroid cells. Embo J 24 2354 2366
49. SnowJW
OrkinSH
2009 Translational isoforms of FOG1 regulate GATA1-interacting complexes. J Biol Chem 284 29310 29319
50. KimSI
BultmanSJ
KieferCM
DeanA
BresnickEH
2009 BRG1 requirement for long-range interaction of a locus control region with a downstream promoter. Proc Natl Acad Sci U S A 106 2259 2264
51. ImH
GrassJA
JohnsonKD
KimSI
BoyerME
2005 Chromatin domain activation via GATA-1 utilization of a small subset of dispersed GATA motifs within a broad chromosomal region. Proc Natl Acad Sci U S A 102 17065 17070
52. ImH
GrassJA
JohnsonKD
BoyerME
WuJ
2004 Measurement of protein-DNA interactions in vivo by chromatin immunoprecipitation. Methods Mol Biol 284 129 146
Štítky
Genetika Reprodukční medicína
Článek Allelic Variation at the 8q23.3 Colorectal Cancer Risk Locus Functions as a Cis-Acting Regulator ofČlánek Allelic Selection of Amplicons in Glioblastoma Revealed by Combining Somatic and Germline AnalysisČlánek Lactic Acidosis Triggers Starvation Response with Paradoxical Induction of TXNIP through MondoAČlánek Rice a Cinnamoyl-CoA Reductase-Like Gene Family Member, Is Required for NH1-Mediated Immunity to pv.Článek Differentiation of Zebrafish Melanophores Depends on Transcription Factors AP2 Alpha and AP2 Epsilon
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2010 Číslo 9
-
Všechny články tohoto čísla
- Optimal Strategy for Competence Differentiation in Bacteria
- Mutational Patterns Cannot Explain Genome Composition: Are There Any Neutral Sites in the Genomes of Bacteria?
- Frail Hypotheses in Evolutionary Biology
- Genetic Architecture of Complex Traits and Accuracy of Genomic Prediction: Coat Colour, Milk-Fat Percentage, and Type in Holstein Cattle as Contrasting Model Traits
- Allelic Variation at the 8q23.3 Colorectal Cancer Risk Locus Functions as a Cis-Acting Regulator of
- Allelic Selection of Amplicons in Glioblastoma Revealed by Combining Somatic and Germline Analysis
- Germline Variation Controls the Architecture of Somatic Alterations in Tumors
- Mice Doubly-Deficient in Lysosomal Hexosaminidase A and Neuraminidase 4 Show Epileptic Crises and Rapid Neuronal Loss
- Analysis of Population Structure: A Unifying Framework and Novel Methods Based on Sparse Factor Analysis
- FliO Regulation of FliP in the Formation of the Flagellum
- Cdc20 Is Critical for Meiosis I and Fertility of Female Mice
- dMyc Functions Downstream of Yorkie to Promote the Supercompetitive Behavior of Hippo Pathway Mutant Cells
- DCAF26, an Adaptor Protein of Cul4-Based E3, Is Essential for DNA Methylation in
- Genome-Wide Double-Stranded RNA Sequencing Reveals the Functional Significance of Base-Paired RNAs in
- An Immune Response Network Associated with Blood Lipid Levels
- Genetic Variants and Their Interactions in the Prediction of Increased Pre-Clinical Carotid Atherosclerosis: The Cardiovascular Risk in Young Finns Study
- The Histone H3K36 Methyltransferase MES-4 Acts Epigenetically to Transmit the Memory of Germline Gene Expression to Progeny
- Long- and Short-Term Selective Forces on Malaria Parasite Genomes
- Lactic Acidosis Triggers Starvation Response with Paradoxical Induction of TXNIP through MondoA
- Identification of Early Requirements for Preplacodal Ectoderm and Sensory Organ Development
- Orphan CpG Islands Identify Numerous Conserved Promoters in the Mammalian Genome
- Analysis of the Basidiomycete Reveals Conservation of the Core Meiotic Expression Program over Half a Billion Years of Evolution
- ETS-4 Is a Transcriptional Regulator of Life Span in
- The SR Protein B52/SRp55 Is Required for DNA Topoisomerase I Recruitment to Chromatin, mRNA Release and Transcription Shutdown
- The Baker's Yeast Diploid Genome Is Remarkably Stable in Vegetative Growth and Meiosis
- Chromatin Landscape Dictates HSF Binding to Target DNA Elements
- The APETALA-2-Like Transcription Factor OsAP2-39 Controls Key Interactions between Abscisic Acid and Gibberellin in Rice
- Accurately Assessing the Risk of Schizophrenia Conferred by Rare Copy-Number Variation Affecting Genes with Brain Function
- Widespread Over-Expression of the X Chromosome in Sterile F Hybrid Mice
- The Characterization of Twenty Sequenced Human Genomes
- The Genome of a Pathogenic : Cooptive Virulence Underpinned by Key Gene Acquisitions
- A Single Element Maintains Repression of the Key Developmental Regulator
- Identification of New Genetic Risk Variants for Type 2 Diabetes
- Effect of Correlated tRNA Abundances on Translation Errors and Evolution of Codon Usage Bias
- Evidence of Selection upon Genomic GC-Content in Bacteria
- Proteomic Changes Resulting from Gene Copy Number Variations in Cancer Cells
- Rice a Cinnamoyl-CoA Reductase-Like Gene Family Member, Is Required for NH1-Mediated Immunity to pv.
- Longitudinal Genome-Wide Association of Cardiovascular Disease Risk Factors in the Bogalusa Heart Study
- Response to Mechanical Stress Is Mediated by the TRPA Channel Painless in the Heart
- DNMT3L Modulates Significant and Distinct Flanking Sequence Preference for DNA Methylation by DNMT3A and DNMT3B
- Identifying Signatures of Natural Selection in Tibetan and Andean Populations Using Dense Genome Scan Data
- Incremental Genetic Perturbations to MCM2-7 Expression and Subcellular Distribution Reveal Exquisite Sensitivity of Mice to DNA Replication Stress
- Loss of Maternal ATRX Results in Centromere Instability and Aneuploidy in the Mammalian Oocyte and Pre-Implantation Embryo
- Comparative Genomic Hybridization (CGH) Reveals a Neo-X Chromosome and Biased Gene Movement in Stalk-Eyed Flies (Genus )
- Differentiation of Zebrafish Melanophores Depends on Transcription Factors AP2 Alpha and AP2 Epsilon
- Gene–Environment Interactions at Nucleotide Resolution
- Dementia Revealed: Novel Chromosome 6 Locus for Late-Onset Alzheimer Disease Provides Genetic Evidence for Folate-Pathway Abnormalities
- Critical Functions of Rpa3/Ssb3 in S-Phase DNA Damage Responses in Fission Yeast
- Preferential Re-Replication of Heterochromatin in the Absence of Geminin
- The Potential for Enhancing the Power of Genetic Association Studies in African Americans through the Reuse of Existing Genotype Data
- Evidence That Mutation Is Universally Biased towards AT in Bacteria
- Perturbation Analysis of Heterochromatin-Mediated Gene Silencing and Somatic Inheritance
- Diversity of Eukaryotic DNA Replication Origins Revealed by Genome-Wide Analysis of Chromatin Structure
- Genetic Deletion of the Desmosomal Component Promotes Tumor Microinvasion in a Mouse Model of Pancreatic Neuroendocrine Carcinogenesis
- The Metabolic Enzyme ManA Reveals a Link between Cell Wall Integrity and Chromosome Morphology
- SNPs Associated with Cerebrospinal Fluid Phospho-Tau Levels Influence Rate of Decline in Alzheimer's Disease
- Synthesizing and Salvaging NAD: Lessons Learned from
- A Central Regulatory System Largely Controls Transcriptional Activation and Repression Responses to Phosphate Starvation in Arabidopsis
- An Insect Herbivore Microbiome with High Plant Biomass-Degrading Capacity
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Synthesizing and Salvaging NAD: Lessons Learned from
- Optimal Strategy for Competence Differentiation in Bacteria
- Long- and Short-Term Selective Forces on Malaria Parasite Genomes
- Identifying Signatures of Natural Selection in Tibetan and Andean Populations Using Dense Genome Scan Data
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



