-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaThe Homeodomain Iroquois Proteins Control Cell Cycle Progression and Regulate the Size of Developmental Fields
The correct development of body organs, with their characteristic size and shape, requires the coordination of cell division and cell differentiation. Here we show that the Iroquois proteins (Irx in vertebrates) slow down cell division in the Drosophila imaginal discs, in addition to their well-known role in cell fate and territorial specification. In humans, inactivating mutations at the Irx genes are associated to several types of cancer, thus allowing their classification as tumour suppressor genes. We have observed that Drosophila Iroquois genes similarly behave as tumour suppressor genes. Iroquois proteins belong to a family of homeodomain-containing transcriptional regulators. However, our results indicate that they control cell division by a transcription independent mechanism based on their physical interaction with Cyclin E containing complexes, a key player in cell-cycle progression. We have identified two evolutionary conserved domains of Iroquois proteins, different from the homeodomain, involved in that interaction. This new function of Iroquois proteins places them in a key position to coordinate growth and differentiation during normal development. Our results further suggest a molecular mechanism for their role in tumour suppression. Future studies of Irx genes should help to determine if a similar mechanism could operate to help cancer progression when Irx activity is compromised.
Published in the journal: . PLoS Genet 11(8): e32767. doi:10.1371/journal.pgen.1005463
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1005463Summary
The correct development of body organs, with their characteristic size and shape, requires the coordination of cell division and cell differentiation. Here we show that the Iroquois proteins (Irx in vertebrates) slow down cell division in the Drosophila imaginal discs, in addition to their well-known role in cell fate and territorial specification. In humans, inactivating mutations at the Irx genes are associated to several types of cancer, thus allowing their classification as tumour suppressor genes. We have observed that Drosophila Iroquois genes similarly behave as tumour suppressor genes. Iroquois proteins belong to a family of homeodomain-containing transcriptional regulators. However, our results indicate that they control cell division by a transcription independent mechanism based on their physical interaction with Cyclin E containing complexes, a key player in cell-cycle progression. We have identified two evolutionary conserved domains of Iroquois proteins, different from the homeodomain, involved in that interaction. This new function of Iroquois proteins places them in a key position to coordinate growth and differentiation during normal development. Our results further suggest a molecular mechanism for their role in tumour suppression. Future studies of Irx genes should help to determine if a similar mechanism could operate to help cancer progression when Irx activity is compromised.
Introduction
Development of the different body parts in multicellular organisms is a stepwise process that entails the specification within developmental fields of territories with the ability to acquire different fates. Morphogens, which orchestrate such territorial specification, are also able to regulate territorial growth [1]. There is increasing evidence that, conversely, the regulation of the size of the developmental fields over which morphogens spread and operate is paramount for territorial specification [2–4]. For instance, in two paradigms of morphogenetic fields—the vertebrate limb primordium and the Drosophila imaginal discs - two sources of morphogens are present at opposite sites. Since activity of one of them is prevented by the action of the other one, the morphogenetic field must reach a critical size for that morphogen to escape from inhibition and be able to initiate the territorial specification program [5–8]. Therefore, the identification of the genes that control cell proliferation in developmental fields is key to a better understanding of how cell proliferation and territorial specification are coordinated during development.
Here we address the role of the Drosophila Iroquois Complex genes (Iro genes) in cell proliferation. The three Iro genes, araucan (ara), caupolican (caup) and mirror (mirr), encode highly related and evolutionarily conserved homeodomain transcription factors of the TALE family [9–11]. They play key roles in development that range from territorial specification to pattern formation (reviewed in [12]). Namely, at the early second larval instar, Iro genes are expressed in sub-regions of the wing and eye imaginal discs where they define the prospective notum and the dorsal compartment of the eye, respectively [13–15]. Iro genes also contribute to the growth of the discs by generating organising borders at the confrontation of Iro-expressing and non-expressing cells [13–15]. In the dorsal compartment of the eye disc, Iro proteins repress the expression of fringe (fng), thus restricting the activation of the Notch pathway at the dorso/ventral (D/V) compartment border. This triggers growth of the entire eye disc and the initiation of retinal differentiation from its posterior rim [14, 16, 17], reviewed in [18]. Moreover, Iro proteins may also have a more direct role in the control of cell proliferation. Thus, clones of iro- cells in the eye disc are larger than the control ones [13, 19] and, conversely, generalized over-expression of ara in the wing disc reduces wing size [9]. Furthermore, vertebrate Irx genes (orthologs of Drosophila Iroquois genes) appear to function as tumour suppressor genes (TSG) for certain types of cancer [20–23].
In this work we show that Iro proteins indeed control cell proliferation, both during normal development and in several established Drosophila tumour-like models. Iro proteins specifically regulate the G1-S transition of the cell cycle by modulating the activity of the CyclinE/ Cyclin dependent kinase 2 (CycE/Cdk2) complex. Unexpectedly for transcription factors, they are able to do so by a non-transcriptional mechanism. Thus, we demonstrate that Caup forms a protein complex with CycE in S2 cells and disclose the function of the evolutionarily-conserved IRO·box domain of Caup for that physical interaction and for cell cycle regulation in vivo. Our results support a direct, cell-autonomous role of Drosophila Iro genes in the regulation of cell cycle progression. This function of the Iro genes uncovers a new layer of regulation of organ size during development and may account for their behaviour as tumour suppressor genes.
Results
Loss of function of Iro genes enhances cell proliferation
We found that iroEGP1 homozygous flies and those harbouring the iroEGP1 allele combined with a deficiency of the whole Iro-C (iroDFM3, S1A Fig) had dorsally enlarged eyes (Fig 1A–1D, 5% of iroEGP1 flies, 36% of the iroEGP1 /iroDFM3 everted flies). The cephalic capsule was morphologically normal, except for alterations in the number of orbital bristles (Fig 1D, arrowhead). In third instar wild-type eye imaginal discs, the three Iro genes are expressed in a dorsal domain ahead of the morphogenetic furrow (S1B and S1C Fig, see also [10, 14]). In contrast, in iroEGP1 /iroDFM3 eye discs the expression of caup was undetectable and that of ara was strongly decreased, while mirr expression was not affected (S1D–S1F Fig). Dorsally enlarged eyes were also found in 51% of the flies depleted of Mirr (by expression of two copies of UAS-mirr RNAi driven by eyGal4 at 25°C).
Fig. 1. Cell-autonomous increase in cell proliferation in iro mutants. 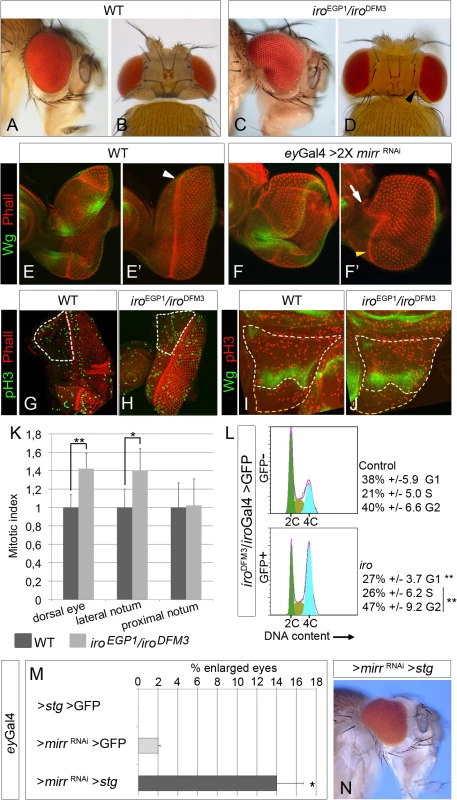
Lateral (A, C) and dorsal (B, D) views of heads of flies of the indicated genotypes. (E- F’) Expression of Wg (green) and Phalloidin staining (red) in wild-type (E, E’) and eyGal4>mirr RNAi (two copies of mirr RNAi, flies raised at 29°C, F, F’) eye discs. (E and E’ and F and F’ are different focal planes of the same disc). Arrowheads and arrow mark the position of the morphogenetic furrow. (G-K) Mitotic patterns (phospho-Histone H3 staining, green, G, H; red I, J) and quantification of the relative mitotic index (K) in Iro-expressing territories (white dotted areas in G-J) and in the prospective proximal notum (yellow dotted areas in I, J). (*p<0.05; **p<0.005). (L) G1/S transition is accelerated in iro mutant cells. Representative profiles of FACS analysis of cells dissociated from iroDFM3/iroGal4 UAS-GFP wing discs. (The differences in the percentages of G1 and (G2+S) cells between the GFP+ and GFP- populations are statistically significant, **p<0.005). (M, N) Reduction of Mirr levels (one copy of UAS-mirr RNAi, larvae raised at 25°C) and over-expression of stg synergistically interact to increase eye size. (M) Quantification of the fraction of enlarged eyes in flies of the indicated genotypes (average from two independent experiments, n>100, *p<0.05). (N) Representative mutant enlarged eye. In this and following figures, the eye discs are oriented dorsal up and posterior to the right, and the wing discs, ventral up and posterior to the right. Quantitative data are shown as arithmetic mean +/- SD (error bars). WT, wild-type. Ectopic D/V organisers, induced by clones of iro mutant cells in the dorsal compartment of the eye disc, can promote dorsally enlarged eyes [14, 19]. In adult eyes, the D/V organizer is visualized as the symmetry axis of the ommatidia field, named the equator (S1G and S1G’ Fig; [24]). However, ectopic equators were not found in retina sections of adult enlarged iroEGP1/iroDFM3 eyes (S1H and S1H’ Fig). Enlarged eyes have also been associated with reduced activity of the Wingless (Wg) pathway, which allows morphogenetic furrow initiation from the lateral margins of the disc [25]. While similar advance of the morphogenetic furrow was found in the dorsal domain in Mirr depleted eye discs (ey-Gal4> 2 X UAS-mirr RNAi, Fig 1F’, arrow), expression of wg was not apparently modified (Fig 1E and 1F, see also [14, 19]). Thus, we can rule out the generation of ectopic D/V organisers or insufficiency for Wg as the cause(s) of the observed eye enlargements.
Next, we monitored the rate of cell proliferation and the occurrence of cell death in iroEGP1/iroDFM3 eye discs, as their modifications might explain the enlarged eyes. Indeed the mitotic index was significantly increased in the Iro expressing domain, as compared to similar regions of wild-type discs (Fig 1G, 1H and 1K). This increase was not specific of the eye disc, since it also occurred in the lateral-notum region of iroEGP1/iroDFM3 wing discs (Fig 1I–1K, the lateral notum is delimited proximally by wg expression and distally by the most proximal of the wing hinge folds). Notably, the mitotic index was not altered in the region of iroEGP1/iroDFM3 wing discs proximal to the domain of wg expression (a region where Iro genes are not expressed at the third instar [12]), when compared to that of a similar region of wild-type discs (Fig 1I–1K).
We analyzed the cell cycle profiles of iro mutant cells using iroDFM3/iroGal4 UAS-GFP wing imaginal discs, which express GFP in the ara/caup domain [26]. iroGal4 is a hypomorphic iro allele [26] and iroDFM3 is a null allele (S1A Fig). We separated the GFP+ and GFP- cell populations by FACS. In wild type wing discs, the cell cycle profile of wing pouch disc cells (mostly Iro non-expressing cells) and that of the rest of the disc (most of them Iro-expressing cells) are very similar [27]. Thus, GFP- cells represented the internal control. Indeed, their cell cycle profile (38% in G1, 21% in S and 40% in G2, Fig 1L) was very similar to that previously described for wild-type cells from whole wing discs [28] and for wing pouch disc cells [27]. However, iro mutant GFP+ cells showed a cell cycle profile statistically different from that of rest of the wing disc cells (27% in G1, 26% in S and 47% in G2, Fig 1L). These alterations in the cell cycle profile resembled those caused by over-expression of cycE [28] and suggested that reduced levels of Iro proteins accelerate the passage through the G1 phase. In sum, these results allow us to conclude that Iro proteins cell-autonomously restrict cell proliferation.
We reasoned that an increase in the rate of the G2-M transition in the iro mutant eye discs should enhance eye overgrowth. Indeed, we found a synergistic effect on dorsal eye growth when string (stg), a phosphatase that drives the G2-M transition [29], was expressed in a background of slightly reduced expression of mirr (Fig 1M and 1N). We conclude that the reduced levels of Iro proteins in the dorsal territory of the iro eye disc induced over proliferation that resulted in dorsal eye overgrowth.
We also found an increased number of apoptotic cells in the iro territories of the mutant discs (S1M–S1P’ Fig). This increased apoptosis might help compensate the excess of proliferation, and reduce the extent of overgrowth especially in the notum (that was only slightly deformed in iroEGP1 mutants, S1I–S1L Fig). It further precludes precise analysis of the doubling time of iro mutant cells.
Over-expression of Iro genes restricts cell proliferation
Next, we tested whether over-expression of Iro genes caused the opposite effect to their loss of activity, that is, a reduction of cell proliferation. Since generalized expression of any of the Iro genes in the eye disc eliminates the D/V organiser and prevents growth of the eye disc and eye formation [14, 16, 17], we examined the effect of caup excess of function in the wing disc. We over-expressed caup-HA (henceforth caup) either in its normal expression domain, the prospective notum (using the apGal4 driver) or in the wing pouch (nubGal4 driver). We assayed the effect of transient over-expressions of caup using of the Gal4/Gal80ts system. We combined the nubGal4 and apGal4lines with a tubGal80ts transgene [30]. At 17°C, (permissive temperature for Gal80ts), Gal80 inhibits Gal4 activity. nubGal4 (or apGal4); tubGal80ts; UAS-caup larvae were raised at 17°C, and transferred to 29°C (to inactivate Gal80ts) 16 hours prior to their dissection at late third larval instar. Both in the nub and ap domains, caup over-expression caused a significant reduction in the mitotic index (Fig 2A–2B’ and 2D; S2J and S2K Fig), Similar reduction in the mitotic index also occurred upon forced expression of ara or mirr (S2D, S2E and S2H Fig). We also observed a decreased incorporation of the thymidine analogue EdU in the cells over-expressing caup (Fig 2E and 2E’). Cell size was not noticeably affected by transient caup over expression (S2A and S2B Fig). Since it also was unmodified by depletion of CycE in similar experimental conditions (S2C Fig), we assume this could be due to the transient over-expression of the transgenes.
Fig. 2. Over-expression of caup inhibits cell cycle progression. 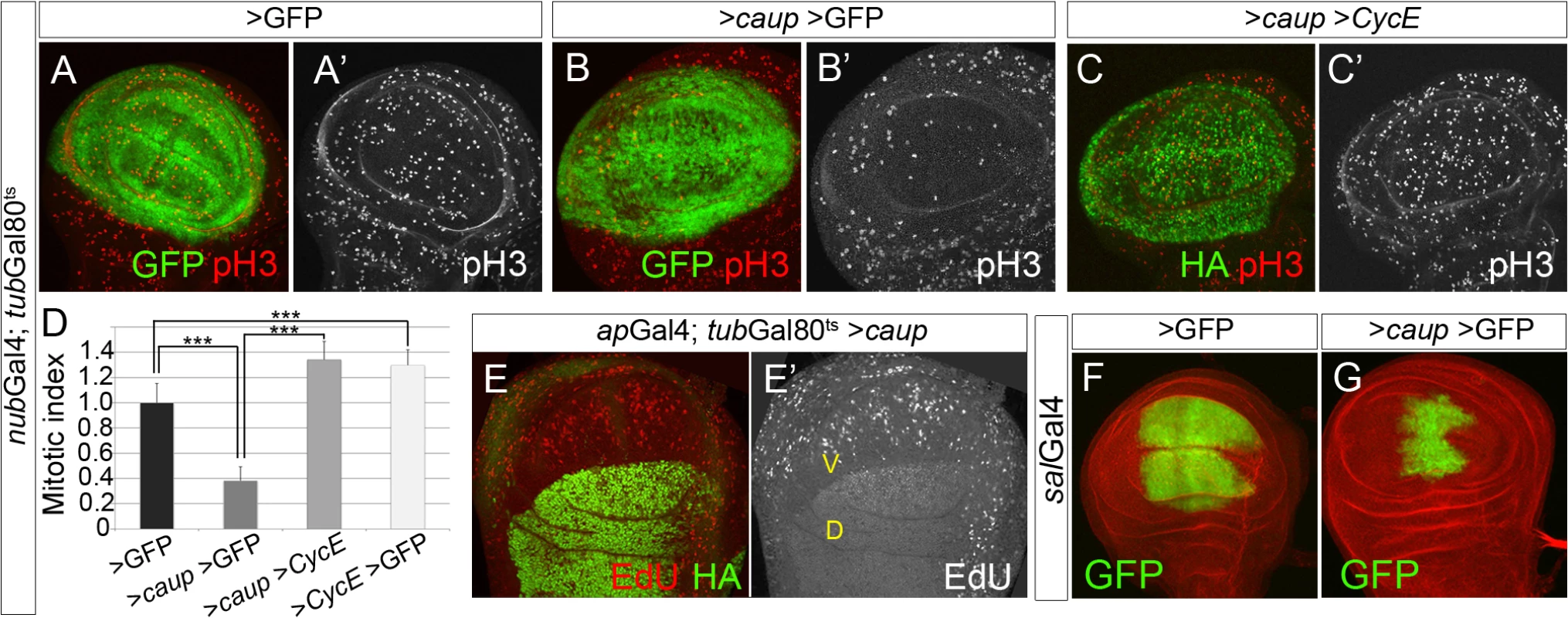
(A-C’) Mitotic pattern (pH3 staining) of wing imaginal discs expressing the indicated transgenes driven by nubGal4 during 16 h prior to dissection (expression domain shown in green). (D) Quantification of the relative mitotic index in the nub territory in the indicated genetic backgrounds (***p<0.00001). (E, E’) Pattern of S phase cells (assayed by EdU incorporation) in wing discs expressing caup-HA (green in E) in the dorsal (D) compartment (apGal4 driver). Compare the pattern of EdU incorporation in the dorsal and control ventral (V) compartment. (F, G) Over-expression of caup driven by salGal4 reduces the size of the sal territory (labelled by GFP, disc counterstained with phalloidin, red). To analyze the effect of much prolonged over-expression of caup in the wing disc, compatible with the development of the adult wing, we resorted to the salGal4 line. This Gal4 line drives expression of UAS genes in the central wing pouch of the wing disc from early third instar until 4h of pupal development (Fig 2F, [31]). Accordingly to a decreased cell proliferation in the sal domain of the wing discs caused by caup over-expression (S2M–S2M” Fig), we found a significant reduction in the size of this domain (Fig 2G, compare with F;) and of the adult wings (Fig 3A, 3B and 3G and S3A Fig). Furthermore, the mutant wings showed altered venation pattern and wing margin notches (Fig 3A and 3B). Wing notches were also found associated to cell cycle arrest caused by depletion of CycE (S3D Fig) and by the over-expression of dacapo (dap), ortholog of the Cyclin-dependent kinase (Cdk) inhibitor p21 [32], (S3E Fig) and could be attributed to reduced wg expression at the prospective wing margin in the salGal4>caup wing discs (S3B–S3C’ Fig). In addition, salGal4>caup wings showed enlarged cells in the sal domain (Fig 3H; S3J and S3K Fig). Small wings with vein patterning defects also result from over-expressing ara [9].
Fig. 3. Genetic interactions of caup with cell cycle regulators. 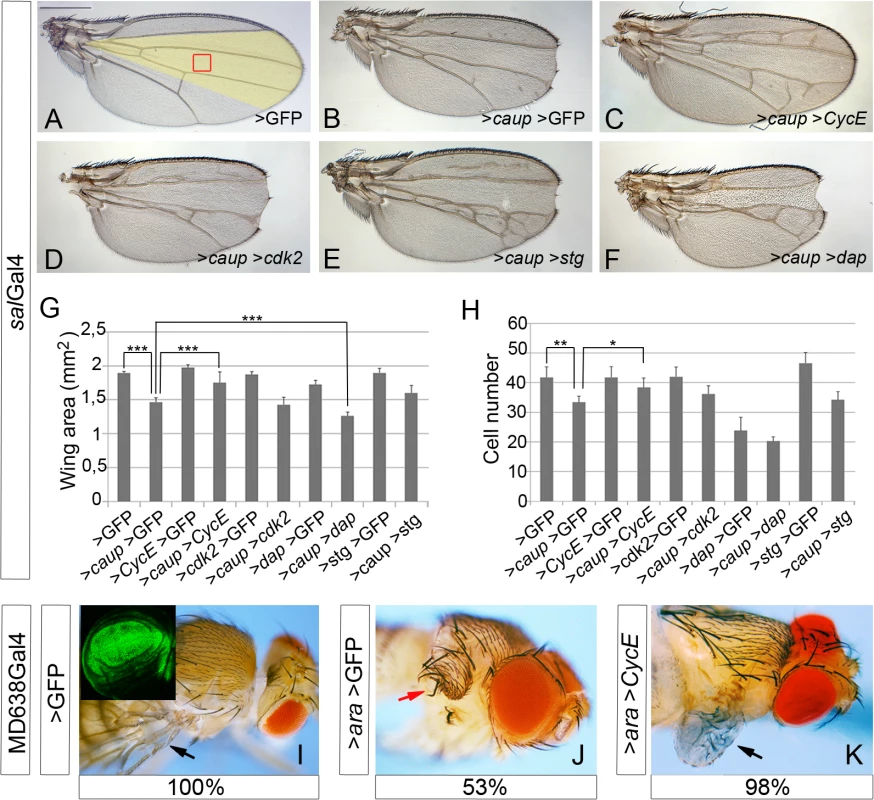
(A-F) Representative wings from flies of the indicated genotypes. Scale bar in A represents 500 μm. The region of the adult wing derived from the sal-expressing domain is shown in yellow in A. (G, H) Quantification of wing area (G, n = 10) and of the number of cells in a fixed wing area, similar to the region boxed in A (H, n = 5, calculated from the number of trichomes) for the indicated genotypes. ***p<0.0001; **p<0.005; *p<0.05. (I-K) Restoring cell proliferation by exogenously provided CycE recovers wing development in flies over-expressing ara (arrows). Transgene expression was driven by MD638Gal4 (expression domain in green in the inset in I). Red arrow in J indicates the notum-like structure that develops after ara over-expression (53% of the cases). The remaining MD638Gal4>ara>GFP flies present a wing stump and do not develop extra notum tissue. 98% of flies co-expressing ara and cycE show partially recovered wings (black arrow in K) and never develop a double notum (n>90). Although some cells entered apoptosis after caup over-expression (S4E and S4E’ Fig), their contribution to the mutant phenotype was apparently minimal. Co-expression of caup with the apoptosis inhibitor DIAP1 [33], reduced apoptosis (S4E–S4F’ Fig) but did not modify either the size, vein pattern or notches of wings over-expressing caup (S4A, S4B and S4I Fig). salGal4 driven expression lasts until 4 h after puparium formation [31]. Thus, to rule out the possibility that cell death during pupal stages could contribute to the mutant phenotype of salGal4>caup flies, we over-expressed caup in heterozygous conditions for Df(3L)H99. This deficiency removes the apoptosis inducing genes reaper, hid and grim [34] and halving the copy number of these genes largely reduces induced cell death [35]. We found that such reduction of apoptotic-inducing proteins did not modify the wing phenotype of salGal4>caup flies (S4C, S4D and S4I Fig).
These results indicate that elevated levels of Iro proteins restrict cell proliferation in the wing imaginal discs.
caup genetically interacts with CycE
To further analyze the role of Iro proteins on cell cycle progression we searched for genetic interactions between caup and several cell cycle regulators. Co-expression of caup (salGal4 driver) with the G2-M regulator stg [32], which on its own only slightly decreased cell size (Fig 3H; S3F Fig), did not rescue the effects of caup over-expression (Fig 3B, 3E, 3G and 3H).
Next we investigated the interaction of caup with G1/S regulators. CycE binds to and activates Cdk2 to drive the G1-S transition [32]. While over-expression of CycE or Cdk2 (salGal4 driver) did not modify wing or cell size (Fig 3G and 3H; S3G and S3H Fig), the co-expression of caup with CycE reverted all aspects of the caup over-expression adult phenotype (Fig 3A–3C, 3G and 3H; S3J–S3L Fig). Nevertheless, no reversion of the phenotype was observed by co-expressing cdk2 (Fig 3D, 3G and 3H). In contrast, the Cdk inhibitor dap [32], whose over-expression reduced wing size and cell number and caused wing notches (Fig 3G and 3H; S3E and S3I Fig), enhanced the caup over-expression effect (Fig 3B and 3F–3H).
These results suggested that CycE, but not Cdk2, becomes a limiting factor for cell proliferation in the presence of high levels of Caup. Therefore, we examined if exogenously provided CycE could recover cell proliferation in cells over-expressing caup (nubGal4 and apGal4 drivers). Fig 2A–2D and S2J–S2L Fig showed that this was indeed the case. Similar interactions were observed between ara or mirr and CycE (S2D–S2I Fig). Conversely, we found that co-expression with the F-box protein Archipelago (Ago), which induces CycE degradation through the proteosome pathway [36], significantly reduced the size of the salGal4>caup wings (S5A–S5D and S5G Fig). However, depletion of Ago (by expression of ago RNAi), which increased wing size (S5E and S5G Fig) did not recover but even enhanced the caup-over expression phenotype (S5B, S5F and S5G Fig). This effect could be attributed to the stabilization of unknown targets of Ago (other than CycE) by the depletion of this protein.
Next, we wonder whether similar insufficiency for CycE and the resulting impaired cell proliferation, could underlie other adult phenotypes caused by Iro genes over-expression. Associated to ectopic expression of ara in the prospective wing pouch, wings are absent and extra notum tissue develops ([37–39] and Fig 3J). Interestingly, co-expression of ara and CycE, which restored cell proliferation, allowed differentiation of a wing, albeit of a reduced size (Fig 3K). These results agree with those of [8], which showed that decreased cell proliferation in the wing pouch from early larval stages causes wing loss and duplication of body wall structures. In sum, these genetic interactions further support the regulation of cell cycle progression by Iro proteins at the G1-S transition suggested by the cell cycle profile analyses.
caup over-expression inhibited the activity of the CycE/Cdk2 complex
In Drosophila, the activity of the CycE/Cdk2 complex is required and sufficient for G1-S transition [32]. We examined the activity of this complex in cells that ectopically express caup by MPM-2 staining. This antibody recognizes a CycE/Cdk2 regulated protein complex that assembles into the histone locus body and is visualized as nuclear dots [40]. As shown in Fig 4A and 4A’, caup over-expressing cells of the posterior compartment (hhGal4 driver) displayed lower punctuated staining than control anterior cells indicating a decreased activity of the CycE/Cdk2 complex.
Fig. 4. Functional and physical interaction of Caup with the CycE/Cdk2 complex. 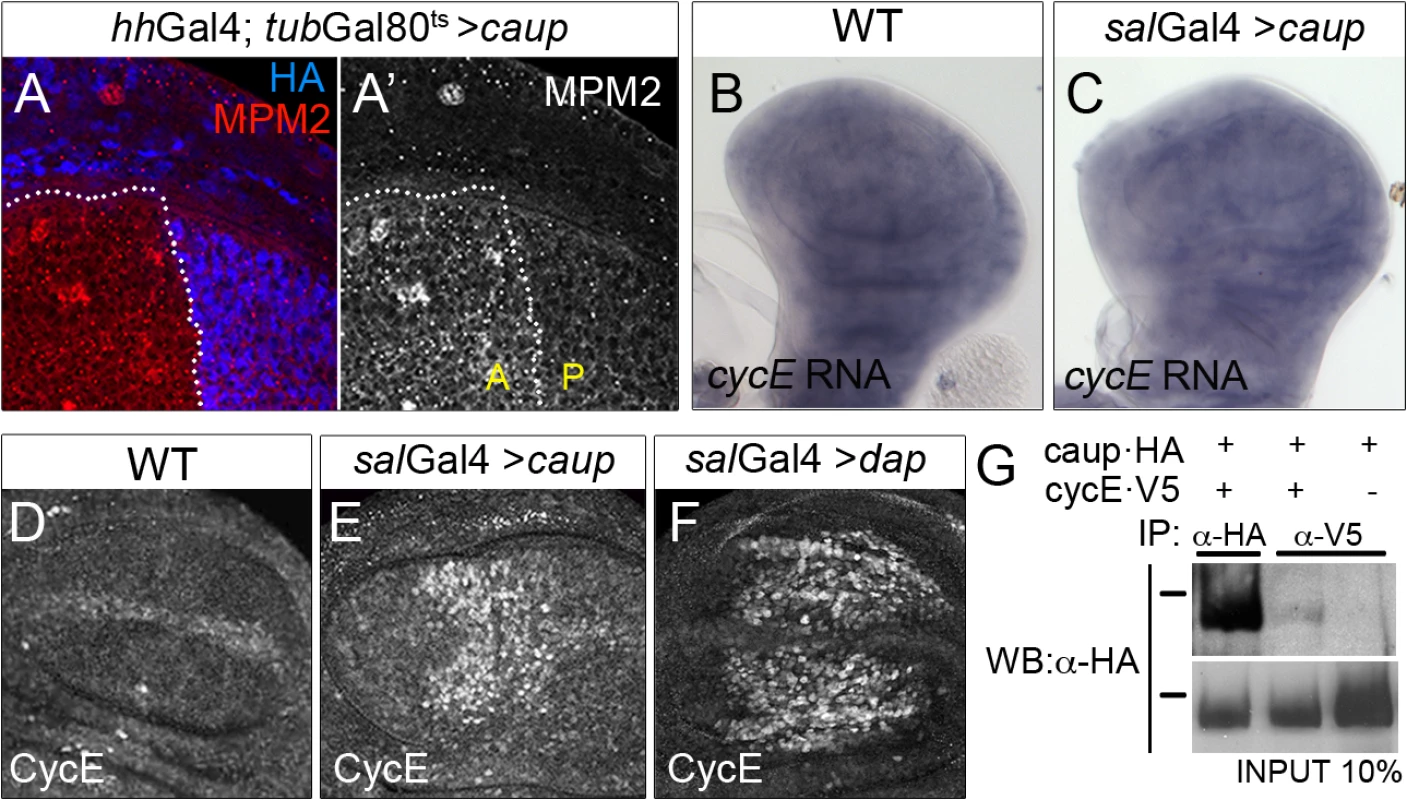
Activity of the CycE/Cdk2 complex, monitored by MPM-2 staining (A, A’); cycE transcription (detected by in situ hybridization, B, C) and CycE accumulation (detected by immunostaining, D-F) in wing imaginal discs of the indicated genotypes. (G) Caup co-immunoprecipitates with CycE in S2 cells. Western blot of protein extracts from S2 cells expressing the indicated tagged proteins, immunoprecipitated with anti-HA or anti-V5 antibodies and probed with anti-HA. Black bars indicate position of the 100 KDa protein marker. As we have shown above, CycE is a limiting component in caup over-expressing cells. Thus, the decreased activity of the CycE/Cdk2 complex could result from repression of CycE expression. However, transcription of CycE in the wing disc was not noticeably modified by caup forced expression (Fig 4B and 4C, see also S6A–S6C Fig). Interestingly, CycE protein levels were strongly increased (Fig 4D and 4E), even when apoptosis was reduced in salGal4>caup discs (S4G–S4H’ and S4J Fig). This suggested the stabilization of CycE protein when caup was over-expressed. Since phosphorylation of CycE by the CycE/Cdk2 complex is essential for its degradation [41], this result also supported that Caup reduced the activity of the CycE/Cdk2 complex. Similar increase in CycE levels was found associated to the inhibition of the CycE/Cdk2 complex by dap over expression (Fig 4F). Since mRNA and protein levels of dap were not modified in caup over-expressing cells (S6D–S6I Fig), the decreased activity of the CycE/Cdk2 complex in caup over expressing cells cannot be attributed to a deficiency of CycE or to excess amount of Dap.
Caup bind to a CycE-containing protein complex
Putative Cyclin-binding sites have been identified in the three Drosophila Iro proteins (Eukaryotic Linear Motiv server, http://elm.eu.org). Hence, we wondered whether the reduction of CycE function in caup over-expressing cells (despite their higher than normal CycE levels) might be due physical interaction of Caup with CycE containing complexes. We tested for this interaction by co-immunoprecipitation of Caup-HA and CycE-V5 from Drosophila S2 cells. As shown in Fig 4G, Caup-HA was present in CycE-containing complexes.
Next, we tested whether the putative Cyc-binding site present in Caup mediated the interaction with CycE and, therefore, its effect on cell cycle regulation. We mutated this site and over-expressed the resulting protein (Caupcyc*, Fig 5A) in wing discs. Caupcyc* was less effective than wild-type Caup in reducing wing size (Fig 5B and 5D) and in repressing cell proliferation (Fig 5H), although it appeared similarly effective than wild-type Caup in inducing CycE accumulation (a read-out of the inhibition of CycE/Cdk2 complex activity, Fig 5E, 5E’ and 5G). In agreement with our working hypothesis, the decreased ability of Caupcyc* to reduce wing size and cell proliferation was paralleled by its compromised ability to co-immunoprecipitate with CycE in S2 cells (Fig 5I).
Fig. 5. Structure-function analysis of Caup. 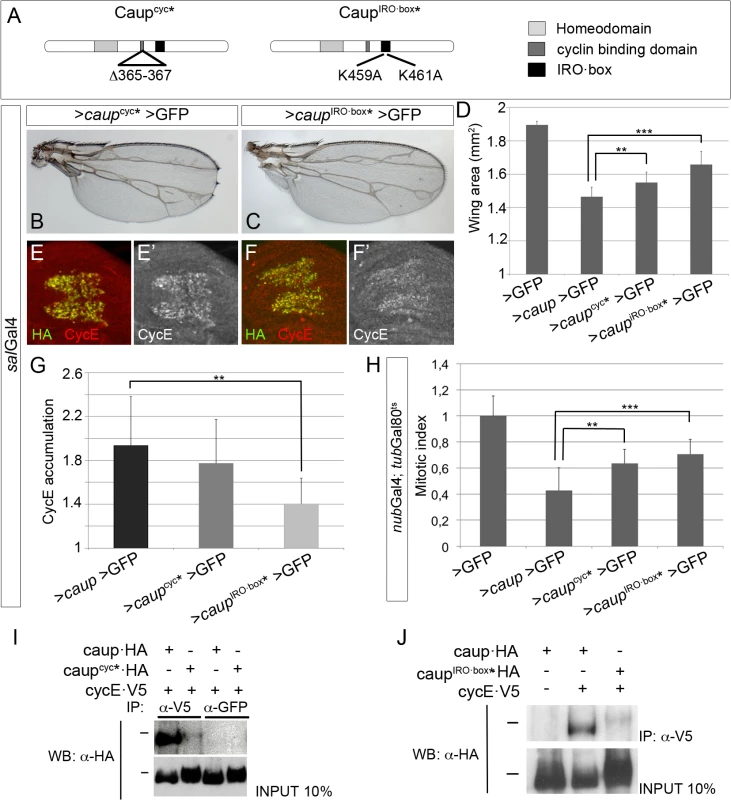
(A) Domain structure of Caup. The amino acid changed in the Cyc-binding domain and IRO·box in the novel mutants are indicated. (B, D) Representative wing phenotypes associated to Caupcyc* and CaupIRO·box* over-expression (B, C) and quantification of wing sizes of flies over-expressing the indicated transgenes (D). (E-G) Accumulation of CycE in wing imaginal cells that ectopically express Caupcyc* or CaupIRO·box*, quantified in G. (B-G, over-expression driven by salGal4). (H) Mitotic index in the nub territory of wing discs over-expressing the indicated transgenes. In all cases, quantifications are shown in relation to those performed in salGal4>caup>GFP or salGal4>GFP control wing discs and wings from larvae reared in parallel to the experimental ones. (I, J) Interaction of the different Caup proteins with CycE-containing complexes. Western blots of protein extracts from S2 cells expressing CycE-V5 and the different Caup-HA proteins, immunoprecipitated with the indicated antibodies and probed with anti-HA. Black bars indicate the position of the 100 kDa protein marker. These results suggest that Caup may be interacting with CycE-containing complexes through additional domain(s). Iro/Irx proteins harbour a conserved stretch of 14 amino acids, the IRO·box, whose function is unknown [11]. We mutagenized it changing its two conserved positively charged amino acids into Alanine (CaupIRO·box*, Fig 5A) and assayed its activity in vivo and its ability to interact with CycE-containing complexes as described for Caupcyc*. CaupIRO·box* was much less effective than wild-type Caup and Caupcyc* in interfering with cell cycle progression as shown by its effect on the mitotic index (Fig 5H), wing size (Fig 5C and 5D) and CycE accumulation (Fig 5F and 5G). Accordingly, CaupIRO·box* showed a highly reduced ability to co-immunoprecipitate with CycE (Fig 5J). Since Caupcyc* and CaupIRO·box* were still able to repress cell proliferation, albeit less than wild-type Caup, we generated a double mutant caupcyc* - IRO·box*. It still reduced wing size when over-expressed to a similar extent than CaupIRO·box* (S7D and S7E Fig).
The functional differences observed between wild-type Caup, Caupcyc* and CaupIRO·box* could not be attributed to an altered sub-cellular localization, significantly different levels of expression or stability, since these were similar (S7A–S7C and S7F Fig). Both Caupcyc* and CaupIRO·box* retained the ability to act as transcriptional regulators (monitored by repression of fng, a direct target of Iro genes [42], Fig 6K–6L’) and accordingly, over-expression of CaupIRO·box* in the eye disc prevented eye development (S7I Fig). Thus, these results suggest that Caup inhibits the activity of the CycE/Cdk2 complex by physical interaction mediated, at least in part, by both the Cyc-binding domain and the IRO·box rather than by a transcriptional-dependent mechanism.
Fig. 6. Functional analysis of homeodomain-mutant Caup proteins. 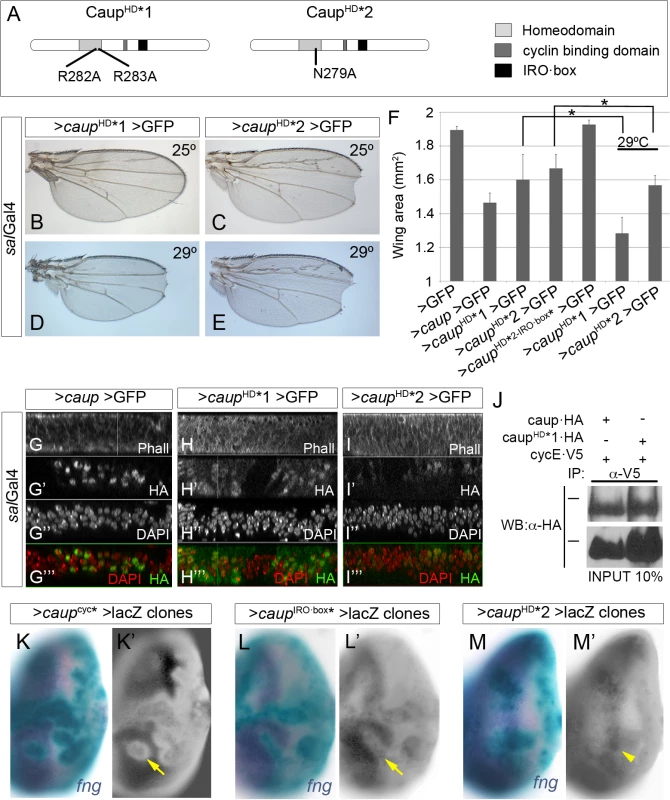
(A) Domain structure of Caup. The position of the point mutations generated in the homeodomain of CaupHD*1 and CaupHD*2 proteins is indicated. (B-E) Representative wing phenotypes associated to the over-expression of caupHD*1 or caupHD*2 at the indicated temperatures. (F) Wing areas of flies expressing the indicated transgenes driven by salGal4 at 25°C, save when otherwise indicated. (G-I”‘) Sub-cellular localization of the different Caup proteins. Wild-type Caup localized to the cell nuclei (G-G”‘). CaupHD*1 (H-H”‘) and CaupHD*2 (I-I”‘) are also found diffusely distributed in the cytosol. H- I”‘ images were taken with higher laser intensity than G-G”‘ because CaupHD*1 and CaupHD*2 accumulate at lower levels than wild-type Caup. (J) Interaction of CaupHD*1 with CycE-containing complexes. Western blots of protein extracts from S2 cells expressing CycE-V5 and the indicated Caup-HA proteins, immunoprecipitated with anti-V5 antibody and probed with anti-HA. Black bars indicate the position of the 100 kDa protein marker. (K-M´) Transcriptional activity of different Caup* proteins. Clones of cells expressing caup* and lacZ are marked by X-Gal staining (green). fng mRNA (in situ hybridization) is shown in blue (K, L and M) and separately in K’, L’ and M’. Caupcyc* (K, K’) and CaupIRO·box* (L, L’) cell- autonomously repress fng expression (arrows) (The apparent decrease in fng expression around the clones over expressing Caupcyc* or CaupIRObox* is due to the epithelial folds that surround them, as previously shown for caup over-expressing clones [60]). CaupHD*2 does not repress fng expression (arrowhead; M, M´). To further support this conclusion, we generated additional Caup mutants devoid of transcription factor activity by point mutations at key amino acids of the recognition helix of the homeodomain [43, 44] (CaupHD* 1 and CaupHD* 2, Fig 6A). These mutant proteins were apparently less effective than wild-type Caup in restricting wing disc growth (Fig 6B, 6C and 6F). However, they were expressed at very reduced levels and showed both cytosolic and nuclear accumulation (Fig 6G–6I’ and S7G Fig), which could account for their low effect. Indeed, when expression was increased (flies raised at 29°C), they strongly reduced wing size (Fig 6D–6F). As expected, and even upon enforced expression, CaupHD* 1 or CaupHD* 2 were unable to repress fng expression and to prevent eye formation, although they notably reduced eye size (Fig 6M and 6M’ and S7J Fig). In S2 cells, CaupHD*1 and CaupHD*2 co-immunoprecipitated with CycE similarly to wild-type Caup (Fig 6J and S7K Fig). Moreover, the ability of CaupHD*2 to reduce wing size was abolished when this protein was additionally mutated at the IRO·box (Fig 6F). Hence these data support the binding of Caup to CycE-containing complexes, mainly through the IRO·box, as the main molecular mechanism for its function in the control of the cell cycle.
Iro proteins regulate growth in Drosophila tumour models
Our results demonstrated the ability of Iro proteins to restrict cell cycle progression during normal development. Next, we addressed whether they were able to do so in Drosophila tumour-like models.
Over-expression of the Notch ligand Delta (Dl) causes the development of slightly enlarged eyes (eyGal4>Dl>lacZ flies, Fig 7A) and provides a sensitized genetic background useful to identify genes affecting cell proliferation and tumorigenesis [45]. We tested whether reduced activity of any of the Iro genes affected the size of eyGal4>Dl>lacZ eyes. Indeed, while partial depletion of Caup on its own had no discernible effect on eye size (S8B Fig), it increased both the size and the number of eyes that showed severe folding (Fig 7A and 7B). Similar enhancement of this mutant phenotype was obtained by co-expressing Dl and RNAi constructs targeted to ara or mirr (S8A, S8C and S8D Fig) or in combination with iroEGP7/+ (Fig 7C, 61% of the eyGal4>Dl>iroEGP7/+ eyes were enlarged compared with 39% of the eyes in eyGal4>Dl control flies).
Fig. 7. The levels of Iro proteins modulate tumour-like growth. 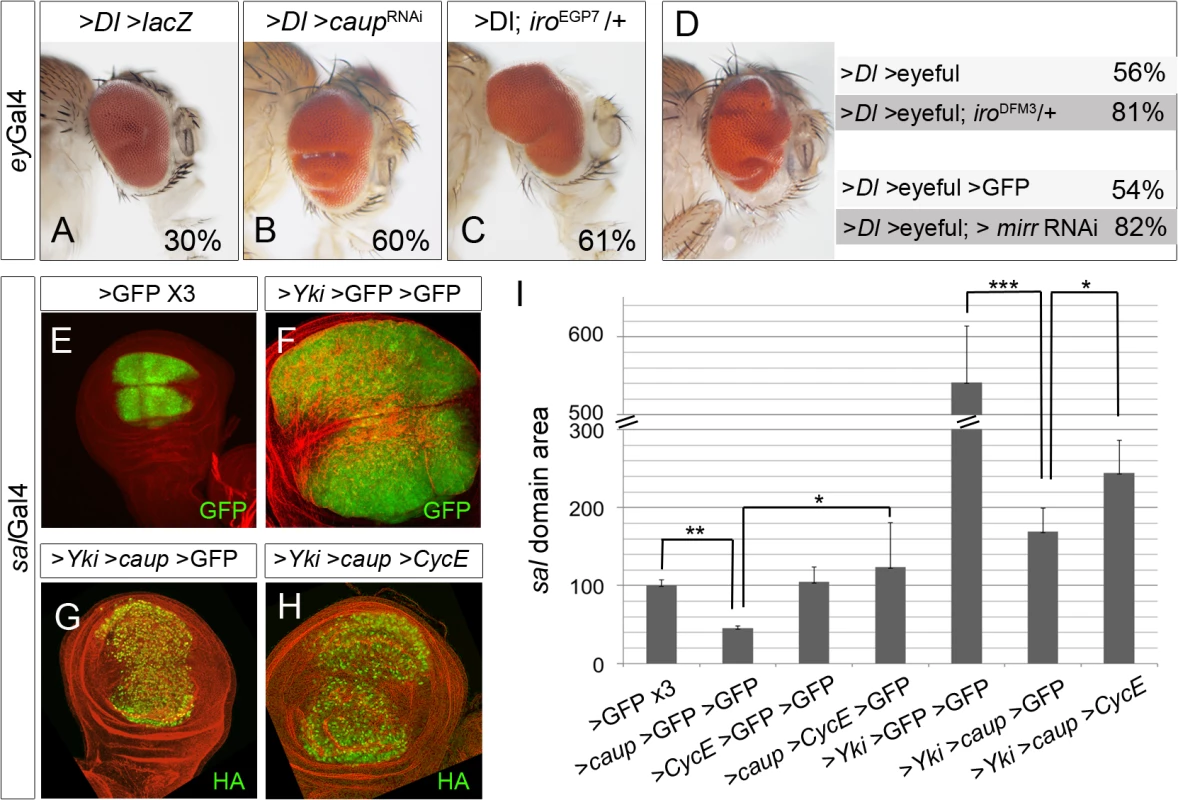
(A-C) Depletion of Iro proteins enhances eye growth in the sensitized background eyGal4>Dl>LacZ (note the enlarged and folded eyes in B, C, compare with A). Representative eyes are shown, along with the percentage of enlarged eyes for each genotype (average from two independent experiments, n>80 each). Flies were raised at 29°C. (D) Reduction of iro function (iroDFM3/+, or mirr depletion) enhances tumour-like growth in the >Dl >eyeful tumour model. (Left) Representative enlarged tumourous eye. (Right) Percentage of enlarged eyes in flies of the indicated genotypes (n>100, average value of three independent experiments). (E-I) Over-expression of caup reduces yki-induced overgrowth by CycE /Cdk2 inhibition. Compare the size of the sal domain (in green) in wing discs of the indicated genotypes. (G) Quantification of the area of the sal domain in third instar wing discs. Size domain was normalized to that of a sal>GFP>GFP>GFP wing discs (*p< 0.0001; **p<0.01). Discs are counterstained with Phalloidin (red). Co-expression in the eye disc (driven by eyGal4) of Dl and the epigenetic silencers Pipsqueak and Lola (referred to as >Dl >eyeful flies) induces the formation of tumour-like overgrowths in the eye [45]. Frequency of tumour formation was enhanced when >Dl >eyeful flies were in addition heterozygous for iroDFM3 or depleted of Mirr (Fig 7D).
Thus, Iro depletion enhanced tumorous growth in the eye. Next we assayed whether, conversely, over-expression of caup reduced the overgrowth of the wing disc in another tumoral model. The Hippo pathway controls organ size in Drosophila and vertebrates by a coordinated regulation of proliferation and apoptosis and its dysfunction is frequently detected in human cancers [46]. Over-expression of the downstream component of the Hippo pathway yorkie (yki, salGal4>yki) increased the size of the territory where it is expressed (Fig 7E, 7F and 7I). We observed that co-expression of caup alleviated the overgrowth caused by yki (Fig 7G and 7I). One of the effects of yki over-expression is the activation of cycE transcription ([47], S6B Fig). Therefore, we hypothesized that the phenotypic suppression by Caup could be due to CycE/Cdk2 inhibition. Indeed, cycE co-expression partially reverted the effect of caup on yki-induced overgrowth (Fig 7E–7I). In sum, our data suggest a role of Iro genes as TSGs in Drosophila.
Discussion
The identification of genes that control cell proliferation is paramount in developmental and cancer biology. The Iroquois proteins play multiple roles in regionalization and patterning during Drosophila development (reviewed in [12]). Here we show that they are also involved in the control of cell proliferation and, interestingly for homeodomain-containing proteins, they appear to do so by a non-transcriptional mechanism. This novel function of Iro genes would help developmental fields to attain their correct size and, if altered by Iro down regulation, could be a critical step for tumour progression.
We have analyzed iro hypomorphic and over-expression conditions and found that Iro proteins negatively control the G1-S transition of the cell cycle. caup over-expression impaired the activity of CycE/Cdk2 complex, while simultaneously increased the level of CycE protein. Still, CycE appears to be a limiting factor since its exogenous administration restores cell proliferation, while its reduction enhances it. The presence of Caup in CycE-containing protein complexes allow us to propose that this physical interaction inhibits CycE/Cdk2 activity thus slowing down cell proliferation. This hypothesis is supported by our observation that Caupcyc* and CaupIRO·box* mutant proteins show both impaired ability to co-immunoprecipitate with CycE and to restrict cell cycle progression. Although not experimentally demonstrated, we speculate that Caup may interact with CycE and Cdk2 containing complexes and inhibit their activity by preventing substrate recognition and/or stabilizing p21 binding. Further work is required to determine more precisely these molecular interactions. Since Caupcyc*IRO·box* still retains some ability to repress cell proliferation, we presume that either the functionality of these domains was not completely abolished by the mutations generated or the existence of additional unidentified interacting sites.
Although other homeobox proteins (and also some epigenetic regulators) have been shown to modulate the activity of cell cycle regulators by protein-protein interaction, many of them do it through transcriptional regulation [48–50]. We can rule out a transcriptional effect of Caup on cell cycle regulation since transcriptionally inactive CaupHD*1 and CaupHD*2 are still able to inhibit cell cycle progression.
Iro proteins play redundant roles in several developmental contexts [14, 15]. Here we show that the three of them are able to repress cell cycle progression when over-expressed and that this effect is abrogated by co-expression of cycE. The presence of putative Cyclin binding motives and the high conservation of the IRO·box in the Iro proteins [11] led us to propose that Ara and Mirr may also physically interact with CycE containing complexes. Since we found that the penetrance of the dorsal eye enlargement phenotype increases by reducing the overall amount of Iro proteins, we suggest that they may act in a redundant manner to modulate CycE/Cdk2 activity. Alternatively, the three Iro proteins may be functioning in a stoichiometric complex, this explaining why depletion of only one of them causes eye enlargement.
The present results suggest a novel role of Iro proteins as cell-autonomous regulators of the growth of the domains of the imaginal discs where they are expressed. Furthermore, our results fit to a current model that suggests that growth of territorial fields modulates the response of cells to morphogens (reviewed in [3]). In the eye discs, the ability of Decapentaplegic (Dpp) to induce retina differentiation is counteracted by Wg emanating from the anterior-most region of the discs (reviewed in [18] until the disc attains a size such that dpp expressing cells are beyond the range of action of Wg [7]. Accordingly, we suggest that the enhanced cell proliferation found in iro mutant discs, would enlarge the physical separation between Wg - and Dpp-expressing cells in the dorsal domain, thus increasing the efficiency of Dpp signalling and causing dorsal eye enlargement.
In analogy with this model for eye disc development, specification of the wing driven by Wg in the distal part of the wing disc is counteracted by the Vein morphogen, which spreads from the most proximal part of the wing disc (reviewed in [3]). In this scenario, reduction of the size of the distal wing disc by inhibition of cell proliferation prevents wing development (with the concomitant generation of a notum-like tissue, as shown in [8] and in this work), by facilitating the inhibition of Wg by Vein. Interestingly, Vein activates Iro gene expression in the notum region [38, 51] while Wg do so in the dorsal eye disc [14, 52–55]. Thus, we propose that Iro genes could provide a molecular mechanism that allow the ligands Vein (in the notum) and Wg (in the dorsal eye) to regulate the size of the morphogenetic field in which they operate.
Our results further suggest that a direct regulation of cell cycle progression by Iro/Irx proteins may be relevant for tumorigenesis. Thus, tumorous-like growth was observed in the eye imaginal discs when iro function was reduced in a sensitized genetic background (such as ey>Dl or ey>Dl >eyeful flies). Conversely, we show the ability of caup over-expression to counteract the overgrowth induced by Yki in imaginal discs, and that this is partially mediated by cycE/cdk2 inactivation. These data suggest a role of Iro genes as TSGs in Drosophila and agree with the association found between loss or reduced expression of members of Irx gene family and certain types of human cancer [20–23]. Note however that the role of Iro/Irx genes in tumorigenesis may be cell type-dependent since in some cases they appear to act as oncogenes [55 56]. Considering the presence of the IRO box [11] and of putative Cyclin-binding domains in Irx proteins (http://elm.eu.org), we hypothesize that some Irx mutations may contribute to cancer progression in vertebrates by increasing the activity of the CycE/Cdk2 complex and thus accelerating the G1-S transition, a key step frequently affected in cancer cells [57].
Materials and Methods
Site-directed mutagenesis of Caup
The following Caup mutations (caup-mut) were generated: Caupcyc*, deletion of amino-acids 365 to 367 (RGL) of the Caup putative Cyclin-binding domain (RGLAP); CaupIRO·box*, substitution of the only two positively charged amino acids of the IRO·box, Lysine 459 and Lysine 461 [11] to Ala; CaupHD*1, substitution of homeodomain Arginine 282 and Arginine 283 to Alanine [43] and CaupHD*2, substitution of homeodomain Asparagine 279 to Alanine [44]. Mutants were obtained by site-directed mutagenesis (Quick-Change system, Stratagene) of wild-type caup cDNA [9] or caup-mut cDNA (this work) with the primers indicated in Supplemental Experimental Procedures.
Over-expression experiments
Larvae expressing UAS-transgenes driven by salGal4; MD638Gal4 or eyGal4 were raised at 25°C unless otherwise indicated. To increase the penetrance of the dorsally enlarged eye phenotype, eyGal4; 2x UAS-mirr RNAi larvae (Fig 1F) were raised at 29°C. To avoid the embryonic lethality associated with caup over-expression driven by ap and hh Gal4, we combined these lines with a tubGal80ts transgene [30]. Below 29°C, Gal80 inhibits Gal4 activity. Gal4 line; UAS-iro gene/tubGal80ts larvae were raised at 17°C, and transferred to 29°C 16 hours prior to dissection. In all experiments, the number of UAS genes was kept constant to avoid differences due to Gal4 titration. UAS-caup-HA and UAS-caup*-HA transgenic flies were obtained by the site-specific integration system at the 51D cytogenetic position [58] to get similar expression levels.
Flow cytometry analysis
50 wing discs were dissected from iroDFM3/iroGal4, UAS-GFP larvae at 100–120h after egg laying. FACS analysis was done according to [28]. Cells were sorted by GFP expression using FACSCVantage SE (BD Biosciences) and cell cycle profiles were determined by Hoescht flourescence using a FACSCalibur flow cytometer (Becton Dickinson). Data from five independent experiments were analyzed using the FlowJo software and Dean-Jett-Fox model.
Cell transfection and co-immunoprecipitation
Drosophila S2 cells were cultured in Insect-XPRESS media (Lonza) supplemented with 7% fetal calf serum and transfected using Nucleofector Technology (Lonza), according to the manufacturer’s specifications. caup-HA [59] and caup-mut-HA (this work) were cloned downstream of the constitutive promoter of the Drosophila Actin 5C gene in the pAc5.1 B plasmid (Invitrogen). The full-length cycE ORF was amplified from DGRC cDNA clone LD22682 using the following primers: 5’GAATCCGGCCGTACAATTATG3’ and 5’TCTAGAGGGATTGCTTCTAC3’ and cloned in pAc5.1 A (Invitrogen). Transfected cells were cultured during 48 hours before obtaining cell lysates by standard procedures. Antibodies used in immunoprecipitations and immunoblots were mouse anti-V5 (Invitrogen), mouse anti-GFP (Roche) and rat anti-HA (Roche). Similar results were obtained in at least two independent experiments.
Wing size, mitotic index and pixel intensity determination
Areas of the sal domain of wing imaginal discs (n = 10) and of wings from female flies (n = 10; mounted in lactic acid /ethanol, 6 : 5) and pixel intensity of CycE-expressing cells were measured with Adobe Photoshop CS4. The values of CycE pixel intensity for each wing disc correspond to the ratio between average pixel intensity at the sal territory and the average pixel intensity at the adjacent territory (n = 10). To calculate the relative mitotic index, the number of pH3 expressing cells per area was quantified with Adobe Photoshop CS4 and then normalized to the mitotic index in the same region in control discs (n = 10).
Statistical analysis
Data are shown as arithmetic mean ± standard deviation (SD, indicated by error bars). The statistical difference between groups of data was examined by Student’s t-test. p<0.05 was considered statistically significant.
Supporting Information
Zdroje
1. Schwank G, Basler K (2010) Regulation of organ growth by morphogen gradients. Cold Spring Harb Perspect Biol. 2: a001669. doi: 10.1101/cshperspect.a001669 20182606
2. Amore G, Casares F (2010) Size matters: the contribution of cell proliferation to the progression of the specification Drosophila eye gene regulatory network. Dev Biol 344 : 569–577. doi: 10.1016/j.ydbio.2010.06.015 20599903
3. Dekanty A, Milan M (2011) The interplay between morphogens and tissue growth. EMBO Rep 12 : 1003–1010. doi: 10.1038/embor.2011.172 21886183
4. Towers M, Tickle C (2009) Growing models of vertebrate limb development. Development 136 : 179–190. doi: 10.1242/dev.024158 19103802
5. Cooper KL, Hu JK, ten Berge D, Fernandez-Teran M, Ros MA, Tabin CJ (2011) Initiation of proximal-distal patterning in the vertebrate limb by signals and growth. Science 332 : 1083–1086. doi: 10.1126/science.1199499 21617075
6. Rosello-Diez A, Ros MA, Torres M (2011) Diffusible signals, not autonomous mechanisms, determine the main proximodistal limb subdivision. Science 332 : 1086–1088. doi: 10.1126/science.1199489 21617076
7. Kenyon KL, Ranade SS, Curtiss J, Mlodzik M, Pignoni F (2003) Coordinating proliferation and tissue specification to promote regional identity in the Drosophila head. Dev Cell 5 : 403–414. 12967560
8. Rafel N, Milan M (2008) Notch signalling coordinates tissue growth and wing fate specification in Drosophila. Development 135 : 3995–4001. doi: 10.1242/dev.027789 18987026
9. Gomez-Skarmeta JL, Diez del Corral R, de la Calle-Mustienes E, Ferres-Marco D, Modolell J (1996) Araucan and caupolican, two members of the novel iroquois complex, encode homeoproteins that control proneural and vein-forming genes. Cell 85 : 95–105. 8620542
10. McNeill H, Yang CH, Brodsky M, Ungos J, Simon MA (1997) mirror encodes a novel PBX-class homeoprotein that functions in the definition of the dorsal-ventral border in the Drosophila eye. Genes Dev 11 : 1073–1082. 9136934
11. Burglin TR (1997) Analysis of TALE superclass homeobox genes (MEIS, PBC, KNOX, Iroquois, TGIF) reveals a novel domain conserved between plants and animals. Nucleic Acids Res 25 : 4173–4180. 9336443
12. Cavodeassi F, Modolell J, Gomez-Skarmeta JL (2001) The Iroquois family of genes: from body building to neural patterning. Development 128 : 2847–2855. 11532909
13. Cavodeassi F, Modolell J, Campuzano S (2000) The Iroquois homeobox genes function as dorsal selectors in the Drosophila head. Development 127 : 1921–1929. 10751180
14. Cavodeassi F, Diez Del Corral R, Campuzano S, Dominguez M (1999) Compartments and organising boundaries in the Drosophila eye: the role of the homeodomain Iroquois proteins. Development 126 : 4933–4942. 10529412
15. Diez del Corral R, Aroca P, Gomez-Skarmeta JL, Cavodeassi F, Modolell J (1999) The Iroquois homeodomain proteins are required to specify body wall identity in Drosophila. Genes Dev 13 : 1754–1761. 10398687
16. Cho KO, Choi KW. (1998) Fringe is essential for mirror symmetry and morphogenesis in the Drosophila eye. Nature 396 : 272–276. 9834034
17. Dominguez M, de Celis JF (1998) A dorsal/ventral boundary established by Notch controls growth and polarity in the Drosophila eye. Nature 396 : 276–278. 9834035
18. Dominguez M, Casares F (2005) Organ specification-growth control connection: new in-sights from the Drosophila eye-antennal disc. Dev Dyn 232 : 673–684. 15704149
19. Pichaud F, Casares F (2000) homothorax and iroquois-C genes are required for the establishment of territories within the developing eye disc. Mech Dev 96 : 15–25. 10940621
20. Bennett KL, Karpenko M, Lin MT, Claus R, Arab K, Dyckhoff G, et al. (2008) Frequently methylated tumor suppressor genes in head and neck squamous cell carcinoma. Cancer Res 68 : 4494–4499. doi: 10.1158/0008-5472.CAN-07-6509 18559491
21. Lu Y, Yu Y, Zhu Z, Xu H, Ji J, Bu L, et al. (2005) Identification of a new target region by loss of heterozygosity at 5p15.33 in sporadic gastric carcinomas: genotype and phenotype related. Cancer Lett 224 : 329–337. 15914283
22. Guo X, Liu W, Pan Y, Ni P, Ji J, Guo L, et al. (2010) Homeobox gene IRX1 is a tumor suppressor gene in gastric carcinoma. Oncogene 29 : 3908–3920. doi: 10.1038/onc.2010.143 20440264
23. Nguyen HH, Takata R, Akamatsu S, Shigemizu D, Tsunoda T, Furihata M, et al. (2012) IRX4 at 5p15 suppresses prostate cancer growth through the interaction with vitamin D receptor, conferring prostate cancer susceptibility. Hum Mol Genet 21 : 2076–2085. doi: 10.1093/hmg/dds025 22323358
24. Wolff T, Ready DF (1993) Pattern formation in the Drosophila retina. In: Bate M, Martinez-Arias A, editors. The development of Drosophila melanogaster. Cold Spring Harbor Lab Press, Vol II p. 1277–1325.
25. Treisman JE, Rubin GM (1995) wingless inhibits morphogenetic furrow movement in the Drosophila eye disc. Development 121 : 3519–3527. 8582266
26. Mazzoni EO, Celik A, Wernet MF, Vasiliauskas D, Johnston RJ, Cook TA, et al. (2008) Iroquois complex genes induce co-expression of rhodopsins in Drosophila. PLoS Biol 6:e97. doi: 10.1371/journal.pbio.0060097 18433293
27. Organista MF, De Celis JF (2013) The Spalt transcription factors regulate cell proliferation, survival and epithelial integrity downstream of the Decapentaplegic signalling pathway. Biol Open 2 : 37–48. doi: 10.1242/bio.20123038 23336075
28. Neufeld TP, de la Cruz AF, Johnston LA, Edgar BA (1998) Coordination of growth and cell division in the Drosophila wing. Cell 93 : 1183–1193. 9657151
29. Edgar BA, O'Farrell PH (1990) The three postblastoderm cell cycles of Drosophila embryogenesis are regulated in G2 by string. Cell 62 : 469–480. 2199063
30. McGuire SE, Roman G, Davis RL (2004) Gene expression systems in Drosophila: a synthesis of time and space. Trends Genet 20 : 384–391. 15262411
31. Cruz C, Glavic A, Casado M, de Celis JF (2009) A gain-of-function screen identifying genes required for growth and pattern formation of the Drosophila melanogaster wing. Genetics 183 : 1005–1026. doi: 10.1534/genetics.109.107748 19737745
32. Lee LA, Orr-Weaver TL (2003) Regulation of cell cycles in Drosophila development: intrinsic and extrinsic cues. Annu Rev Genet 37 : 545–578. 14616073
33. Wang SL, Hawkins CJ, Yoo SJ, Muller HA, Hay BA (1999) The Drosophila caspase inhibitor DIAP1 is essential for cell survival and is negatively regulated by HID. Cell 98 : 453–463. 10481910
34. Chen P, Nordstrom W, Gish B, Abrams JM (1996) grim, a novel cell death gene in Drosophila. Genes Dev 10 : 1773–1782. 8698237
35. Waldron JA, Jones CI, Towler BP, Pashler AL, Grima DP, Hebbes S, et al. (2015) Xrn1/Pacman affects apoptosis and regulates expression of hid and reaper. Biol Open. 4 : 649–660. doi: 10.1242/bio.201410199 25836675
36. Moberg KH, Bell DW, Wahrer DC, Haber DA, Hariharan IK (2001) Archipelago regulates Cyclin E levels in Drosophila and is mutated in human cancer cell lines. Nature 413 : 311–316. 11565033
37. Aldaz S, Morata G, Azpiazu N (2003) The Pax-homeobox gene eyegone is involved in the subdivision of the thorax of Drosophila. Development 130 : 4473–4482. 12900462
38. Wang SH, Simcox A, Campbell G (2000) Dual role for Drosophila epidermal growth factor re.ceptor signaling in early wing disc development. Genes Dev 14 : 2271–2276. 10995384
39. de Navascues J, Modolell J (2007) tailup, a LIM-HD gene, and Iro-C cooperate in Drosophila dorsal mesothorax specification. Development 134 : 1779–1788. 17409113
40. White AE, Leslie ME, Calvi BR, Marzluff WF, Duronio RJ (2007) Developmental and cell cycle regulation of the Drosophila histone locus body. Mol Biol Cell 18 : 2491–2502. 17442888
41. Clurman BE, Sheaff RJ, Thress K, Groudine M, Roberts JM (1996) Turnover of cyclin E by the ubiquitin-proteasome pathway is regulated by cdk2 binding and cyclin phosphorylation. Genes Dev 10 : 1979–1690. 8769642
42. Bilioni A, Craig G, Hill C, McNeill H (2005) Iroquois transcription factors recognize a unique motif to mediate transcriptional repression in vivo. Proc Natl Acad Sci U S A 102 : 14671–14676. 16203991
43. Noyes MB, Christensen RG, Wakabayashi A, Stormo GD, Brodsky MH, Wolfe SA (2008) Analysis of homeodomain specificities allows the family-wide prediction of preferred recognition sites. Cell 133 : 1277–1289. doi: 10.1016/j.cell.2008.05.023 18585360
44. Ades SE, Sauer RT (1995) Specificity of minor-groove and major-groove interactions in a homeodomain-DNA complex. Biochemistry 34 : 14601–14608. 7578067
45. Ferres-Marco D, Gutierrez-Garcia I, Vallejo DM, Bolivar J, Gutierrez-Avino FJ, Dominguez M (2006) Epigenetic silencers and Notch collaborate to promote malignant tumours by Rb silencing. Nature 439 : 430–436. 16437107
46. Yu FX, Guan KL (2013) The Hippo pathway: regulators and regulations. Genes Dev 27 : 355–371. doi: 10.1101/gad.210773.112 23431053
47. Zhao B, Tumaneng K, Guan KL (2011) The Hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nature Cell Biol 13 : 877–883. doi: 10.1038/ncb2303 21808241
48. Del Bene F, Wittbrodt J (2005). Cell cycle control by homeobox genes in development and disease. Semin Cell Dev Biol 16 : 449–460. 15840452
49. Baig J, Chanut F, Kornberg TB, Klebes A (2010) The chromatin-remodeling protein Osa interacts with CyclinE in Drosophila eye imaginal discs. Genetics 184 : 731–44. doi: 10.1534/genetics.109.109967 20008573
50. Lecona E, Rojas LA, Bonasio R, Johnston A, Fernandez-Capetillo O, Reinberg D (2013) Polycomb protein SCML2 regulates the cell cycle by binding and modulating CDK/CYCLIN/p21 complexes. PLoS Biol 11: e1001737. doi: 10.1371/journal.pbio.1001737 24358021
51. Zecca M, Struhl G (2002) Control of growth and patterning of the Drosophila wing imaginal disc by EGFR-mediated signaling. Development 129 : 1369–1376. 11880346
52. Heberlein U, Borod ER, Chanut FA (1998) Dorsoventral patterning in the Drosophila retina by wingless. Development 125 : 567–577. 9435278
53. Lee JD, Treisman JE (2001) The role of Wingless signaling in establishing the anteroposterior and dorsoventral axes of the eye disc. Development 128 : 1519–1529. 11290291
54. Maurel-Zaffran C, Treisman JE (2000) pannier acts upstream of wingless to direct dorsal eye disc development in Drosophila. Development 127 : 1007–1016. 10662640
55. Myrthue A, Rademacher BL, Pittsenbarger J, Kutyba-Brooks B, Gantner M, Qian DZ, et al. (2008) The iroquois homeobox gene 5 is regulated by 1,25-dihydroxyvitamin D3 in human prostate cancer and regulates apoptosis and the cell cycle in LNCaP prostate cancer cells. Clin Cancer Res 14 : 3562–3570. doi: 10.1158/1078-0432.CCR-07-4649 18519790
56. Martorell O, Barriga FM, Merlos-Suarez A, Stephan-Otto Attolini C, Casanova J, Batlle E, et al. (2014) Iro/IRX transcription factors negatively regulate Dpp/TGF-beta pathway activity during intestinal tumorigenesis. EMBO Rep 15 : 1210–1218. doi: 10.15252/embr.201438622 25296644
57. Hwang HC, Clurman BE (2005) Cyclin E in normal and neoplastic cell cycles. Oncogene 24 : 2776–2786. 15838514
58. Bischof J, Maeda RK, Hediger M, Karch F, Basler K (2007) An optimized transgenesis system for Drosophila using germ-line-specific phiC31 integrases. Proc Natl Acad Sci U S A 104 : 3312–3317. 17360644
59. Carrasco-Rando M, Tutor AS, Prieto-Sanchez S, Gonzalez-Perez E, Barrios N, Letizia A, et al. (2011) Drosophila araucan and caupolican integrate intrinsic and signalling inputs for the acquisition by muscle progenitors of the lateral transverse fate. PLoS Genet 7: e1002186. doi: 10.1371/journal.pgen.1002186 21811416
60. Villa-Cuesta E, Gonzalez-Perez E, Modolell J (2007) Apposition of iroquois expressing and non-expressing cells leads to cell sorting and fold formation in the Drosophila imaginal wing disc. BMC Dev Biol 7 : 106. 17880703
Štítky
Genetika Reprodukční medicína
Článek Loss and Gain of Natural Killer Cell Receptor Function in an African Hunter-Gatherer PopulationČlánek Let-7 Represses Carcinogenesis and a Stem Cell Phenotype in the Intestine via Regulation of Hmga2Článek Binding of Multiple Rap1 Proteins Stimulates Chromosome Breakage Induction during DNA ReplicationČlánek SLIRP Regulates the Rate of Mitochondrial Protein Synthesis and Protects LRPPRC from DegradationČlánek Protein Composition of Infectious Spores Reveals Novel Sexual Development and Germination Factors inČlánek The Formin Diaphanous Regulates Myoblast Fusion through Actin Polymerization and Arp2/3 RegulationČlánek Runx1 Transcription Factor Is Required for Myoblasts Proliferation during Muscle Regeneration
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2015 Číslo 8
-
Všechny články tohoto čísla
- Putting the Brakes on Huntington Disease in a Mouse Experimental Model
- Identification of Driving Fusion Genes and Genomic Landscape of Medullary Thyroid Cancer
- Evidence for Retromutagenesis as a Mechanism for Adaptive Mutation in
- TSPO, a Mitochondrial Outer Membrane Protein, Controls Ethanol-Related Behaviors in
- Evidence for Lysosome Depletion and Impaired Autophagic Clearance in Hereditary Spastic Paraplegia Type SPG11
- Loss and Gain of Natural Killer Cell Receptor Function in an African Hunter-Gatherer Population
- Trans-Reactivation: A New Epigenetic Phenomenon Underlying Transcriptional Reactivation of Silenced Genes
- Early Developmental and Evolutionary Origins of Gene Body DNA Methylation Patterns in Mammalian Placentas
- Strong Selective Sweeps on the X Chromosome in the Human-Chimpanzee Ancestor Explain Its Low Divergence
- Dominance of Deleterious Alleles Controls the Response to a Population Bottleneck
- Transient 1a Induction Defines the Wound Epidermis during Zebrafish Fin Regeneration
- Systems Genetics Reveals the Functional Context of PCOS Loci and Identifies Genetic and Molecular Mechanisms of Disease Heterogeneity
- A Genome Scale Screen for Mutants with Delayed Exit from Mitosis: Ire1-Independent Induction of Autophagy Integrates ER Homeostasis into Mitotic Lifespan
- Non-synonymous FGD3 Variant as Positional Candidate for Disproportional Tall Stature Accounting for a Carcass Weight QTL () and Skeletal Dysplasia in Japanese Black Cattle
- The Relationship between Gene Network Structure and Expression Variation among Individuals and Species
- Calmodulin Methyltransferase Is Required for Growth, Muscle Strength, Somatosensory Development and Brain Function
- The Wnt Frizzled Receptor MOM-5 Regulates the UNC-5 Netrin Receptor through Small GTPase-Dependent Signaling to Determine the Polarity of Migrating Cells
- Nbs1 ChIP-Seq Identifies Off-Target DNA Double-Strand Breaks Induced by AID in Activated Splenic B Cells
- CCNYL1, but Not CCNY, Cooperates with CDK16 to Regulate Spermatogenesis in Mouse
- Evidence for a Common Origin of Blacksmiths and Cultivators in the Ethiopian Ari within the Last 4500 Years: Lessons for Clustering-Based Inference
- Of Fighting Flies, Mice, and Men: Are Some of the Molecular and Neuronal Mechanisms of Aggression Universal in the Animal Kingdom?
- Hypoxia and Temperature Regulated Morphogenesis in
- The Homeodomain Iroquois Proteins Control Cell Cycle Progression and Regulate the Size of Developmental Fields
- Evolution and Design Governing Signal Precision and Amplification in a Bacterial Chemosensory Pathway
- Rac1 Regulates Endometrial Secretory Function to Control Placental Development
- Let-7 Represses Carcinogenesis and a Stem Cell Phenotype in the Intestine via Regulation of Hmga2
- Functions as a Positive Regulator of Growth and Metabolism in
- The Nucleosome Acidic Patch Regulates the H2B K123 Monoubiquitylation Cascade and Transcription Elongation in
- Rhoptry Proteins ROP5 and ROP18 Are Major Murine Virulence Factors in Genetically Divergent South American Strains of
- Exon 7 Contributes to the Stable Localization of Xist RNA on the Inactive X-Chromosome
- Regulates Refractive Error and Myopia Development in Mice and Humans
- mTORC1 Prevents Preosteoblast Differentiation through the Notch Signaling Pathway
- Regulation of Gene Expression Patterns in Mosquito Reproduction
- Molecular Basis of Gene-Gene Interaction: Cyclic Cross-Regulation of Gene Expression and Post-GWAS Gene-Gene Interaction Involved in Atrial Fibrillation
- The Spalt Transcription Factors Generate the Transcriptional Landscape of the Wing Pouch Central Region
- Binding of Multiple Rap1 Proteins Stimulates Chromosome Breakage Induction during DNA Replication
- Functional Divergence in the Role of N-Linked Glycosylation in Smoothened Signaling
- YAP1 Exerts Its Transcriptional Control via TEAD-Mediated Activation of Enhancers
- Coordinated Evolution of Influenza A Surface Proteins
- The Evolutionary Potential of Phenotypic Mutations
- Genome-Wide Association and Trans-ethnic Meta-Analysis for Advanced Diabetic Kidney Disease: Family Investigation of Nephropathy and Diabetes (FIND)
- New Routes to Phylogeography: A Bayesian Structured Coalescent Approximation
- SLIRP Regulates the Rate of Mitochondrial Protein Synthesis and Protects LRPPRC from Degradation
- Satellite DNA Modulates Gene Expression in the Beetle after Heat Stress
- SHOEBOX Modulates Root Meristem Size in Rice through Dose-Dependent Effects of Gibberellins on Cell Elongation and Proliferation
- Reduced Crossover Interference and Increased ZMM-Independent Recombination in the Absence of Tel1/ATM
- Suppression of Somatic Expansion Delays the Onset of Pathophysiology in a Mouse Model of Huntington’s Disease
- Protein Composition of Infectious Spores Reveals Novel Sexual Development and Germination Factors in
- The Evolutionarily Conserved LIM Homeodomain Protein LIM-4/LHX6 Specifies the Terminal Identity of a Cholinergic and Peptidergic . Sensory/Inter/Motor Neuron-Type
- SmD1 Modulates the miRNA Pathway Independently of Its Pre-mRNA Splicing Function
- piRNAs Are Associated with Diverse Transgenerational Effects on Gene and Transposon Expression in a Hybrid Dysgenic Syndrome of .
- Retinoic Acid Signaling Regulates Differential Expression of the Tandemly-Duplicated Long Wavelength-Sensitive Cone Opsin Genes in Zebrafish
- The Formin Diaphanous Regulates Myoblast Fusion through Actin Polymerization and Arp2/3 Regulation
- Genome-Wide Analysis of PAPS1-Dependent Polyadenylation Identifies Novel Roles for Functionally Specialized Poly(A) Polymerases in
- Runx1 Transcription Factor Is Required for Myoblasts Proliferation during Muscle Regeneration
- Regulation of Mutagenic DNA Polymerase V Activation in Space and Time
- Variability of Gene Expression Identifies Transcriptional Regulators of Early Human Embryonic Development
- The Drosophila Gene Interacts Genetically with and Shows Female-Specific Effects of Divergence
- Functional Activation of the Flagellar Type III Secretion Export Apparatus
- Retrohoming of a Mobile Group II Intron in Human Cells Suggests How Eukaryotes Limit Group II Intron Proliferation
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Exon 7 Contributes to the Stable Localization of Xist RNA on the Inactive X-Chromosome
- YAP1 Exerts Its Transcriptional Control via TEAD-Mediated Activation of Enhancers
- SmD1 Modulates the miRNA Pathway Independently of Its Pre-mRNA Splicing Function
- Molecular Basis of Gene-Gene Interaction: Cyclic Cross-Regulation of Gene Expression and Post-GWAS Gene-Gene Interaction Involved in Atrial Fibrillation
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



