-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
Reklama, , and mutations in early-onset Alzheimer disease: A genetic screening study of familial and sporadic cases
In a cohort including patients with familial Alzheimer disease as well as sporadic cases of early-onset Alzheimer disease, Dominique Campion and colleagues identify previously unreported mutations to PSEN1 and PSEN2.
Published in the journal: . PLoS Med 14(3): e32767. doi:10.1371/journal.pmed.1002270
Category: Research Article
doi: https://doi.org/10.1371/journal.pmed.1002270Summary
In a cohort including patients with familial Alzheimer disease as well as sporadic cases of early-onset Alzheimer disease, Dominique Campion and colleagues identify previously unreported mutations to PSEN1 and PSEN2.
Introduction
Alzheimer disease (AD) (MIM #104300) is the most common form of dementia. However, early-onset AD (EOAD) constitutes a minority of patients, with an estimated prevalence of 41.2 per 100,000 persons at risk [1]. Among these forms, presenilin-1 (PSEN1) (MIM #104311), presenilin-2 (PSEN2) (MIM #600759) [2–5], and amyloid protein precursor (APP) (MIM #104760) mutations [6–8] and duplications [9] cause autosomal-dominant EOAD (AD-EOAD), the prevalence of which is estimated to be 5.3 per 100,000 persons at risk [1]. PSEN1 is the most commonly involved gene, with 221 mutations reported as pathogenic in the Alzforum database (www.alzforum.org/mutations). The second most commonly involved gene is APP, with 32 pathogenic mutations described, while 19 different PSEN2 pathogenic mutations have been reported. APP encodes the amyloid-β precursor protein, the processing of which by the β-secretase and the γ-secretase complex leads to the production of the amyloid β (Aβ) peptide, a key event in AD pathogeny. The aggregation of the Aβ peptide in the brain’s parenchyma indeed triggers a cascade of events leading to AD. Its aggregation in cerebromeningeal vessels leads to cerebral amyloid angiopathy (CAA), a condition frequently associated with AD and responsible for recurrent haemorrhagic strokes and white matter lesions. PSEN1 and PSEN2 encode the presenilins, which constitute the catalytic subunit of the γ-secretase complex (for review, see [10,11]). AD-EOAD causative mutations are thought to be responsible for the increased aggregation of the Aβ peptide in the brain’s parenchyma through one of the two following mechanisms: increased overall production of all Aβ species (e.g., APP duplications or APP mutations located around the β cleavage site) or production of a more aggregation-prone form of the Aβ peptide.
The power to detect genetic variations has dramatically improved over the last few years, but the interpretation of rare variants remains a challenge in a high proportion of cases. The pathogenicity of most APP, PSEN1, and PSEN2 variants has not yet been assessed through in vitro functional experiments. In cases of insufficient genetic evidence (i.e., lack or limited familial segregation or recurrence), definite pathogenicity of a given variant may therefore remain uncertain. An algorithm was proposed to classify those variants, based on (i) intrafamilial segregation, (ii) recurrence of the mutation in independent cases and association in case–control samples, (iii) residue conservation between PSEN1 and PSEN2 and residue localization on functional domains, and (iv) functional tests, when available [12]. Reporting patients carrying novel as well as previously known mutations along with the associated phenotypes will aid in classification of these variants and will eventually allow genetic counseling and inclusion in preventive trials for presymptomatic carriers [13].
We had previously described the PSENs and APP mutational spectrum in a large French series of families with an EOAD diagnosis in at least two first-degree relatives from two generations [14]. The aim of the present article is to report mutations in additional families included since our last 2012 update [14]. Furthermore, we add the results of the genetic screening of 129 sporadic EOAD patients with an age of onset (AOO) before 51. The involvement of PSEN1, PSEN2, and APP mutations in the genetics of sporadic EOAD has been scarcely studied. In particular, systematic genetic assessments of series of patients with youngest AOO who are at high risk to carry an AD-EOAD mutation were not reported before. In these patients, the family history can remain negative because of a censoring effect (i.e., death of the transmitting parent before EOAD onset) [15] or if the mutation occurs de novo (i.e., if it is not found in parents but occurs in the parental germline or as a postzygotic event) [16].
Materials and methods
The study was approved by the Paris Ile de France II ethics committee.
Subjects
EOAD subjects were referred to the National Reference Center for Early-Onset Alzheimer Patients (CNR-MAJ) from 28 university hospitals across France. For each patient, AD diagnosis was established using the National Institute of Aging–Alzheimer’s Association (NIA–AA) criteria [17]. All patients underwent a comprehensive clinical examination, including personal medical and family history and neuropsychological assessment. Search for mutations in APP, PSEN1, and PSEN2 genes was performed (i) in AD-EOAD presentations (i.e., if at least two first-degree relatives suffered from EOAD [AOO ≤65 y]) in two generations or (ii) in sporadic presentations if a patient without family history of AD had an age of onset before 51 y. No other exclusion criteria were applied. Familial cases (n = 63 mutation carriers belonging to 53 families, 42% males, mean AOO = 48 ± 5 y) were included in the 2012–2016 interval, whereas sporadic cases (n = 129, 44% males, mean AOO = 45 ± 2 y) were included from 1999 onwards. All patients were from European origin with the exception of five patients from African descent: three familial and two sporadic cases. Cerebrospinal fluid (CSF) AD biomarkers were assessed in 65% of the mutation carriers, and neuropathological examination was performed in 3 mutation carriers. A written consent to participate to the study was signed by every patient.
CSF analysis
CSF samples were obtained using a Sprotte needle in polypropylene collection tubes and aliquoted after centrifugation into polypropylene tubes (catalog number 62.610.201; Sarstedt, Nümbrecht, Germany), then frozen at −80°C within 1 h. Aβ42, Tau, and P-Tau measurements were performed using enzyme-linked immunosorbent assays (ELISA) (Fujirebio Europe N.V., Ghent, Belgium) according to the manufacturer’s instructions. The analysis of all biomarkers was performed in two duplicates and averaged for statistical analyses. Following values were used to define biochemical AD signature: Aβ42 < 700 pg/mL; Tau > 350 pg/mL, and P-Tau > 60 pg/mL. Each subject was classified according to the Paris, Lyon, Marseille (PLM) scale [18]: class 0, corresponding to no pathologic biomarkers; class 1, corresponding to 1/3 pathologic biomarkers; class 2, corresponding to 2/3 pathologic biomarkers; and class 3, with all three biomarkers being pathologic.
Genetic analyses
Genetic analyses were performed on DNA extracted from whole blood. Exons 2–12 of PSEN1 (NM_000021.3), exons 4–13 of PSEN2 (NM_000447.2), and exons 16 and 17 of APP (NM_000484.3) were analysed by Sanger sequencing. APP duplications and PSEN1 exon 9/10 deletion were detected using QMPSF (quantitative multiplex PCR of short fluorescent fragments). APOE genotype was determined for each subject by Sanger sequencing. Primers are available upon request. Guerreiro’s algorithm [12] and Alzforum (www.alzforum.org/mutations) database were used to classify each mutation’s pathogenicity.
In sporadic cases, when DNA was available for both unaffected parents, parenthood was checked using a package of four microsatellites markers, each with a heterozygosity index from 79 to 88%, and the presence of the mutation identified in the proband was assessed by Sanger sequencing.
Results
Update of the EOAD French series
We identified mutations in 53 previously unreported AD-EOAD families and in 18/129 sporadic cases, including 44 PSEN1, 2 PSEN2, and 20 APP mutations as well as five APP duplications. The total number of mutation carriers including affected relatives in AD-EOAD families was n = 81 patients (Tables 1–4). Overall, 12 PSEN1 mutations and 1 PSEN2 mutation were previously unreported (Tables 1 and 2, in bold). In the next sections, we describe the mutation spectrum, with a particular focus on novel mutations.
Tab. 1. Previously unreported French families with AD-EOAD and sporadic cases carrying a PSEN1 mutation. 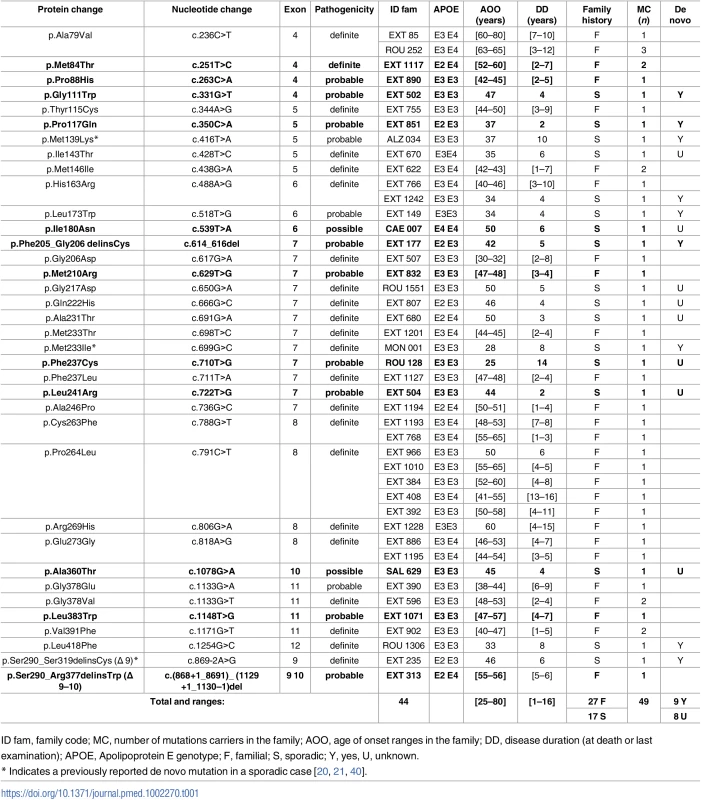
Novel mutations appear in bold. Tab. 2. Previously unreported French families with AD-EOAD carrying a PSEN2 mutation. 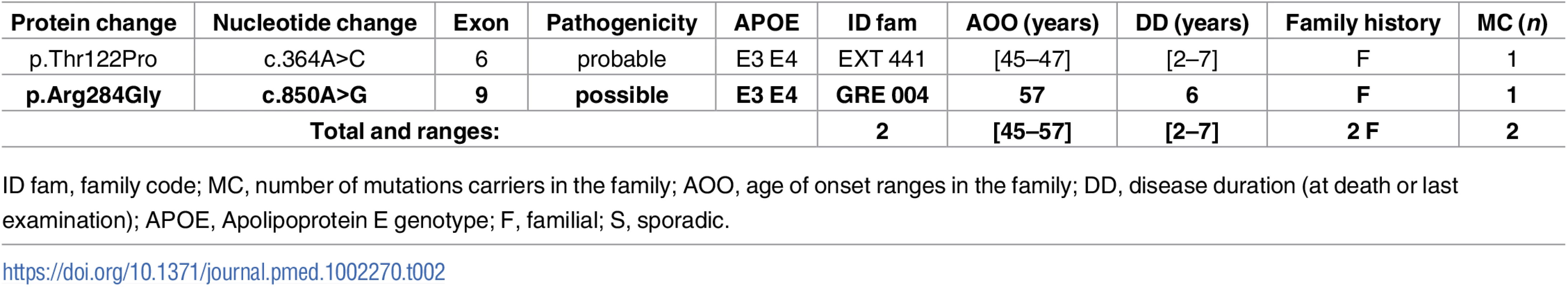
Novel mutations appear in bold. Tab. 3. Previously unreported French families with AD-EOAD carrying an APP mutation. 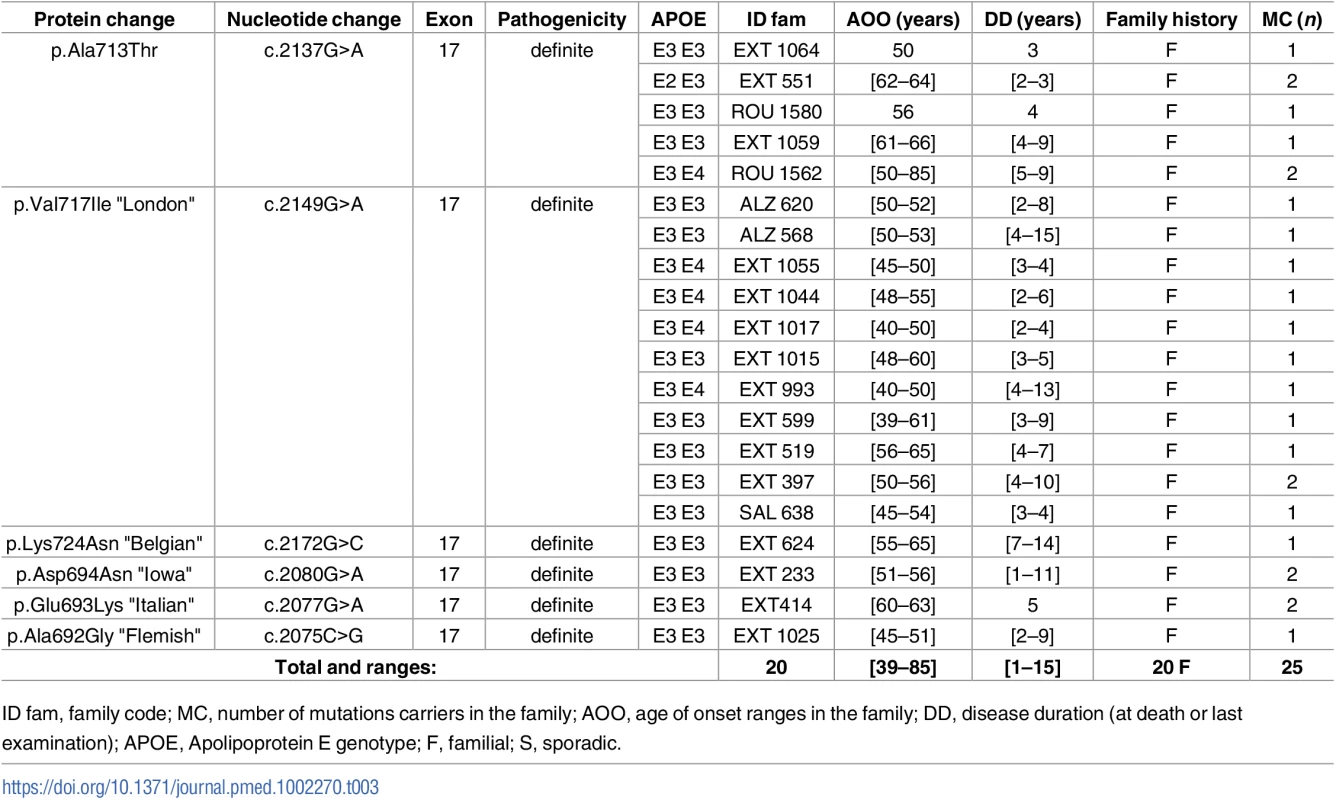
ID fam, family code; MC, number of mutations carriers in the family; AOO, age of onset ranges in the family; DD, disease duration (at death or last examination); APOE, Apolipoprotein E genotype; F, familial; S, sporadic. Tab. 4. Previously unreported French families with AD-EOAD and sporadic cases carrying an APP duplication. 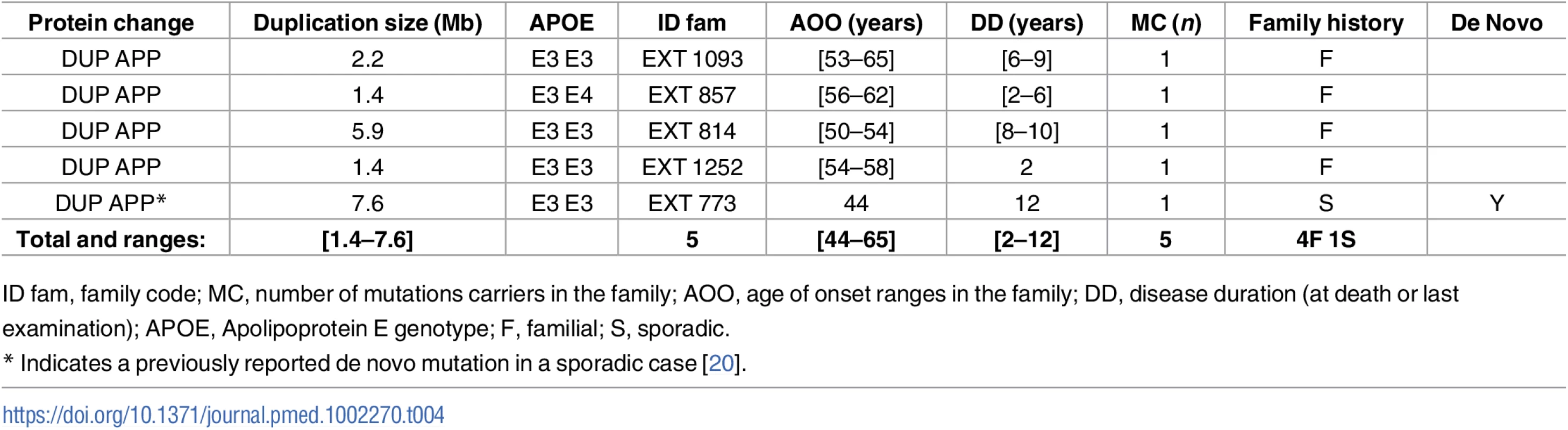
ID fam, family code; MC, number of mutations carriers in the family; AOO, age of onset ranges in the family; DD, disease duration (at death or last examination); APOE, Apolipoprotein E genotype; F, familial; S, sporadic. PSEN1
Five of the 12 novel PSEN1 mutations were identified in AD-EOAD families: a sister and the mother of the patient carrying the c.251T>C, p.(Met84Thr) mutation were also affected with AD (age at death: 61 and 64 y, respectively); the father of the patient carrying the c.263C>A, p.(Pro88His) mutation died at age 47 with an AD diagnosis; the father of the patient carrying the c.629T>G, p.(Met210Arg) mutation died from AD at age 50, with an AOO of 47 y; the mother and the maternal grandmother of the patient carrying the c.1148T>G, p.(Leu383Trp) mutation died from AD at 54 and 50 y, respectively (AOO was 47 y for both). We also detected in an AD-EOAD family a novel genomic in-frame deletion encompassing PSEN1 exons 9 and 10: c.(868+1_869–1)_(1129+1_1130–1)del, p.Ser290_Arg1129delinsTrp, thereafter named Δ9–10, which resulted in a missense change from serine to tryptophan at the aberrant exon 8–11 junction (Table 1). The remaining 7 novel PSEN1 mutations were found in patients with sporadic EOAD. Among these mutations, a censoring effect was observed in families of patients carrying the c.772T>C, p.(Leu241Arg), the c.539T>A, p.(Ile180Asn), and the c.710T>G, p.(Phe237Cys) substitutions, while the c.331G>T, p.(Gly111Trp), the c.350C>A, p.(Pro117Gln), and the c.614_616del, p.(Phe205_Gly206delinsCys) mutations occurred de novo. The seventh patient carried the c.1078G>A p.(Ala360Thr) variant. No censoring effect was noted in his family, but parental DNA was not available to verify the de novo occurrence of the mutation (Table 1). Among carriers of the PSEN1 mutation, the clinical presentation was mainly isolated progressive cognitive decline, but six patients carrying either the p.(Pro264Leu), p.(Leu173Trp), p.(Gln222His), or the Δ9–10 PSEN1 mutation displayed an associated phenotype of spastic paraparesis. Another patient carrying the PSEN1 p.(Gly378Glu) substitution also exhibited an atypical presentation: cerebellar ataxia and extra pyramidal syndrome.
PSEN2
Only one novel PSEN2 mutation, c.850A>G, p.(Arg284Gly), and a previously known mutation, p.(Thr122Pro), were identified during this screen (Table 2). No atypical phenotype was noticed.
APP
In the APP gene, no novel mutation was found. We identified a previously reported mutation in 25 patients from 20 AD-EOAD families (Table 3). The most frequent one was the c.2149G>A, p.(Val717Ile) substitution, which was present in 12 subjects from 11 families. Clinical features were typical of AD with amnestic presentation. The c.2137G>A, p.(Ala713Thr) mutation was found in 7 patients from 5 unrelated families. They exhibited a progressive cognitive decline starting from age 50 to 66 y. Notably, the mother of a patient who carried the mutation together with an APOE 4–4 genotype had no cognitive impairment until the age of 85, when she presented recurrent lobar hematoma. In addition, 5 subjects from 3 families carried mutations located within the coding sequence of the Aβ peptide: one carried the “Flemish” APP mutation c.2075C>G, p.(Ala692Gly), two carried the “Italian” mutation c.2077G>A, p.(Glu693Lys), and another two carried the “Iowa” mutation c.2080G>A, p.(Asp694Asn). A complete description of the phenotype of these 5 patients is provided in Sellal et al. [19].
APP duplications
Four subjects in four distinct AD-EOAD families and a sporadic case carried an APP duplication (Table 4). All patients exhibited progressive cognitive impairment. Only one presented signs of CAA and suffered from intracerebral hematoma at the age of 60.
CSF biomarkers
CSF biomarkers were available for 53 out of 81 mutation carriers (65%) (Table 5). There was no significant difference in Aβ42, Tau, and P-Tau mean values between patients bearing PSEN1 and APP mutations or duplications (two groups, p-values = 0.78, 0.19, and 0.16, respectively, Mann–Whitney U test). Among the 53 patients, 46 (87%) were classified PLM 3, 5 (9%) were classified PLM 2, and 2 (4%) were classified PLM 1; no patient was classified PLM 0. Among the 5 patients classified PLM 2, 2 had low Aβ42 and elevated Tau levels, and 3 had elevated Tau and P-Tau with normal Aβ42 CSF level. Two of the latter 3 patients carried a PSEN1 mutation: 1 carried the p.(Leu383Trp) with AOO at 57 y and 4 y of evolution, and the other carried the p.(Ala231Thr) with AOO at 50 y and 3 y of evolution. The third one carried an APP p.(Val717Ile) mutation with an AOO at 56 y and 4 y of evolution. The two patients classified PLM 1 had low Aβ42 value, without Tau or P-Tau elevation. One carried a p.(Ala360Thr) PSEN1 mutation with AOO at 45 y and 3 y of evolution; the second carried a p.(Ala692Gly) APP mutation with AOO at 45 y and 2 y of evolution.
Tab. 5. CSF biomarkers levels in mutation carriers (pg/mL). 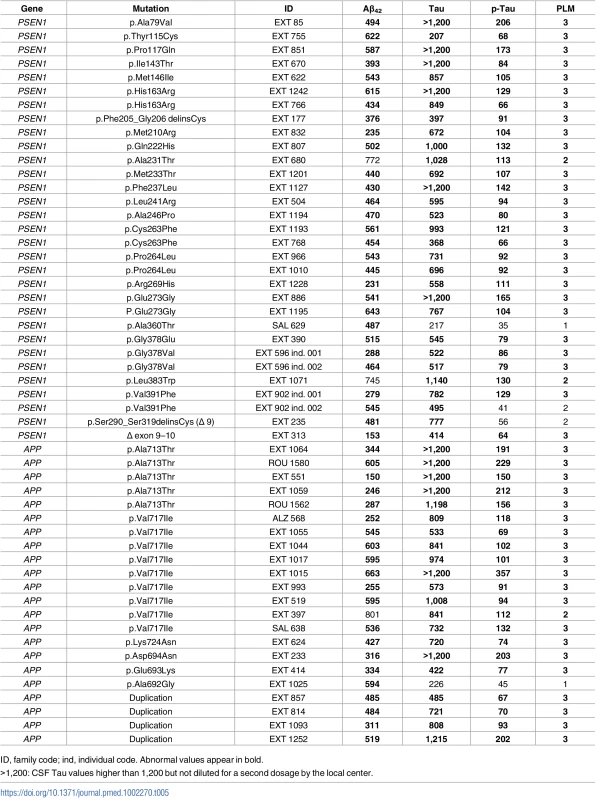
ID, family code; ind, individual code. Abnormal values appear in bold. Neuropathology
Neuropathological examination was available for three subjects. For patient EXT 773, who carried an APP duplication, the diagnosis was definite AD with Braak stage VI, Thal stage V. There was amyloid deposition in vessel walls in the insula and basal ganglia. Signs of severe CAA were found in middle frontal gyrus, superior temporal gyrus, inferior parietal cortex, and primary motor area. Lewy bodies were found in the amygdala, locus niger, nucleus basalis of Meynert, and entorhinal cortex.
For patient EXT 149, who carried the c.518T>G, p.Leu173Trp de novo PSEN1 mutation, rare senile plaques associated with numerous cotton wool deposits and neurofibrillary tangles were present in hippocampal regions and cortical areas. Lewy bodies were found in the amygdala and limbic cortex as well as the frontal, temporal, and parietal cortices and cingulum. CAA was noted in hippocampal regions, the temporal lobe, and the cerebellum.
For patient EXT 1117, who carried the c.251T>C, p.Met84Thr PSEN1 mutation, neuropathological examination showed global atrophy, particularly in temporal lobes. Samples from the cerebellum and the frontal, temporal, and parietal cortices showed numerous senile plaques and neurofibrillary tangles associated with severe CAA. No Lewy bodies were observed.
Mutational spectrum in the whole French EOAD series
Adding this sample to our previous reports [1,8,9,14,15,19,21–24], a total of 170 AD-EOAD families and 18 sporadic cases carrying mutations in genes known to cause EOAD have now been identified by our national reference center. Ninety distinct mutations (78 PSEN1, 4 PSEN2, and 8 APP, including APP duplication) were represented by respectively 127, 9, 34, and 18 occurrences in this whole sample (S1 Table). For each distinct mutation, the frequency reported in the Exome Aggregation Consortium (ExAC) database [25], which colligates human exome data from ~60,000 individuals, is null or very low (S1 Table).
The mean AOO for PSEN1 mutation carriers was 44.4 y (range 24–80), 53.9 y (range 45–69) for PSEN2 mutation carriers, 50.9 y (range 39–85) for APP mutation carriers, and 51.1 y (range 41–69) for patients carrying APP duplications. Variation of AOO by mutated gene was similar to the one reported by Ryman et al. (2014) [26].
Sporadic cases and de novo mutations
Among the 129 patients with a sporadic presentation and an AOO before 51 y for whom a mutation screening was performed, we identified 18 mutations, including 17 PSEN1 mutations and 1 APP duplication (Tables 1 and 4). For 10 patients, DNA of the unaffected parents was available, and analysis of parental DNA showed that the 10 mutations had occurred de novo: 7 patients carried a de novo PSEN1 missense mutation, 1 carried a de novo splicing PSEN1 mutation, 1 carried a de novo PSEN1 indel, and another 1 carried an APP de novo duplication. Interestingly, 5 out of 7 missense de novo PSEN1 mutations occurred at a position already known to be hit by pathogenic mutations. Parental DNA was not available for the remaining PSEN1 mutation carriers, but we noted a strong censoring effect due to a young age at death in two families, and the parents were unknown in three other families. For the remaining 3 patients, the absence of both a censoring effect and AD history in the parents is suggestive of a de novo occurrence, but this could not be proved by parental DNA analysis.
Discussion
We have studied two samples of EOAD patients and identified 10 novel missense mutations, 1 novel indel, and 1 novel genomic deletion in PSEN1 and 1 novel missense mutation in PSEN2. According to the Guerreiro’s algorithm [12], pathogenicity was considered as definite for 1 mutation, probable for 9, and possible for 3. Considering the whole French EOAD series, 90 distinct mutations (including the APP duplication) are now reported, and pathogenicity is considered definite for 69 mutations (77%), probable for 16 (18%), and possible for 3 (5%).
The pathological effect of three known mutations deserves discussion because of incomplete penetrance, nonpathogenicity, or wide range of AOO.
The PSEN1 c.236C>T, p.(Ala79Val) substitution is currently considered pathogenic and leads to an increase in Aβ42 level and Aβ42/Aβ40 ratios in cell cultures [27]. However, this variant seems to be associated with a later onset compared to the other PSEN1 variants. It was found in several families with late-onset AD (LOAD) [26,28,29]. Four mutation carriers from one family had a definite, neuropathological diagnosis of AD and an AOO after 75 y [28]. Of note, this variant has been reported once in the ExAC database [25] (among ~60,000 controls). Considering that it was also found in subjects with EOAD [30,31], these data suggest that this mutation is associated with a large range of AOO (53–78 y), which could lead to underestimation of its frequency and is of importance for genetic counseling.
Second, the PSEN2 c.211T>C, p.(Arg71Trp) variant was initially found in patients with LOAD [12,32,33]. We previously reported this variant in two EOAD families [14], but we removed it from our complete list because it is now considered as nonpathogenic. It did not segregate with AD in several families [32], including 8/14 affected individuals not carrying this variant in one large family [29]. It was found with an allele frequency of 0.034% (1.95% in the Finnish population) in the ExAC database. When coexpressed in HEK293 cells with APP, the variant did not alter the Aβ42/Aβ40 ratio in vitro [34]. As previously discussed, these elements lead us to consider this mutation as nonpathogenic [15].
Third, since our first report in a patient with sporadic probable AD [8], the APP c.2137 G>A p.(Ala713Thr) mutation has now been found in 24 patients from 11 families [5,35–39], including the 6 patients from 5 families included here. Although cerebrovascular lesions were described in brain imaging of some of these patients [36–38], the clinical presentation was a progressive cognitive decline in all but one of the reported cases. Interestingly, AOO ranged from 49 to 85 y, and several asymptomatic carriers were also reported, including one 88-y-old woman [8]. In one family, the mutation was found homozygous in 3 patients [38], and the disease onset was not different from the heterozygous carriers. In the present report, the mother of the proband ROU-1562 had no cognitive impairment until the age of 85, when a diagnosis of probable CAA was made. Of note, this variant has been reported with an allele frequency of 0.0058% in the ExAC database. Taken together, this suggests that the p.Ala713Thr substitution is a pathological variant with reduced penetrance, which is unusual compared to other APP mutations and is of main consequence for genetic counseling.
A notable finding of this study, as compared with the state of the literature, is the number (n = 10) of de novo PSEN1 or APP mutations detected in this set of 129 sporadic cases with onset below age 51. Furthermore, this could be underestimated, as parental DNA was not available for all cases. To our knowledge, only four de novo mutations had previously been reported in APP or PSEN1, including three by our group [20,22,23,40]. To our knowledge, there is no evidence to suggest that the PSEN1 gene is a hot spot of de novo mutations. Following the estimations by Samocha and coauthors [41] provided on the ExAC database [25], the probability to observe a PSEN1 de novo missense mutation in an individual is 1.29 x 10−5. This probability is that of an average gene since 56% of genes are more mutable and 44% less mutable than PSEN1. Thus, the discrepancy between the low number of previously reported de novo PSEN1 mutations in sporadic EOAD patients and the present report is likely to reflect a lack of inclusion of these patients in previous mutational screenings, which focused on familial cases. This underscores the need to systematically include patients with sporadic presentation and very early AOO in genetic screening. Consequences for genetic counseling are important, as the offspring of a mutation carrier has a same 50% risk to be a mutation carrier regardless of the familial or sporadic presentation of the affected parent; the offspring can then (i) be accurately informed, (ii) ask for a presymptomatic testing, and (iii) be a possible candidate for preventive clinical trials [13].
Concerning CSF biomarkers, 48/53 (91%) of patients with available CSF exhibit signs of both Aβ and Tau pathology, and 87% of the mutation carriers were classified PLM 3. This is higher than the 76% reported in our previous series [14]. This difference can be explained by the change in the Aβ42 cutoff (<700 versus <500 pg/ml in our previous series) according to the 2013 recommendations of the PLM network, whose aim is to homogenize preanalytical treatment for CSF biomarkers across French centers [42]. Overall, no AD mutation carrier presented with normal CSF biomarkers, suggesting that when all three CSF biomarkers are in normal ranges, genes involved in other neurodegenerative diseases should be screened in the first instance.
Our primary goal was to provide to clinicians a list of variants that can accurately be used in genetic counseling. Considering our whole series, this goal is achieved for 60/78 (77%) of PSEN1, 1/4 of PSEN2, and 8/8 of APP mutations reported in the French population. However, despite a large effort, too many mutations in AD-EOAD genes remain insufficiently characterized, and some are incompletely penetrant. The recent analysis of ~60,000 human exomes by the ExAC consortium has revealed an implausibly high per-individual burden of variants reported as causing disease in databases listing Mendelian disease alleles. These findings cast doubt on the validity of these databases and lead to a reclassification of numerous variants as benign [25]. In this context, it is reassuring to see that all variants reported here have a null or very low frequency in ExAC, which is a strong argument for pathogenicity.
A limitation of this study is the absence of functional assessment of the possibly and probably pathogenic variants, which should help their classification. Moreover, only three genes were analyzed. It is possible that de novo mutations in other genes are also involved in the genetic determinism of sporadic forms. To address this latter issue, the next step is now to perform exome sequencing on negatively screened families and sporadic cases. Indeed, this approach already enabled us to show that (i) rare variations in the SORL1 gene might be responsible of a subset of AD-EOAD families [43] or at least constitute a penetrant risk factor for familial EOAD [44] and (ii) a set of genes defining an Aβ-centered genetic network are enriched in de novo mutations in sporadic cases [20].
Our findings suggest that a nonnegligible fraction of PSEN1 mutations occur de novo. The practical implication for clinicians is to highlight the need to systematically include patients with sporadic presentation and very early AOO in genetic screening for the APP, PSEN1, and PSEN2 genes. In addition, the need to pursue the effort to classify variants should be emphasized since, based on our results, definite pathogenicity is currently established for only 77% of identified mutations in these genes.
Supporting Information
Zdroje
1. Campion D, Dumanchin C, Hannequin D, Dubois B, Belliard S, Puel M, et al. Early-onset autosomal dominant Alzheimer disease: prevalence, genetic heterogeneity, and mutation spectrum. American journal of human genetics. 1999;65(3):664–70. doi: 10.1086/302553 10441572
2. Sherrington R, Rogaev EI, Liang Y, Rogaeva EA, Levesque G, Ikeda M, et al. Cloning of a gene bearing missense mutations in early-onset familial Alzheimer's disease. Nature. 1995;375(6534):754–60. doi: 10.1038/375754a0 7596406
3. Levy-Lahad E, Wijsman EM, Nemens E, Anderson L, Goddard KA, Weber JL, et al. A familial Alzheimer's disease locus on chromosome 1. Science. 1995;269(5226):970–3. 7638621
4. Rogaev EI, Sherrington R, Rogaeva EA, Levesque G, Ikeda M, Liang Y, et al. Familial Alzheimer's disease in kindreds with missense mutations in a gene on chromosome 1 related to the Alzheimer's disease type 3 gene. Nature. 1995;376(6543):775–8. doi: 10.1038/376775a0 7651536
5. Sherrington R, Froelich S, Sorbi S, Campion D, Chi H, Rogaeva EA, et al. Alzheimer's disease associated with mutations in presenilin 2 is rare and variably penetrant. Human molecular genetics. 1996;5(7):985–8. 8817335
6. Chartier-Harlin MC, Crawford F, Houlden H, Warren A, Hughes D, Fidani L, et al. Early-onset Alzheimer's disease caused by mutations at codon 717 of the beta-amyloid precursor protein gene. Nature. 1991;353(6347):844–6. doi: 10.1038/353844a0 1944558
7. Goate A, Chartier-Harlin MC, Mullan M, Brown J, Crawford F, Fidani L, et al. Segregation of a missense mutation in the amyloid precursor protein gene with familial Alzheimer's disease. Nature. 1991;349(6311):704–6. doi: 10.1038/349704a0 1671712
8. Carter DA, Desmarais E, Bellis M, Campion D, Clerget-Darpoux F, Brice A, et al. More missense in amyloid gene. Nature genetics. 1992;2(4):255–6. doi: 10.1038/ng1292-255 1303275
9. Rovelet-Lecrux A, Hannequin D, Raux G, Le Meur N, Laquerriere A, Vital A, et al. APP locus duplication causes autosomal dominant early-onset Alzheimer disease with cerebral amyloid angiopathy. Nature genetics. 2006;38(1):24–6. doi: 10.1038/ng1718 16369530
10. Haass C, Kaether C, Thinakaran G, Sisodia S. Trafficking and proteolytic processing of APP. Cold Spring Harbor perspectives in medicine. 2012;2(5):a006270. doi: 10.1101/cshperspect.a006270 22553493
11. Campion D, Pottier C, Nicolas G, Le Guennec K, Rovelet-Lecrux A. Alzheimer disease: modeling an Abeta-centered biological network. Molecular psychiatry. 2016;21(7):861–71. doi: 10.1038/mp.2016.38 27021818
12. Guerreiro RJ, Baquero M, Blesa R, Boada M, Bras JM, Bullido MJ, et al. Genetic screening of Alzheimer's disease genes in Iberian and African samples yields novel mutations in presenilins and APP. Neurobiology of aging. 2010;31(5):725–31. doi: 10.1016/j.neurobiolaging.2008.06.012 18667258
13. Moulder KL, Snider BJ, Mills SL, Buckles VD, Santacruz AM, Bateman RJ, et al. Dominantly Inherited Alzheimer Network: facilitating research and clinical trials. Alzheimer's research & therapy. 2013;5(5):48.
14. Wallon D, Rousseau S, Rovelet-Lecrux A, Quillard-Muraine M, Guyant-Marechal L, Martinaud O, et al. The French series of autosomal dominant early onset Alzheimer's disease cases: mutation spectrum and cerebrospinal fluid biomarkers. Journal of Alzheimer's disease: JAD. 2012;30(4):847–56. doi: 10.3233/JAD-2012-120172 22475797
15. Nicolas G, Wallon D, Charbonnier C, Quenez O, Rousseau S, Richard AC, et al. Screening of dementia genes by whole-exome sequencing in early-onset Alzheimer disease: input and lessons. European journal of human genetics: EJHG. 2016;24(5):710–6. doi: 10.1038/ejhg.2015.173 26242991
16. Acuna-Hidalgo R, Veltman JA, Hoischen A. New insights into the generation and role of de novo mutations in health and disease. Genome biology. 2016;17(1):241. doi: 10.1186/s13059-016-1110-1 27894357
17. McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR Jr., Kawas CH, et al. The diagnosis of dementia due to Alzheimer's disease: recommendations from the National Institute on Aging-Alzheimer's Association workgroups on diagnostic guidelines for Alzheimer's disease. Alzheimer's & dementia: the journal of the Alzheimer's Association. 2011;7(3):263–9.
18. Lehmann S, Dumurgier J, Schraen S, Wallon D, Blanc F, Magnin E, et al. A diagnostic scale for Alzheimer's disease based on cerebrospinal fluid biomarker profiles. Alzheimer's research & therapy. 2014;6(3):38.
19. Sellal F, Wallon D, Martinez-Almoyna L, Marelli C, Dhar A, Oesterle H, et al. APP Mutations in Cerebral Amyloid Angiopathy with or without Cortical Calcifications: Report of Three Families and a Literature Review. Journal of Alzheimer's disease: JAD. 2017;56(1):37–46. doi: 10.3233/JAD-160709 27858710
20. Rovelet-Lecrux A, Charbonnier C, Wallon D, Nicolas G, Seaman MN, Pottier C, et al. De novo deleterious genetic variations target a biological network centered on Abeta peptide in early-onset Alzheimer disease. Molecular psychiatry. 2015;20(9):1046–56. doi: 10.1038/mp.2015.100 26194182
21. Raux G, Guyant-Marechal L, Martin C, Bou J, Penet C, Brice A, et al. Molecular diagnosis of autosomal dominant early onset Alzheimer's disease: an update. Journal of medical genetics. 2005;42(10):793–5. doi: 10.1136/jmg.2005.033456 16033913
22. Portet F, Dauvilliers Y, Campion D, Raux G, Hauw JJ, Lyon-Caen O, et al. Very early onset AD with a de novo mutation in the presenilin 1 gene (Met 233 Leu). Neurology. 2003;61(8):1136–7. 14581682
23. Dumanchin C, Brice A, Campion D, Hannequin D, Martin C, Moreau V, et al. De novo presenilin 1 mutations are rare in clinically sporadic, early onset Alzheimer's disease cases. French Alzheimer's Disease Study Group. Journal of medical genetics. 1998;35(8):672–3. 9719376
24. Dumanchin C, Tournier I, Martin C, Didic M, Belliard S, Carlander B, et al. Biological effects of four PSEN1 gene mutations causing Alzheimer disease with spastic paraparesis and cotton wool plaques. Human mutation. 2006;27(10):1063.
25. Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536(7616):285–91. doi: 10.1038/nature19057 27535533
26. Ryman DC, Acosta-Baena N, Aisen PS, Bird T, Danek A, Fox NC, et al. Symptom onset in autosomal dominant Alzheimer disease: a systematic review and meta-analysis. Neurology. 2014;83(3):253–60. doi: 10.1212/WNL.0000000000000596 24928124
27. Kumar-Singh S, Theuns J, Van Broeck B, Pirici D, Vennekens K, Corsmit E, et al. Mean age-of-onset of familial alzheimer disease caused by presenilin mutations correlates with both increased Abeta42 and decreased Abeta40. Human mutation. 2006;27(7):686–95. doi: 10.1002/humu.20336 16752394
28. Kauwe JS, Jacquart S, Chakraverty S, Wang J, Mayo K, Fagan AM, et al. Extreme cerebrospinal fluid amyloid beta levels identify family with late-onset Alzheimer's disease presenilin 1 mutation. Annals of neurology. 2007;61(5):446–53. doi: 10.1002/ana.21099 17366635
29. Cruchaga C, Haller G, Chakraverty S, Mayo K, Vallania FL, Mitra RD, et al. Rare variants in APP, PSEN1 and PSEN2 increase risk for AD in late-onset Alzheimer's disease families. PLoS ONE. 2012;7(2):e31039. doi: 10.1371/journal.pone.0031039 22312439
30. Cruts M, van Duijn CM, Backhovens H, Van den Broeck M, Wehnert A, Serneels S, et al. Estimation of the genetic contribution of presenilin-1 and -2 mutations in a population-based study of presenile Alzheimer disease. Human molecular genetics. 1998;7(1):43–51. 9384602
31. Finckh U, Muller-Thomsen T, Mann U, Eggers C, Marksteiner J, Meins W, et al. High prevalence of pathogenic mutations in patients with early-onset dementia detected by sequence analyses of four different genes. American journal of human genetics. 2000;66(1):110–7. doi: 10.1086/302702 10631141
32. Sleegers K, Roks G, Theuns J, Aulchenko YS, Rademakers R, Cruts M, et al. Familial clustering and genetic risk for dementia in a genetically isolated Dutch population. Brain: a journal of neurology. 2004;127(Pt 7):1641–9.
33. Brouwers N, Sleegers K, Van Broeckhoven C. Molecular genetics of Alzheimer's disease: an update. Annals of medicine. 2008;40(8):562–83. doi: 10.1080/07853890802186905 18608129
34. To MD, Gokgoz N, Doyle TG, Donoviel DB, Knight JA, Hyslop PS, et al. Functional characterization of novel presenilin-2 variants identified in human breast cancers. Oncogene. 2006;25(25):3557–64. doi: 10.1038/sj.onc.1209397 16474849
35. Armstrong J, Boada M, Rey MJ, Vidal N, Ferrer I. Familial Alzheimer disease associated with A713T mutation in APP. Neuroscience letters. 2004;370(2–3):241–3. doi: 10.1016/j.neulet.2004.08.026 15488330
36. Rossi G, Giaccone G, Maletta R, Morbin M, Capobianco R, Mangieri M, et al. A family with Alzheimer disease and strokes associated with A713T mutation of the APP gene. Neurology. 2004;63(5):910–2. 15365148
37. Bernardi L, Geracitano S, Colao R, Puccio G, Gallo M, Anfossi M, et al. AbetaPP A713T mutation in late onset Alzheimer's disease with cerebrovascular lesions. Journal of Alzheimer's disease: JAD. 2009;17(2):383–9. doi: 10.3233/JAD-2009-1061 19363265
38. Conidi ME, Bernardi L, Puccio G, Smirne N, Muraca MG, Curcio SA, et al. Homozygous carriers of APP A713T mutation in an autosomal dominant Alzheimer disease family. Neurology. 2015;84(22):2266–73. doi: 10.1212/WNL.0000000000001648 25948718
39. Barber IS, Garcia-Cardenas JM, Sakdapanichkul C, Deacon C, Zapata Erazo G, Guerreiro R, et al. Screening exons 16 and 17 of the amyloid precursor protein gene in sporadic early-onset Alzheimer's disease. Neurobiology of aging. 2016;39 : 220 e1–7.
40. Golan MP, Styczynska M, Jozwiak K, Walecki J, Maruszak A, Pniewski J, et al. Early-onset Alzheimer's disease with a de novo mutation in the presenilin 1 gene. Experimental neurology. 2007;208(2):264–8. doi: 10.1016/j.expneurol.2007.08.016 17931627
41. Samocha KE, Robinson EB, Sanders SJ, Stevens C, Sabo A, McGrath LM, et al. A framework for the interpretation of de novo mutation in human disease. Nature genetics. 2014;46(9):944–50. doi: 10.1038/ng.3050 25086666
42. Gabelle A, Dumurgier J, Vercruysse O, Paquet C, Bombois S, Laplanche JL, et al. Impact of the 2008–2012 French Alzheimer Plan on the use of cerebrospinal fluid biomarkers in research memory center: the PLM Study. Journal of Alzheimer's disease: JAD. 2013;34(1):297–305. doi: 10.3233/JAD-121549 23186986
43. Pottier C, Hannequin D, Coutant S, Rovelet-Lecrux A, Wallon D, Rousseau S, et al. High frequency of potentially pathogenic SORL1 mutations in autosomal dominant early-onset Alzheimer disease. Molecular psychiatry. 2012;17(9):875–9. doi: 10.1038/mp.2012.15 22472873
44. Nicolas G, Charbonnier C, Wallon D, Quenez O, Bellenguez C, Grenier-Boley B, et al. SORL1 rare variants: a major risk factor for familial early-onset Alzheimer's disease. Molecular psychiatry. 2016;21(6):831–6. doi: 10.1038/mp.2015.121 26303663
Štítky
Interní lékařství
Článek vyšel v časopisePLOS Medicine
Nejčtenější tento týden
2017 Číslo 3- Berberin: přírodní hypolipidemikum se slibnými výsledky
- Léčba bolesti u seniorů
- Příznivý vliv Armolipidu Plus na hladinu cholesterolu a zánětlivé parametry u pacientů s chronickým subklinickým zánětem
- Červená fermentovaná rýže účinně snižuje hladinu LDL cholesterolu jako vhodná alternativa ke statinové terapii
- Jak postupovat při výběru betablokátoru − doporučení z kardiologické praxe
-
Všechny články tohoto čísla
- 2016 Reviewer and Editorial Board Thank You
- , , and mutations in early-onset Alzheimer disease: A genetic screening study of familial and sporadic cases
- Differential associations of plasma lipids with incident dementia and dementia subtypes in the 3C Study: A longitudinal, population-based prospective cohort study
- Mixed pathologies and neural reserve: Implications of complexity for Alzheimer disease drug discovery
- -related risk of mild cognitive impairment and dementia for prevention trials: An analysis of four cohorts
- Neuropathological diagnoses and clinical correlates in older adults in Brazil: A cross-sectional study
- Early diagnosis of mild cognitive impairment and mild dementia through basic and instrumental activities of daily living: Development of a new evaluation tool
- Potentially modifiable lifestyle factors, cognitive reserve, and cognitive function in later life: A cross-sectional study
- Association between fatty acid metabolism in the brain and Alzheimer disease neuropathology and cognitive performance: A nontargeted metabolomic study
- Fine-mapping of the human leukocyte antigen locus as a risk factor for Alzheimer disease: A case–control study
- What’s the “Take Home” from Research on Dementia Trends?
- Cultural representations of dementia
- Dementia and aging populations—A global priority for contextualized research and health policy
- Dementia in the oldest old: Beyond Alzheimer disease
- Rehabilitation for people living with dementia: A practical framework of positive support
- Dementia in low-income and middle-income countries: Different realities mandate tailored solutions
- Challenges and opportunities in understanding dementia and delirium in the acute hospital
- Dementia incidence trend over 1992-2014 in the Netherlands: Analysis of primary care data
- Association between delirium superimposed on dementia and mortality in hospitalized older adults: A prospective cohort study
- Development of an adaptive, personalized, and scalable dementia care program: Early findings from the Care Ecosystem
- Genetic assessment of age-associated Alzheimer disease risk: Development and validation of a polygenic hazard score
- Age-related cognitive decline and associations with sex, education and apolipoprotein E genotype across ethnocultural groups and geographic regions: a collaborative cohort study
- The impact of individual Cognitive Stimulation Therapy (iCST) on cognition, quality of life, caregiver health, and family relationships in dementia: A randomised controlled trial
- Effectiveness of an intervention to facilitate prompt referral to memory clinics in the United Kingdom: Cluster randomised controlled trial
- Subjective and objective cognitive function among older adults with a history of traumatic brain injury: A population-based cohort study
- Association of lifelong exposure to cognitive reserve-enhancing factors with dementia risk: A community-based cohort study
- Multimorbidity and healthcare utilization among home care clients with dementia in Ontario, Canada: A retrospective analysis of a population-based cohort
- PLOS Medicine
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Effectiveness of an intervention to facilitate prompt referral to memory clinics in the United Kingdom: Cluster randomised controlled trial
- , , and mutations in early-onset Alzheimer disease: A genetic screening study of familial and sporadic cases
- Challenges and opportunities in understanding dementia and delirium in the acute hospital
- Early diagnosis of mild cognitive impairment and mild dementia through basic and instrumental activities of daily living: Development of a new evaluation tool
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



