-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaNovel and Conserved Protein Macoilin Is Required for Diverse Neuronal Functions in
Neural signals are processed in nervous systems of animals responding to variable environmental stimuli. This study shows that a novel and highly conserved protein, macoilin (MACO-1), plays an essential role in diverse neural functions in Caenorhabditis elegans. maco-1 mutants showed abnormal behaviors, including defective locomotion, thermotaxis, and chemotaxis. Expression of human macoilin in the C. elegans nervous system weakly rescued the abnormal thermotactic phenotype of the maco-1 mutants, suggesting that macoilin is functionally conserved across species. Abnormal thermotaxis may have been caused by impaired locomotion of maco-1 mutants. However, calcium imaging of AFD thermosensory neurons and AIY postsynaptic interneurons of maco-1 mutants suggest that macoilin is required for appropriate responses of AFD and AIY neurons to thermal stimuli. Studies on localization of MACO-1 showed that C. elegans and human macoilins are localized mainly to the rough endoplasmic reticulum. Our results suggest that macoilin is required for various neural events, such as the regulation of neuronal activity.
Published in the journal: . PLoS Genet 7(5): e32767. doi:10.1371/journal.pgen.1001384
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1001384Summary
Neural signals are processed in nervous systems of animals responding to variable environmental stimuli. This study shows that a novel and highly conserved protein, macoilin (MACO-1), plays an essential role in diverse neural functions in Caenorhabditis elegans. maco-1 mutants showed abnormal behaviors, including defective locomotion, thermotaxis, and chemotaxis. Expression of human macoilin in the C. elegans nervous system weakly rescued the abnormal thermotactic phenotype of the maco-1 mutants, suggesting that macoilin is functionally conserved across species. Abnormal thermotaxis may have been caused by impaired locomotion of maco-1 mutants. However, calcium imaging of AFD thermosensory neurons and AIY postsynaptic interneurons of maco-1 mutants suggest that macoilin is required for appropriate responses of AFD and AIY neurons to thermal stimuli. Studies on localization of MACO-1 showed that C. elegans and human macoilins are localized mainly to the rough endoplasmic reticulum. Our results suggest that macoilin is required for various neural events, such as the regulation of neuronal activity.
Introduction
Genome projects have determined DNA sequences of various organisms and identified locations of predicted genes in genomes. Surprisingly, numbers of genes are relatively similar across evolutionally divergent animal species. For example, estimated gene numbers are 20000–25000 in human [1], 14000 in Drosophila melanogaster [2], and 19000 in Caenorhabditis elegans [3]. Of the estimated genes in C. elegans, more than 40 percent of the predicted protein products show significant matches to those from other animals [3]. Despite this species homology, functions of these genes are poorly understood. Therefore, further characterization of these proteins could lead to novel insights about important cellular processes, such as gene regulation, protein maturation, signal transduction, and intracellular transport.
Animals use the nervous system to detect stimuli in the surrounding environment. These stimuli are then assessed based on past experiences, and converted into adaptive behaviors. Many neurons are required to act in coordination to execute these processes. However, it is difficult to address individual neuronal function in relation to other neurons in neural circuits because animal brains generally consist of enormous numbers of neurons. The anatomically characterized nervous system of C. elegans is composed of only 302 neurons [4], making it an effective model for studying neuronal function in circuits. Despite their simple nervous systems, C. elegans can sense an array of environmental stimuli and produce appropriate behavioral outputs. For example, worms remember cultivation temperatures in association with food conditions [5]–[7]. This thermotaxis behavior can be divided into two behavioral aspects. First, after cultivation C. elegans with food at a certain temperature for several hours, they will migrate to this cultivation temperature along a temperature gradient [5]–[7]. Previous studies have suggested that this migratory behavior could be explained by the following two opposing strategies: migrations up thermal gradients when located at temperatures below the cultivation temperature (i.e., thermophilic drive); and migrations down thermal gradients when at a higher temperature (i.e., cryophilic drive) [5]–[7]. Work by several groups has been unable to confirm thermophilic drive in C. elegans [8]–[11]. However, recent theoretical and experimental studies have shown that the steepness of the thermal gradient drastically changes migration behavior [12], [13]. The second behavioral aspect of thermotaxis involves worms leaving isothermal tracks (IT) on a temperature gradient as they migrate to the cultivation temperature associated with food (Figure 1B and [5], [6]). Previous reports have suggested that the neuron-specific calcium sensor-1 (NCS-1) plays an important role in optimizing IT [14]. Specifically, reports showed that a knockout phenotype of ncs-1 animals could be rescued by introducing a wild-type NCS-1 into the AIY interneuron, which is a component of the thermotaxis neural circuit, and that NCS-1 overexpression enhanced the frequency of IT behavior [14].
Fig. 1. Phenotype of maco-1. 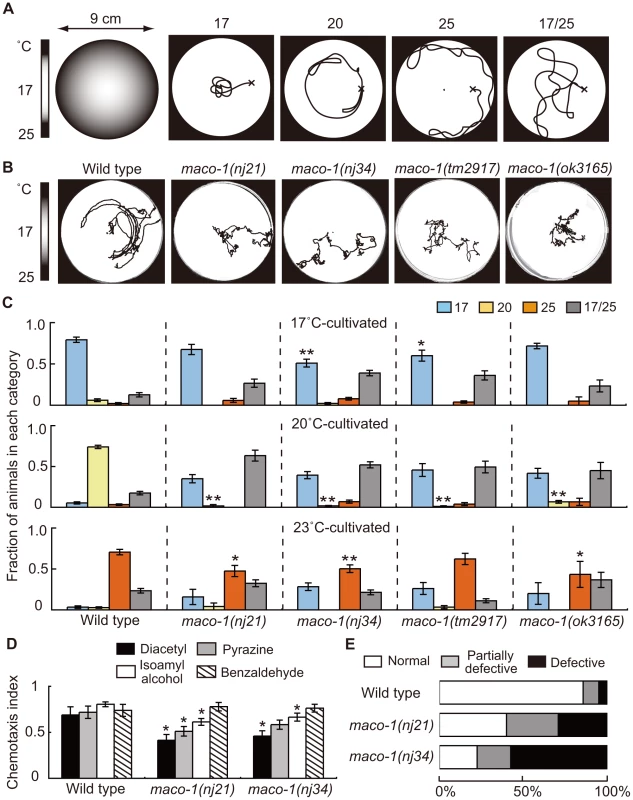
(A) Thermotaxis assay with a radial temperature gradient. (Left panel of A) The center of the assay plate (a 9 cm-diameter petri dish) is approximately 17°C and the periphery is approximately 25°C. Tracks of animals were categorized into four groups after one-hour of free movement. Animals that moved to the 17°C region (center) were classified as ‘17’. Animals that moved to the 20°C region or 25°C region (periphery) were classified as ‘20’ or ‘25’, respectively. Animals that moved randomly around both 17°C and 25°C regions were classified as ‘17/25’. (B) Tracks of wild-type and maco-1 mutant animals showing thermotaxis behaviors on a radial temperature gradient ranging from 17°C (center) to 25°C (periphery). Animals were grown at 20°C. (C) Thermotaxis of wild-type and maco-1 mutant animals cultivated at 17°C (upper graph), 20°C (middle graph) and 23°C (lower graph). n = 60–380. Single and double asterisks indicate p<0.05 and p<0.01 by ANOVA with a Dunnett's post hoc test, respectively. (D) Chemotaxis of wild-type and maco-1 mutant animals for odorants. Odorant concentrations are described in the Materials and Methods section. For each assay, 50–300 animals were analyzed. A single asterisk indicates p<0.05 compared to wild-type by ANOVA with a Dunnett's post hoc test. Bars represent the mean of three independent assays with error bars showing the standard error of the mean (SEM). (E) Chemotaxis to sodium chloride (NaCl). For each genotype, 40–120 animals were individually assayed. The phenotypic categories are described in the Materials and Methods section. Mean fractions categorized as ‘Normal’ were 0.86±0.05 among wild-type animals, 0.40±0.04 in maco-1(nj21) mutants and 0.23±0.18 in maco-1(nj34) mutants. Statistically significant differences were observed between the fraction of ‘Normal’ of wild-type and maco-1 mutants (p<0.01 by ANOVA with a Dunnett's post hoc test). The thermotaxis phenotype of C. elegans, which was achieved by laser-ablating specific neurons, has been examined previously, and results led to the hypothesis of a neural circuit for thermotaxis [6]. This neural circuit is presently one of the simplest circuits to regulate animal behavior. By capitalizing on this simple structure, we expect that the study of thermotaxis behavior will reveal molecular mechanisms of thermosensation and concepts of neural plasticity and neuronal function. Several molecular components related to thermotaxis behavior have been identified. The three guanylyl cyclases, GCY-8, GCY-18 and GCY-23, which are expressed in AFD thermosensory neurons, appear to redundantly produce cyclic guanosine monophosphate (cGMP) as a second messenger. The cGMP-dependent cation channel, which is composed of TAX-2 and TAX-4, is crucial in activating AFD neurons [15]–[17]. TAX-6 (calcineurin) and TTX-4 (PKC-1; nPKC-epsilon/eta), both of which act as negative regulators of AFD thermosensory signaling, have also been reported [18], [19]. Calcium imaging using the genetically encoded calcium indicator cameleon can display fluorescence changes from variable intracellular calcium concentrations [20]. This imaging has shown that AFD neurons respond to thermal stimuli in a cultivation temperature-dependent manner [21]. This response occurred even after disconnecting the AFD dendrites from their cell bodies [22], indicating that temperature memories are stored in AFD neurons. Consistent with findings in AFD neurons, recent calcium imaging analyses also suggest that AWC thermosensory neurons can also store temperature memories [23], [24].
In this study, we found that the novel protein MACO-1 is required for diverse neuronal functions in C. elegans. maco-1/ttx-8 mutants exhibited abnormalities in thermotaxis and chemotaxis to odorants and sodium chloride (NaCl). The maco-1 gene encodes a conserved novel protein that is expressed in many neurons. A previous study reported that the mouse ortholog of MACO-1, macoilin (C61), showed neuron-specific expression [25], although its function remained largely unknown. Rescue experiments revealed that simultaneous expression of MACO-1 in AFD, AIY and AIZ neurons of maco-1 mutants is required for thermotaxis behavior. We also found that maco-1 mutants showed abnormal locomotion resulting from frequent turns. Although defective locomotion of maco-1 mutants might have caused abnormal thermotaxis, calcium imaging indicated that MACO-1 is required for activation of AFD and AIY neurons. In vivo co-localization studies have shown that both MACO-1 and human macoilin are localized to the rough endoplasmic reticulum (rER). All together, our results suggest that macoilin is required for a variety of intracellular events that influence basic neuronal function across species.
Results
maco-1/ttx-8 mutants showed prominent thermotaxis defects and slight chemotaxis defects
To elucidate molecular mechanisms of thermotaxis, we performed a genetic screen and isolated several mutants that were thermotaxis defective [19]. Two of these mutants, maco-1(nj21) and maco-1(nj34), were isolated as novel mutants. Two other mutant alleles, maco-1(tm2917) and maco-1(ok3165), were isolated by the National Bioresource Project (Japan) and the C. elegans Gene Knockout Consortium, respectively. All four alleles were genetically recessive (data not shown). Despite obvious defects in thermotaxis after cultivation at 20°C (Figure 1B and 1C), maco-1(nj21) and maco-1(nj34) mutants were slightly defective in chemotaxis to odorants (Figure 1D and Figure S1) or to NaCl (Figure 1E). These results suggest that MACO-1 is necessary for animals to respond normally to various stimuli, including temperature, odorants and NaCl. Strong thermotaxis defects in maco-1 mutants after cultivation at 20°C suggest that MACO-1 may play an important role in thermotaxis, especially after cultivation at 20°C. Alternatively, the thermotaxis defects in maco-1 mutants could have been more easily detected in thermotaxis assays for 20°C-grown animals (20°C TTX assay) than 17°C or 23°C TTX assays. Specifically, during 20°C TTX assays, animals must migrate and remain in the 20°C region, which is shown as the light grey zone in the left schematic of Figure 1A. By contrast, during 17°C and 23°C TTX assays, animals can achieve thermotactic behaviors by migrating to 17°C and 23°C regions at the center and the periphery of a 9 cm plate, respectively. These areas are places where animals tend to remain due to the nature of assay plates and non-linear temperature gradients.
The maco-1 gene encodes a novel protein
The maco-1 gene was identified by mapping it with the snip-single nucleotide polymorphisms (SNPs) method [26] and conducting subsequent rescue experiments with genomic or polymerase chain reaction (PCR) fragments. Defective thermotaxis of maco-1 mutants was rescued by introduction of the cosmid C09C9. However, thermotaxis was not rescued by the cosmid C15E12 or C03F5 (Figure 2A), suggesting that maco-1 is located in the region between cosmids C15E12 and C03F5 within the cosmid C09C9. In addition, a PCR fragment containing D2092.4 gene (PCR1) did not rescue abnormal thermotaxis behavior of maco-1 mutants. However, another PCR fragment containing D2092.5 gene (PCR2) could rescue this behavior (Figure 2A), suggesting that D2092.5 is identical to the maco-1 gene. D2092.5/maco-1 gene encodes a novel protein that was predicted to have five transmembrane domains in the N-terminus and four coiled coil domains in the C-terminus as determined by the TMHMM Server v. 2.0 [27] and by COILS server [28], respectively (Figure 2B). Blast searches of the U.S. National Center for Biotechnology Inventory (NCBI) inventory identified maco-1 orthologs in several species, including the invertebrates Drosophila melanogaster, and vertebrates Xenopus laevis, Mus musculus and Homo sapiens (Figure 2B, 2C and Table S1). The nj21 mutant has a G-to-A substitution at the splice acceptor site of the 9th intron. The nj34 mutant has two nonsense mutations (W231stop and Q761stop) and a missense mutation (A805T) (Figure 2D). The tm2917 mutant has a 780-base pair (bp) deletion and 1-bp insertion just before the first coiled coil domain (Figure 2D). The ok3165 mutant has a 972-bp deletion and 1-bp insertion in the coiled coil region (Figure 2D).
Fig. 2. Genetic and molecular analysis of maco-1. 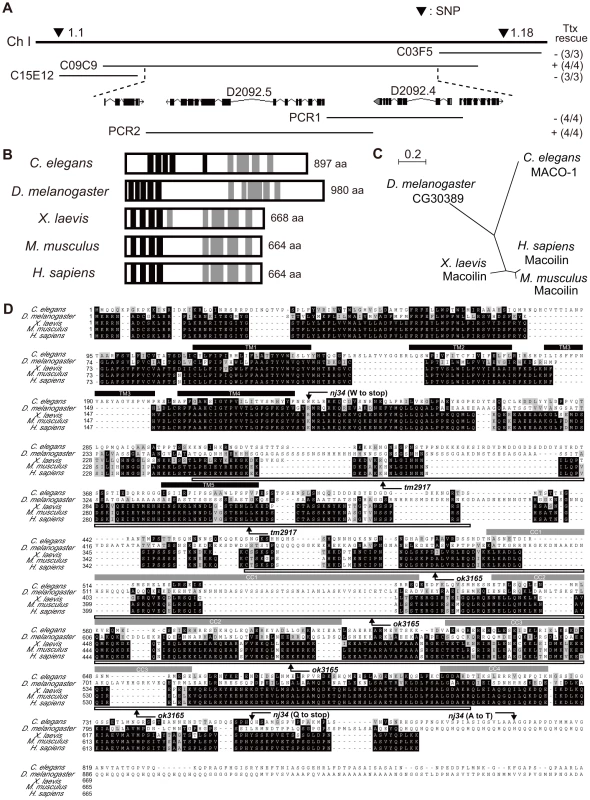
(A) Position of maco-1 on chromosome I. Arrowheads show locations of single nucleotide polymorphisms (SNPs). Results of rescue experiments among thermotaxis-defective maco-1 mutants are indicated as + (rescued) or − (not rescued) on the right side. Numbers in parentheses indicate the fraction of rescued lines. For each genotype, 120–180 animals were individually assayed for thermotaxis. The maco-1 gene was mapped to a 100–kb region between two SNPs (1.1 and 1.18) on chromosome I. We conducted rescue experiments using overlapping cosmids to encompass this 100-kb region. (B) Predicted structure of maco-1 gene family. Black and gray boxes indicate transmembrane domains and coiled coil domains, respectively. Numbers of amino acids are shown on the right. (C) Unrooted dendrogram of MACO-1. The dendrogram was generated using the freely available program, Clustal W [52] (http://clustalw.genome.jp/) and NJplot [53]. (D) Comparison of sequences of MACO-1 and orthologs of MACO-1 in other species and mutation sites of maco-1. Black and gray boxes highlight identical and similar residues, respectively. Black bars and gray bars above the sequences indicate predicted transmembrane domains (TM1–TM5) and coiled coil domains (CC1–CC4) of MACO-1 in C. elegans, respectively. Boxes below the amino-acid alignment correspond to mutation sites of tm2917 and ok3165. MACO-1 is mainly localized to the rER
To study subcellular localization of MACO-1, we introduced the almost full-length genomic maco-1::GFP, which can rescue abnormal thermotaxis of maco-1 mutants (Figure S2A), into wild-type animals and observed green fluorescent protein (GFP) fluorescence. Expression of this fusion protein was observed in the cell body but not in the neurite (Figure 3A and 3B). Because MACO-1 was found in the cell body and is predicted to be a membrane protein, we investigated whether the protein is localized to organelles, such as the ER or the Golgi bodies. We made several transgenic lines co-expressing cyan fluorescent protein (CFP) - or yellow fluorescent protein (YFP)-fused MACO-1 and one of the following organelle markers: Translocating-chain associating membrane protein (CFP::TRAM) as a rER marker; Mannosidase (MANS::CFP) as a Golgi marker and membrane anchored YFP (YFP::GPI) as a plasma membrane marker [29]. We found that functional YFP::MACO-1 fusion protein was co-localized with the rER marker, CFP::TRAM in AFD neurons (Figure 3C–3E, Figure S2C and Text S1). Similarly, YFP fusion protein with human macoilin, YFP::FLJ10747, was also co-localized with CFP::TRAM in AFD neurons (Figure 3F–3H). Further, little to no co-localization of YFP::MACO-1 was observed with the Golgi marker, MANS:CFP (Figure 3I–3K), and CFP::MACO-1 was not co-localized with the plasma membrane marker, YFP::GPI (Figure 3L–3N). These results suggest that MACO-1 is mainly localized to the rER. Consistent with these findings, Arellano-Carbajal et al. showed that an antibody against MACO-1 was predominantly localized to the rER of C. elegans neurons (Arellano-Carbajal et al., personal communication).
Fig. 3. Subcellular localization of MACO-1. 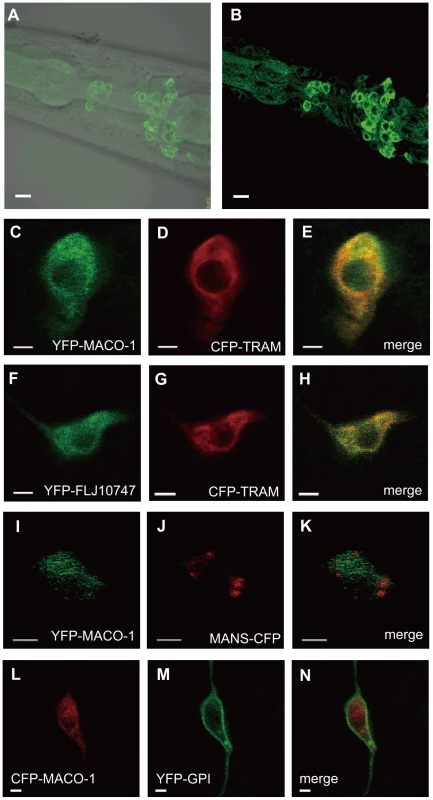
(A, B) Subcellular localization of the almost full-length genomic maco-1::GFP, including predicted transmembrane domains and coiled coil domains in wild-type animals. (A) Merged differential interference contrast (DIC) and green fluorescent protein (GFP) images. (B) GFP image. Anterior is to the left and dorsal is up. (C–N) Co-expression of MACO-1 or human ortholog of MACO-1 (FLJ10747) fused to yellow fluorescent protein (YFP) or cyan fluorescent protein (CFP) with organelle markers in AFD thermosensory neurons of wild-type animal. The following markers were used: CFP-tagged translocating chain-associating membrane protein (CFP::TRAM) as a marker of rough endoplasmic reticulum (rER); CFP-tagged mannosidase (MANS::CFP) as a marker of Golgi bodies; membrane anchored YFP (YFP::GPI) as a marker of plasma membranes [29]. (C–E) Co-expression of YFP::MACO-1 with CFP::TRAM. (F–H) Co-expression of YFP::FLJ10747 with CFP::TRAM. (I–K) Co-expression of YFP::MACO-1 with MANS::CFP. (L–N) Co-expression of YFP::GPI with CFP::MACO-1. Scale bars: A and B = 5 µm; C–N = 2 µm. MACO-1 functions in neurons and is conserved across species
To investigate the expression pattern of maco-1, we constructed the maco-1 promoter::GFP reporter that used an approximately 5-kilobase (kb) upstream region from the ATG initiation codon as a maco-1 promoter. This construct was then introduced into wild-type animals. We found that strong expression was observed in many neurons (Figure S3A and Text S1), while very weak expression was also detected in tissues. Thermotaxis defects were consistently rescued when maco-1 cDNA was expressed in almost all neurons of maco-1 mutants using the unc-14 promoter, indicating that MACO-1 acts in neurons (lower left panel of Figure 4A and Figure 4D).
Fig. 4. Cell-specific rescue experiments for thermotaxis defects in maco-1(nj34) mutants. 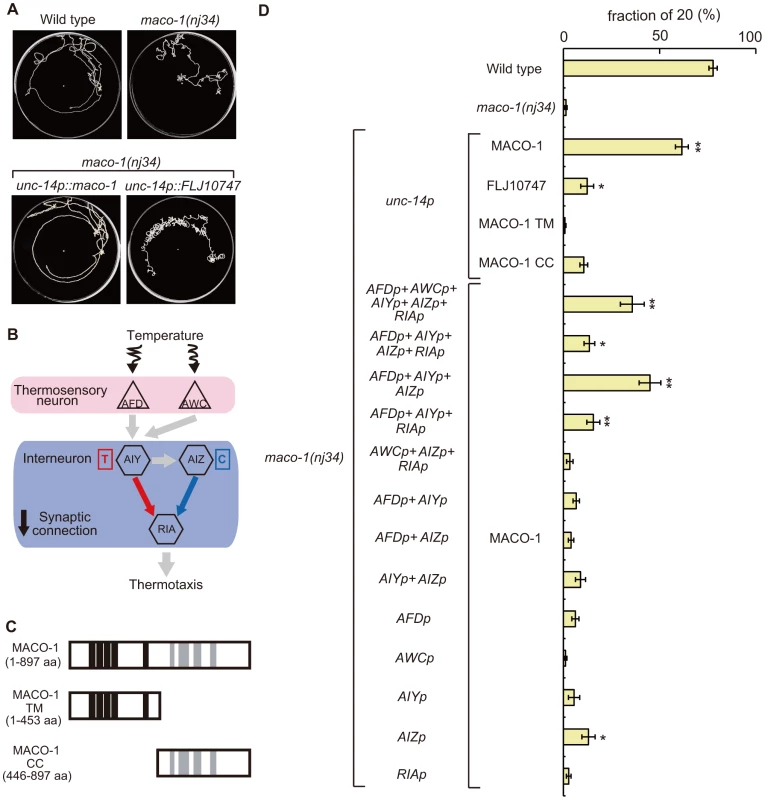
(A) Tracks of animals showing thermotaxis behavior when grown at 20°C. Upper left panel is wild-type animal. Upper right panel is maco-1(nj34) mutant. Lower left panel is maco-1(nj34) expressing maco-1 cDNA in almost all neurons. Lower right panel is maco-1(nj34) expressing human ortholog of MACO-1 in almost all neurons. The unc-14 promoter was used for expression in almost all neurons. (B) The proposed neural circuit model for thermotaxis [6]. Temperature is sensed by AFD and AWC sensory neurons. The AIY-RIA circuit is involved in thermophilic behavior (i.e., movements to temperatures higher than the cultivation temperature indicated as ‘T’). The AIZ-RIA circuit is involved in cryophilic behavior (i.e., movements to temperatures lower than the cultivation temperature indicated as ‘C’). RIA neurons are proposed to integrate these signals. (C) Truncated forms of MACO-1. MACO-1 TM (from 1 to 453 amino acids) lacks coiled coil regions and MACO-1 CC (from 446 to 897 amino acids) lacks transmembrane regions. (D) Rescue of the thermotaxis defect in maco-1 mutants by specific expression of maco-1 cDNA, human ortholog of maco-1 cDNA, MACO-1 TM or MACO-1 CC in almost all neurons using the unc-14 promoter. Specific observed expressions of maco-1 cDNA using several cell-specific promoters. Expression patterns of each promoter are listed in Table S2. Bars show the percentage of animals that moved to the cultivation temperature (20°C). Error bar indicates the standard error of the mean (SEM). Asterisks indicate statistically significant differences between fractions of 20 of 20°C-cultivated transgenic strains relative to fractions of 20 of 20°C-cultivated maco-1 mutants. One asterisk denotes statistical significance at p<0.05 and two asterisks denote statistical significance at p<0.01 (ANOVA with a Dunnett's post hoc test; n = 120–680). To examine whether MACO-1 is conserved functionally among species, we introduced FLJ10747, a predicted human ortholog of MACO-1, into almost all neurons of maco-1 mutants using the unc-14 promoter. We then evaluated the thermotaxis phenotype of the transgenic strain (lower right panel of Figure 4A, Figure 4D, Figure S2B and Text S1). FLJ10747 rescued, albeit weakly, abnormal thermotaxis phenotype of maco-1 mutants, suggesting that MACO-1 is functionally conserved across species.
Both transmembrane domains and coiled coil domains are required for appropriate function of MACO-1
MACO-1 was predicted to have transmembrane domains in the N-terminus and coiled coil domains in the C-terminus (Figure 2B). To examine whether these regions are necessary for MACO-1 function, we constructed two truncated proteins, MACO-1 TM (lacks coiled coil region) and MACO-1 CC (lacks transmembrane region), and conducted rescue experiments using each protein (Figure 4C and 4D). Expression of MACO-1 TM did not rescue in nearly all neurons of maco-1 mutants using the unc-14 promoter, while MACO-1 CC slightly rescued the abnormal thermotaxis of maco-1 mutants (Figure 4D). We also conducted rescue experiments using YFP::MACO-1 TM or YFP::MACO-1 CC fusion proteins to address whether degradation of these truncated proteins or ER quality influenced rescue experiment results. We verified that YFP fluorescence was observed in both truncated fusion proteins (Figure S4J–S4M and Text S1). YFP::MACO-1 TM did not rescue and YFP::MACO-1 CC slightly rescued defective thermotaxis of maco-1 mutants (Figure S2C and Text S1). We also found that YFP::MACO-1 TM appeared to localize to the rER (Figure S4D–S4F and Text S1). However, YFP::MACO-1 CC localization to the rER was minimal, but rather was localized to cell bodies and neurites (Figure S4G–S4I, S4L, S4M and Text S1). These results suggest that MACO-1 TM regions are necessary for appropriate MACO-1 localization, and MACO-1 CC regions are important for proper function of MACO-1.
Proper function of MACO-1 is necessary for normal thermotaxis in many neurons
To identify cells that require MACO-1 activity for sensory behavior, maco-1 cDNA was expressed by using various cell-specific promoters. Expression of MACO-1 in AFD, AWC, AIY, AIZ and RIA neurons, which are essential for thermotaxis behavior (Figure 4B and [6], [24]), partially rescued the thermotaxis defect of maco-1 mutants (Figure 4D). By contrast, specific expression of maco-1 cDNA in AFD, AWC, AIY AIZ or RIA neurons individually did not rescue or only slightly rescued the abnormal thermotaxis phenotype of maco-1 mutants (Figure 4D). Thus, we propose that simultaneous expression of MACO-1 in multiple neurons of the thermotaxis neural circuit is necessary to rescue abnormal thermotaxis among maco-1 mutants. We attempted but were unable to express MACO-1 in several combinations of neurons in maco-1 mutants to find the minimum set of neurons required to ensure full rescue of thermotaxis behavior. However, the expression of MACO-1 in AFD, AIY and AIZ neurons rescued maco-1 mutant defects similar to the expression of MACO-1 in AFD, AWC, AIY, AIZ and RIA neurons (Figure 4D). Notably, expression of MACO-1 in any other combinations of AFD, AWC, AIY, AIZ and RIA neurons only weakly rescued the defect (Figure 4D). All together, these results suggest that MACO-1 activity in AFD, AIY and AIZ neurons is required for thermotaxis behavior, and MACO-1 activity is needed in many neurons to achieve full rescue.
Abnormal thermotaxis behavior is not attributable to indirect effects of defective locomotion in maco-1 mutants
Cell-specific rescue experiments have implicated that a functional defect of MACO-1 in AFD, AIY and AIZ neurons caused defects in thermotaxis of maco-1 mutants. However, we also observed that maco-1 mutants changed their direction of movement more frequently than wild-type animals based on their tracks left during TTX assays (Figure 1B). This observation raised another possibility that locomotory defects secondarily causes the abnormal thermotaxis behavior of maco-1 mutants.
Spontaneous locomotion of C. elegans is roughly classified into two categories. The first category, runs, includes long series of sinusoidal-swimming movement. The second category, turns, is divided into two types. Omega turns involve the animal curling its head toward the tail and continues to propel forward. Reversals involve the animal moving backwards for several seconds and then moving forwards in a new direction [30], [31]. To quantify the potential locomotory defect, we constructed curvature maps of animal bodies along the head-to-tail axis using a tracking system and homemade MATLAB programs. Locomotion was represented by patterns in the curvature map (Figure 5A and 5C–5E). Wild-type animals spent most of their time moving forward (Figure 5C and left graph of 5F). Conversely, forward movement was interrupted by omega turns at short intervals among maco-1 mutant animals (Figure 5D and middle graph of 5F). A similar tendency was observed in transgenic maco-1 mutants (Figure 5E and left graph of 5F), whose abnormal thermotaxis was partially rescued by expression of maco-1 cDNA in AFD, AWC, AIY, AIZ and RIA neurons (Figure 4D). This finding supports that defective locomotion does not cause abnormal thermotaxis in maco-1 mutants.
Fig. 5. Curvature maps of animal body along the head-to-tail axis. 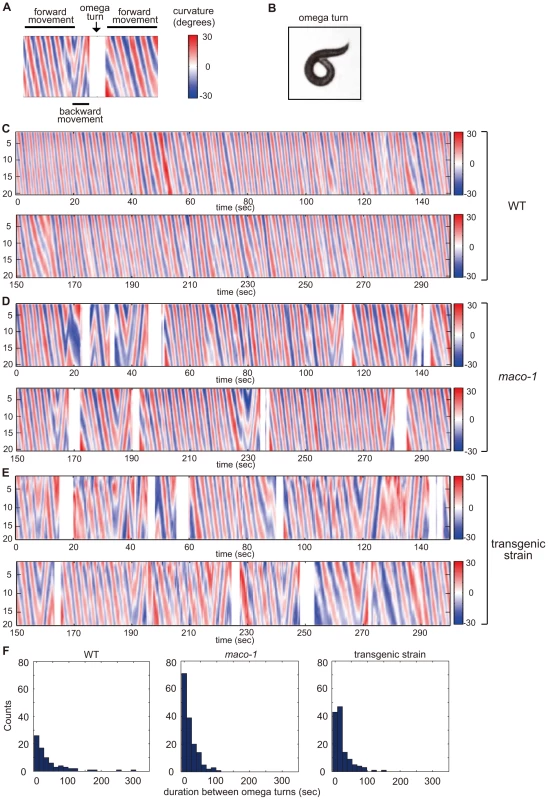
(A) Examples of locomotor states. Black bars above and below the curvature map indicate the state of forward movement and backward movement, respectively. The state of omega turn was indicated as white space (arrow). Although animals sometimes showed modest changes in direction, it is not reflected in this curvature map. (B) An image of omega turn. The head of animal curls to touch its own body. (C–E) Curvature maps of individual animals. Well-fed single animals were placed on TTX plates. Observations began 10 min after transfer. Freely moving individual animals were captured by video camera for 30 min. The x-axis and y-axis indicate time and the position along animal midlines, respectively. The color-map to the right of the graph (numbered from −30 to 30) depicts curvatures (see the Materials and Methods section for calculation procedures). (C) Wild-type. (D) maco-1(nj34). (E) maco-1(nj34) expressing maco-1 cDNA in AFD, AWC, AIY, AIZ and RIA neurons. This transgenic strain showed a partially rescued phenotype in thermotaxis behavior (Figure 4D). (F) Duration between omega turns. Initial 1000-second video images of each animal were analyzed by image-processing software, MATLAB. n = 4 (wild-type), 3 (maco-1) and 2 (transgenic strain). MACO-1 is required for proper activation of AFD and AIY neurons
Cell-specific rescue experiments indicated that MACO-1 activity, at least in AFD, AIY and AIZ neurons, is necessary for normal thermotaxis behavior (Figure 4D). To examine whether the MACO-1 functional defect influenced activities of these neurons, we conducted calcium imaging experiments and monitored responses of AFD and AIY neurons from maco-1(nj34) mutants to thermal stimuli. We found that the ratio change in AFD neurons of maco-1(nj34) mutants was significantly less than that of wild-type animals (Figure 6A). Previous work has shown that the ratio change in AFD neurons of tax-4(p678) mutants was hardly detected [21]. AFD neurons of the tax-4(p678) mutant do not respond properly to temperature because of defective cGMP-dependent cation channels composed of TAX-2 and TAX-4. The average of maximum ratio change from baseline ± SEM in maco-1(nj34) mutants was slightly larger (4.3±0.8) relative to tax-4(p678) mutants (3.3±0.6) [21] (Figure 6A), suggesting that AFD neuronal response to thermal stimuli among maco-1(nj34) mutants was not completely lost but partially reduced. Similarly, the ratio change in AIY neurons of maco-1 mutants was also lower than that of wild-type animals (Figure 6B). Taken together, our data suggest that activities of both AFD and AIY neurons in maco-1 mutants are partially decreased in response to thermal change, further indicating that MACO-1 is required for activation of AFD and AIY neurons. We also examined morphologies of these neurons in maco-1 mutants by expressing GFP in AFD and AIY neurons to determine whether the decreased activity of AFD and AIY neurons was caused by developmental or functional defects. The AFD and AIY neurons in maco-1 mutants appeared normal (Figure S3B, S3C and Text S1), suggesting that MACO-1 is not required for development but rather for proper function of AFD and AIY neurons.
Fig. 6. Calcium imaging of neurons in the thermotaxis circuit among wild-type or maco-1 mutants. 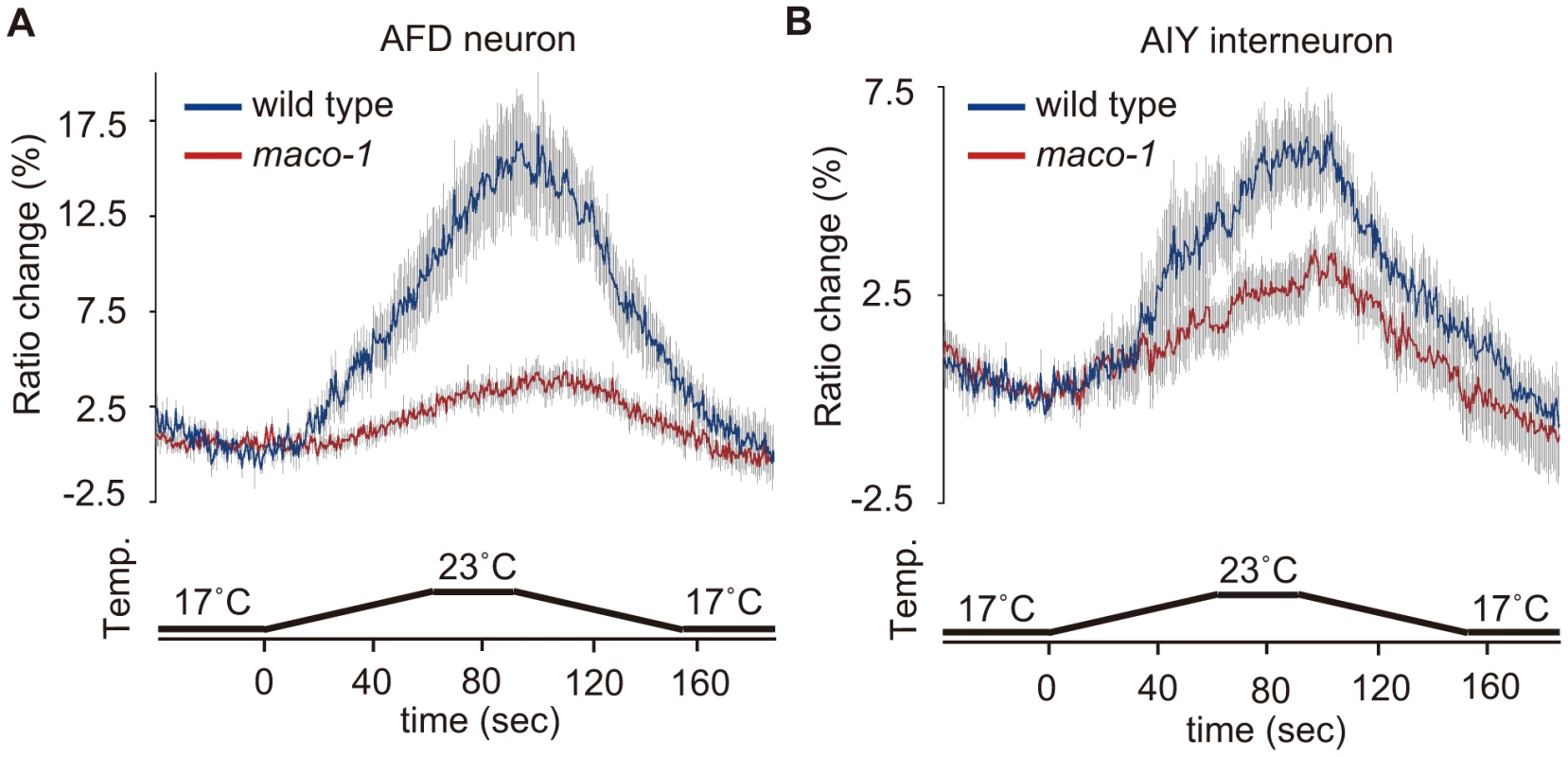
In vivo Ca2+ ratio imaging in individual genotypes cultivated at 20°C. (A) AFD thermosensory neuron. (B) AIY interneuron. Relative increases or decreases in the intracellular Ca2+ concentration were measured as increases or decreases in fluorescence of the yellow fluorescent protein/cyan fluorescent protein (YFP/CFP) of the cameleon protein ratio (ratio change). Blue and red traces represent the mean value of ratio changes of wild type and maco-1(nj34) animals, respectively, during temperature changes as indicated in the bottom panel of each graph (n = 10). Error bars (gray line) indicate the standard error of the mean (SEM). The average of the maximum ratio change from baseline ± SEM was as follows: 17.1±1.6 for wild-type (AFD); 4.3±0.8 for maco-1 (AFD); 6.3±1.0 for wild-type (AIY) and 3.8±0.6 for maco-1 (AIY). Discussion
MACO-1 is required for proper activation of neurons
We found that proper function of MACO-1 in AFD, AIY and AIZ neurons is necessary for normal thermotaxis behavior. Consistent with this result, calcium imaging suggested that MACO-1 is required for accurate activation of AFD and AIY neurons. Rescue experiments also indicate that appropriate MACO-1 function in many other interneurons and motor neurons is probably required for full rescue of abnormal thermotaxis behavior. These results suggest that MACO-1 is required broadly for proper neural activation. Consistent with this postulate, maco-1 mutants showed additional behavioral defects in chemotaxis and locomotion.
MACO-1 likely plays a role in assisting fundamental neuronal function
Co-localization studies revealed that MACO-1 was mainly localized to the rER. The rER is generally thought to be required for protein synthesis, folding and translocation of secretory and transmembrane proteins. Thus, functional defects of MACO-1 in the rER might alter these events, resulting in abnormal phenotypes in maco-1 mutants.
Previous studies have reported that Resistant to Inhibitors of Cholinesterase 3 (RIC-3) is localized to the ER and required for the maturation (the folding, assembly or transport) of nicotinic acetylcholine receptors [32]–[39]. RIC-3 is predicted to have two transmembrane domains followed by coiled-coil domains. Similar structural motifs and intracellular localization patterns of RIC-3 and MACO-1 suggest that MACO-1 might be involved in the maturation of membrane proteins like RIC-3. To explore this possibility, we observed the localization of several membrane proteins that could influence abnormal behaviors of maco-1 mutants. We found slightly increased spacing between adjacent SNB-1::GFP puncta (presynaptic marker) in DD motor neurons of maco-1 mutants compared with wild-type animals (Figure S5, Text S1 and [40], [41]). This change could be attributable to MACO-1 being directly associated with the maturation of presynaptic proteins. However, we cannot exclude the possibility of reduced excitability or altered Ca2+ signaling in maco-1 mutant neurons resulting in weak mislocalization of presynaptic proteins. Further studies on MACO-1 targets may reveal its molecular function and involvement in several intracellular events, such as the maturation of presynaptic proteins and proper neuronal activity.
Materials and Methods
Strains
C. elegans strains were maintained and grown according to standard procedures [42]. The following strains were used: wild-type Bristol strain (N2), wild-type Hawaiian strain (CB4856) for mapping with the snip-SNPs method [26], IK732 maco-1(nj21) I, IK734 maco-1(nj34) I, IK809 maco-1(tm2917) I, IK811 maco-1(ok3165) I, N2; Is[flp-13p::snb-1::gfp] II [41], and many transgenic strains derived from them. maco-1(nj21) and maco-1(nj34) mutants were isolated in genetic screens that bypassed temperature-dependent dauer formation of daf-7(e1372) backgrounds [43]. All mutant strains used in this study were backcrossed to N2 one to ten times before characterization.
Behavioral analysis
Animals were grown at 20°C for all behavioral assays in this study, unless otherwise described. The procedure for the thermotaxis assay using radial temperature gradients was based on previously published research by Mori and Ohshima [6] with some modification. Evaluation of thermotaxis was consistent with methods of Mohri et al. [7]. Thermotaxis of individual animals on a radial temperature gradient was evaluated using the following four phenotypic categories: animals that moved to the cold region (i.e., center of plate) were classified as ‘17’; animals that moved to the 20°C region were classified as ‘20’; animals that moved to warm regions (i.e., plate periphery) were classified as ‘25’; animals that moved to cold and warm regions were classified as ‘17/25’. Chemotaxis to odorants was assayed according to Bargmann et al. [44], except that the assay plates contained a slightly different medium (2% agar, 1 mM magnesium sulfate (MgSO4), 1 mM calcium chloride (CaCl2) and 25 mM potassium phosphate (pH 6)). Dilutions of odorants with ethanol were as follows: 1∶1000 diacetyl, 1 mg/ml pyrazine, 1∶100 isoamyl alcohol, and 1∶100 benzaldehyde. The chemotaxis index was calculated according to Bargmann et al. [44]. Chemotaxis to NaCl was essentially as described previously [15]. Briefly, animals were scored as ‘normal’ when they migrated repeatedly to the concentration peak or remained at the peak during the assay. The animal was scored as ‘partially defective’ when it migrated toward the peak only once or stayed very briefly at the peak. The animal was scored as ‘defective’ when it failed to migrate to the peak.
Molecular biology
The maco-1 promoter::GFP (pMYA3) used in Figure S3A was constructed by PCR amplification of the maco-1 promoter region (i.e., the 4900-bp upstream sequence of maco-1). This fragment was then inserted into the GFP vector pPD95.77 to generate pMYA3. The almost full-length genomic maco-1::GFP (pMYA52) was constructed by PCR amplification of the 5′ 5-kilobase (kb) region of maco-1 gene and 2-kb fragment downstream of 5′ 5-kb fragment using the cosmid C09C9 and wild-type genomic DNA, respectively. These fragments were inserted between the maco-1 promoter and GFP of pMYA3 to generate the almost full-length genomic maco-1::GFP (pMYA52). The almost full-length genomic maco-1 contains a maco-1 promoter region and a 6621-bp fragment of a 7536-bp full-length genomic maco-1 sequence, which includes predicted transmembrane domains and coiled coil domains. The full-length maco-1 cDNA was constructed by PCR amplifications of the 5′ partial maco-1 cDNA (723-bp region of 2694-bp full-length maco-1 cDNA) and the rest of maco-1 cDNA region using a cDNA library and yk1296a05 clone, respectively. These fragments were ligated into pPD49.26 to generate pMYA17. All cell-specific promoter::maco-1 cDNA plasmids were generated by inserting each cell-specific promoter fragment into pMYA17. Cell-specific promoters are as follows: unc-14p for almost all neurons [45]; gcy-8p and gcy-18p for AFD neurons [17], [46]; ttx-3p for AIY neurons [43]; lin-11p for AIZ neurons (unpublished data); odr-1p for AWC neurons [47] and glr-3p for RIA neurons (unpublished data). Detailed expression patterns of these cell-specific promoters are shown in Table S2. The fluorescent protein fused maco-1 cDNA and several organelle markers used in Figure 3 were also constructed. Specifically, the glr-1 promoter of glr-1p::YFP::TRAM [29] was replaced with gcy-8 promoter to generate gcy-8p::YFP::TRAM (pMYA161). The TRAM of pMYA161 was replaced with maco-1 cDNA to generate gcy-8p::YFP::maco-1 (pMYA174). The YFP of pMYA161 was replaced with CFP to generate gcy-8p::CFP::TRAM (pMYA212). The TRAM of pMYA161 was replaced with FLJ10747 cDNA to generate gcy-8p::YFP::FLJ10747 (pMYA182). The glr-1 promoter of glr-1p::MANS::YFP [29] was replaced with gcy-8 promoter to generate gcy-8p::MANS::YFP (pMYA145), and then YFP of pMYA145 was replaced with CFP to generate gcy-8p::MANS::CFP (pMYA208). The glr-1 promoter of glr-1p::YFP::GPI [29] was replaced with the gcy-8 promoter to generate gcy-8p::YFP::GPI (pMYA153). The glr-1 promoter of glr-1p::CFP::PIS [29] was replaced with the gcy-8 promoter to generate gcy-8p::CFP::PIS (pMYA141). The PIS of pMYA141 was replaced with maco-1 cDNA to generate gcy-8p::CFP::maco-1 (pMYA176). Two types of truncated forms of MACO-1, MACO-1 TM (1–453 amino acid) and MACO-1 CC (446–897 amino acid) (Figure 4), were PCR amplified from unc-14p::maco-1 cDNA and replaced with full-length maco-1 cDNA of unc-14p::maco-1 to generate unc-14p::MACO-1 TM (pMYA107) and unc-14p::MACO-1 CC (pMYA109), respectively. The unc-14p::FLJ10747 shown in Figure 4 was constructed by ligation of the unc-14 promoter into pPD49.26 vector to generate pMYA171. The full-length cDNA of human macoilin, FLJ10747 cDNA, was PCR amplified from a Mammalian Gene Collection (MGC) clone (Invitrogen), which contained full-length FLJ10747 cDNA. This product was inserted into pMYA171 to generate unc-14p::FLJ10747 (pMYA172).
Germline transformation
Transgenic strains were constructed by injecting DNA into the gonad of adult animals [48]. The almost full-length genomic maco-1::GFP used in Figure 3A, 3B and Figure S2A was injected into wild-type animals at a concentration of 50 ng/µl. Transgenic animals were recognized with neuronal GFP expression. Figure 3C–3E depicts the co-injection of gcy-8p::YFP::maco-1 (pMYA174) (20 ng/µl) and gcy-8p::CFP::TRAM (pMYA212) (20 ng/µl) into wild-type animals. This injection was coupled with 120 ng/µl of pTAN124.5 (ges-1p::DsRed) as a co-injection marker. Figure 3F–3H shows that gcy-8p::YFP::FLJ10747 (pMYA182) (50 ng/µl) and gcy-8p::CFP::TRAM (pMYA212) (4 ng/µl) were co-injected into wild-type animals with ges-1p::DsRed (pTAN124.5) (120 ng/µl) as a co-injection marker. Figure 4D shows co-injections of test DNA and the injection marker, ges-1p::NLS-GFP (pKDK66) [49], into the strain IK734, maco-1(nj34) at 2 and 50 ng/µl, respectively.
Video analysis
Animal behavior was recorded using a custom made tracking and live imaging system. Well-fed single animal was placed on a TTX plate for tracking experiments. An ordinal analog video camera (Olympus) captured freely moving individual animal for 30 min with adequate sampling rate (30 frames/sec). A computer-controlled microscope stage was automatically moved to center animals in the visual field using a custom image analysis algorithm with microscope software (MetaMorph: Universal Imaging Corporation). Midlines of recorded animals were extracted from each image, and curvatures at certain positions were calculated by analyzing points along midline with constant intervals. Spatio-temporal curvature maps were constructed by expressing the quantity of curvatures with a color-map (x-axis = time and y-axis = position along midline of animal). A smoothing filter (3×3 dimensions) was applied to the curvature map to reduce noise effects. Omega turn states (i.e., head of the animal curls to touch its own body; Figure 5B) were identified by an independent algorithm and expressed as white color in the curvature map. All video images were analyzed with MATLAB (Mathworks, Natick, MA).
In vivo calcium imaging and data analysis
In vivo calcium imaging was performed according to previous studies [21], [50]. Well-fed animals expressing yellow cameleon 3.60 [51] caused by AFD and AIY promoters, gcy-8p::yc3.60 (pSAS309) and AIYp::yc3.60 (pNR86.79), were glued onto a 2% agar pad on glass, immersed in M9 buffer, and covered by cover glass. The agar pad and M9 buffer were kept at the initial imaging temperature. Sample preparation was completed within one minute. The sample was then placed onto a Peltier-based thermocontroller (Tokai Hit, Japan) on the stage of a Nikon E600 or Olympus BX61WI microscope at the initial imaging temperature for two minutes. Fluorescence was introduced into W-View (Hamamatsu Photonics, Japan) or Dual-View (Molecular devices, USA) optic systems. Cyan fluorescent protein (CFP; F480) and yellow fluorescent protein (YFP; F535) images were simultaneously captured by a CCD camera HiSCA or EM-CCD camera C9000-13 ImagEM (Hamamatsu Photonics). Images were taken with a 200-ms exposure time with 4×4 or 2×2 binning. Temperatures on the agar pad were monitored by the thermometer system, DCM-20 (Tokai Hit and Hamamatsu Photonics). For each imaging experiment, fluorescence intensities of F535 and F480 were measured using AquaCosmos (Hamamatsu Photonics) or MetaMorph (Molecular Device) imaging analysis systems. Relative increases or decreases in intracellular Ca2+ concentrations were measured as increases or decreases in YFP/CFP fluorescence ratios of cameleon protein (ratio change).
Statistical analysis
Error bars in figures indicate standard errors (SEM). The statistical analysis for behavioral experiments was performed by one-way analysis of variance (ANOVA) for multiple comparisons followed by post hoc Dunnett's multiple comparison. A single asterisk and double asterisk indicate statistical significance at p<0.05 and p<0.01, respectively. The value of p is probability.
Supporting Information
Zdroje
1. International Human Genome Sequencing Consortium 2004 Finishing the euchromatic sequence of the human genome. Nature 431 931 945
2. AdamsMD
CelnikerSE
HoltRA
EvansCA
GocayneJD
2000 The genome sequence of Drosophila melanogaster. Science 287 2185 2195
3. The C. elegans Sequencing Consortium 1998 Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282 2012 2018
4. WhiteJ
SouthgateE
ThompsonJ
BrennerS
1986 The structure of the nervous system of the nematode C. elegans. Philosophical Transactions of the Royal Society of London - Series B Biological Sciences 314 1 340
5. HedgecockEM
RussellRL
1975 Normal and mutant thermotaxis in the nematode Caenorhabditis elegans. Proceedings of the National Academy of Sciences of the United States of America 72 4061 4065
6. MoriI
OhshimaY
1995 Neural regulation of thermotaxis in Caenorhabditis elegans. Nature 376 344 348
7. MohriA
KodamaE
KimuraKD
KoikeM
MizunoT
2005 Genetic control of temperature preference in the nematode Caenorhabditis elegans. Genetics 169 1437 1450
8. RyuWS
SamuelAD
2002 Thermotaxis in Caenorhabditis elegans analyzed by measuring responses to defined Thermal stimuli. J Neurosci 22 5727 5733
9. YamadaY
OhshimaY
2003 Distribution and movement of Caenorhabditis elegans on a thermal gradient. The Journal of experimental biology 206 2581 2593
10. ClarkDA
GabelCV
LeeTM
SamuelAD
2007 Short-term adaptation and temporal processing in the cryophilic response of Caenorhabditis elegans. Journal of neurophysiology 97 1903 1910
11. RamotD
MacInnisBL
LeeHC
GoodmanMB
2008 Thermotaxis is a robust mechanism for thermoregulation in Caenorhabditis elegans nematodes. J Neurosci 28 12546 12557
12. NakazatoK
MochizukiA
2009 Steepness of thermal gradient is essential to obtain a unified view of thermotaxis in C. elegans. Journal of theoretical biology 260 56 65
13. JuradoP
KodamaE
TanizawaY
MoriI
2010 Distinct thermal migration behaviors in response to different thermal gradients in Caenorhabditis elegans. Genes, brain, and behavior 9 120 127
14. GomezM
De CastroE
GuarinE
SasakuraH
KuharaA
2001 Ca2+ signaling via the neuronal calcium sensor-1 regulates associative learning and memory in C. elegans. Neuron 30 241 248
15. KomatsuH
MoriI
RheeJS
AkaikeN
OhshimaY
1996 Mutations in a cyclic nucleotide-gated channel lead to abnormal thermosensation and chemosensation in C. elegans. Neuron 17 707 718
16. MoriI
1999 Genetics of chemotaxis and thermotaxis in the nematode Caenorhabditis elegans. Annual review of genetics 33 399 422
17. InadaH
ItoH
SatterleeJ
SenguptaP
MatsumotoK
2006 Identification of guanylyl cyclases that function in thermosensory neurons of Caenorhabditis elegans. Genetics 172 2239 2252
18. KuharaA
InadaH
KatsuraI
MoriI
2002 Negative regulation and gain control of sensory neurons by the C. elegans calcineurin TAX-6. Neuron 33 751 763
19. OkochiY
KimuraKD
OhtaA
MoriI
2005 Diverse regulation of sensory signaling by C. elegans nPKC-epsilon/eta TTX-4. The EMBO journal 24 2127 2137
20. MiyawakiA
LlopisJ
HeimR
McCafferyJM
AdamsJA
1997 Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388 882 887
21. KimuraKD
MiyawakiA
MatsumotoK
MoriI
2004 The C. elegans thermosensory neuron AFD responds to warming. Curr Biol 14 1291 1295
22. ClarkDA
BironD
SenguptaP
SamuelAD
2006 The AFD sensory neurons encode multiple functions underlying thermotactic behavior in Caenorhabditis elegans. J Neurosci 26 7444 7451
23. BironD
WassermanS
ThomasJH
SamuelAD
SenguptaP
2008 An olfactory neuron responds stochastically to temperature and modulates Caenorhabditis elegans thermotactic behavior. Proceedings of the National Academy of Sciences of the United States of America 105 11002 11007
24. KuharaA
OkumuraM
KimataT
TanizawaY
TakanoR
2008 Temperature sensing by an olfactory neuron in a circuit controlling behavior of C. elegans. Science 320 803 807
25. KuvbachievaA
BestelAM
TissirF
MaloumI
GuimiotF
2004 Identification of a novel brain-specific and Reelin-regulated gene that encodes a protein colocalized with synapsin. The European journal of neuroscience 20 603 610
26. WicksSR
YehRT
GishWR
WaterstonRH
PlasterkRH
2001 Rapid gene mapping in Caenorhabditis elegans using a high density polymorphism map. Nature genetics 28 160 164
27. KroghA
LarssonB
von HeijneG
SonnhammerEL
2001 Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. Journal of molecular biology 305 567 580
28. LupasA
Van DykeM
StockJ
1991 Predicting coiled coils from protein sequences. Science 252 1162 1164
29. RollsMM
HallDH
VictorM
StelzerEH
RapoportTA
2002 Targeting of rough endoplasmic reticulum membrane proteins and ribosomes in invertebrate neurons. Molecular biology of the cell 13 1778 1791
30. CrollNA
1975 Behavioural analysis of nematode movement. Advances in parasitology 13 71 122
31. CrollNA
1975 Components and patterns in the behaviour of the nematode Caenorhabditis elegans. J Zool 176 159 176
32. HaleviS
McKayJ
PalfreymanM
YassinL
EshelM
2002 The C. elegans ric-3 gene is required for maturation of nicotinic acetylcholine receptors. The EMBO journal 21 1012 1020
33. HaleviS
YassinL
EshelM
SalaF
SalaS
2003 Conservation within the RIC-3 gene family. Effectors of mammalian nicotinic acetylcholine receptor expression. The Journal of biological chemistry 278 34411 34417
34. CastilloM
MuletJ
GutierrezLM
OrtizJA
CastelanF
2005 Dual role of the RIC-3 protein in trafficking of serotonin and nicotinic acetylcholine receptors. The Journal of biological chemistry 280 27062 27068
35. LansdellSJ
GeeVJ
HarknessPC
DowardAI
BakerER
2005 RIC-3 enhances functional expression of multiple nicotinic acetylcholine receptor subtypes in mammalian cells. Molecular pharmacology 68 1431 1438
36. WilliamsME
BurtonB
UrrutiaA
ShcherbatkoA
Chavez-NoriegaLE
2005 Ric-3 promotes functional expression of the nicotinic acetylcholine receptor alpha7 subunit in mammalian cells. The Journal of biological chemistry 280 1257 1263
37. GottschalkA
SchaferWR
2006 Visualization of integral and peripheral cell surface proteins in live Caenorhabditis elegans. Journal of neuroscience methods 154 68 79
38. ChengA
BollanKA
GreenwoodSM
IrvingAJ
ConnollyCN
2007 Differential subcellular localization of RIC-3 isoforms and their role in determining 5-HT3 receptor composition. The Journal of biological chemistry 282 26158 26166
39. LansdellSJ
CollinsT
YabeA
GeeVJ
GibbAJ
2008 Host-cell specific effects of the nicotinic acetylcholine receptor chaperone RIC-3 revealed by a comparison of human and Drosophila RIC-3 homologues. Journal of neurochemistry 105 1573 1581
40. KimK
LiC
2004 Expression and regulation of an FMRFamide-related neuropeptide gene family in Caenorhabditis elegans. The Journal of comparative neurology 475 540 550
41. Sakaguchi-NakashimaA
MeirJY
JinY
MatsumotoK
HisamotoN
2007 LRK-1, a C. elegans PARK8-related kinase, regulates axonal-dendritic polarity of SV proteins. Curr Biol 17 592 598
42. BrennerS
1974 The genetics of Caenorhabditis elegans. Genetics 77 71 94
43. HobertO
MoriI
YamashitaY
HondaH
OhshimaY
1997 Regulation of interneuron function in the C. elegans thermoregulatory pathway by the ttx-3 LIM homeobox gene. Neuron 19 345 357
44. BargmannCI
HartwiegE
HorvitzHR
1993 Odorant-selective genes and neurons mediate olfaction in C. elegans. Cell 74 515 527
45. OguraK
ShirakawaM
BarnesTM
HekimiS
OhshimaY
1997 The UNC-14 protein required for axonal elongation and guidance in Caenorhabditis elegans interacts with the serine/threonine kinase UNC-51. Genes & development 11 1801 1811
46. WedelBJ
GarbersDL
1997 New insights on the functions of the guanylyl cyclase receptors. FEBS letters 410 29 33
47. L'EtoileND
BargmannCI
2000 Olfaction and odor discrimination are mediated by the C. elegans guanylyl cyclase ODR-1. Neuron 25 575 586
48. MelloCC
KramerJM
StinchcombD
AmbrosV
1991 Efficient gene transfer in C.elegans: extrachromosomal maintenance and integration of transforming sequences. The EMBO journal 10 3959 3970
49. FukushigeT
SchroederDF
AllenFL
GoszczynskiB
McGheeJD
1996 Modulation of gene expression in the embryonic digestive tract of C. elegans. Developmental biology 178 276 288
50. KuharaA
MoriI
2006 Molecular physiology of the neural circuit for calcineurin-dependent associative learning in Caenorhabditis elegans. J Neurosci 26 9355 9364
51. NagaiT
YamadaS
TominagaT
IchikawaM
MiyawakiA
2004 Expanded dynamic range of fluorescent indicators for Ca(2+) by circularly permuted yellow fluorescent proteins. Proceedings of the National Academy of Sciences of the United States of America 101 10554 10559
52. ThompsonJD
HigginsDG
GibsonTJ
1994 CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic acids research 22 4673 4680
53. PerriereG
GouyM
1996 WWW-query: an on-line retrieval system for biological sequence banks. Biochimie 78 364 369
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2011 Číslo 5
-
Všechny články tohoto čísla
- Structural and Functional Differences in the Long Non-Coding RNA in Mouse and Human
- Identification, Replication, and Functional Fine-Mapping of Expression Quantitative Trait Loci in Primary Human Liver Tissue
- A −436C>A Polymorphism in the Human Gene Promoter Associated with Severe Childhood Malaria
- A Decline in p38 MAPK Signaling Underlies Immunosenescence in
- The Operon Balances the Requirements for Vegetative Stability and Conjugative Transfer of Plasmid R388
- Novel and Conserved Protein Macoilin Is Required for Diverse Neuronal Functions in
- Ixr1 Is Required for the Expression of the Ribonucleotide Reductase Rnr1 and Maintenance of dNTP Pools
- Genome of Strain SmR1, a Specialized Diazotrophic Endophyte of Tropical Grasses
- A Deficiency of Ceramide Biosynthesis Causes Cerebellar Purkinje Cell Neurodegeneration and Lipofuscin Accumulation
- A Latent Pro-Survival Function for the Mir-290-295 Cluster in Mouse Embryonic Stem Cells
- Association of Genetic Variants in Complement Factor H and Factor H-Related Genes with Systemic Lupus Erythematosus Susceptibility
- DNA Methylation Dynamics in Human Induced Pluripotent Stem Cells over Time
- Prion Formation and Polyglutamine Aggregation Are Controlled by Two Classes of Genes
- Integrated Genome-Scale Prediction of Detrimental Mutations in Transcription Networks
- Post-Embryonic Nerve-Associated Precursors to Adult Pigment Cells: Genetic Requirements and Dynamics of Morphogenesis and Differentiation
- A Novel Mouse Synaptonemal Complex Protein Is Essential for Loading of Central Element Proteins, Recombination, and Fertility
- STAT Is an Essential Activator of the Zygotic Genome in the Early Embryo
- A Genetic and Structural Study of Genome Rearrangements Mediated by High Copy Repeat Ty1 Elements
- A Missense Mutation in Causes a Major QTL Effect on Ear Size in Pigs
- A Flurry of Folding Problems: An Interview with Susan Lindquist
- Meiotic Recombination Intermediates Are Resolved with Minimal Crossover Formation during Return-to-Growth, an Analogue of the Mitotic Cell Cycle
- A Nervous Origin for Fish Stripes
- The ISWI Chromatin Remodeler Organizes the hsrω ncRNA–Containing Omega Speckle Nuclear Compartments
- The Telomerase Subunit Est3 Binds Telomeres in a Cell Cycle– and Est1–Dependent Manner and Interacts Directly with Est1
- Nodal-Dependent Mesendoderm Specification Requires the Combinatorial Activities of FoxH1 and Eomesodermin
- SHINE Transcription Factors Act Redundantly to Pattern the Archetypal Surface of Arabidopsis Flower Organs
- Characterizing Genetic Risk at Known Prostate Cancer Susceptibility Loci in African Americans
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Nodal-Dependent Mesendoderm Specification Requires the Combinatorial Activities of FoxH1 and Eomesodermin
- SHINE Transcription Factors Act Redundantly to Pattern the Archetypal Surface of Arabidopsis Flower Organs
- Association of Genetic Variants in Complement Factor H and Factor H-Related Genes with Systemic Lupus Erythematosus Susceptibility
- STAT Is an Essential Activator of the Zygotic Genome in the Early Embryo
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



