-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaThe TALE Class Homeobox Gene Defines the Anterior Compartment for Head Regeneration
Planaria continue to blossom as a model system for understanding all aspects of regeneration. They provide an opportunity to understand how the replacement of missing tissues from preexisting adult tissue is orchestrated at the molecular level. When amputated along any plane, planaria are capable of regenerating all missing tissue and rescaling all structures to the new size of the animal. Recently, rapid progress has been made in understanding the developmental pathways that control planarian regeneration. In particular Wnt/beta-catenin signaling is central in promoting posterior fates and inhibiting anterior identity. Currently the mechanisms that actively promote anterior identity remain unknown. Here, Smed-prep, encoding a TALE class homeodomain, is described as the first gene necessary for correct anterior fate and patterning during planarian regeneration. Smed-prep is expressed at high levels in the anterior portion of whole animals, and Smed-prep(RNAi) leads to loss of the whole brain during anterior regeneration, but not during lateral regeneration or homeostasis in intact worms. Expression of markers of different anterior fated cells are greatly reduced or lost in Smed-prep(RNAi) animals. We find that the ectopic anterior structures induced by abrogation of Wnt signaling also require Smed-prep to form. We use double knockdown experiments with the S. mediterranea ortholog of nou-darake (that when knocked down induces ectopic brain formation) to show that Smed-prep defines an anterior fated compartment within which stem cells are permitted to assume brain fate, but is not required directly for this differentiation process. Smed-prep is the first gene clearly implicated as being necessary for promoting anterior fate and the first homeobox gene implicated in establishing positional identity during regeneration. Together our results suggest that Smed-prep is required in stem cell progeny as they form the anterior regenerative blastema and is required for specifying anterior cell fates and correct patterning.
Published in the journal: . PLoS Genet 6(4): e32767. doi:10.1371/journal.pgen.1000915
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1000915Summary
Planaria continue to blossom as a model system for understanding all aspects of regeneration. They provide an opportunity to understand how the replacement of missing tissues from preexisting adult tissue is orchestrated at the molecular level. When amputated along any plane, planaria are capable of regenerating all missing tissue and rescaling all structures to the new size of the animal. Recently, rapid progress has been made in understanding the developmental pathways that control planarian regeneration. In particular Wnt/beta-catenin signaling is central in promoting posterior fates and inhibiting anterior identity. Currently the mechanisms that actively promote anterior identity remain unknown. Here, Smed-prep, encoding a TALE class homeodomain, is described as the first gene necessary for correct anterior fate and patterning during planarian regeneration. Smed-prep is expressed at high levels in the anterior portion of whole animals, and Smed-prep(RNAi) leads to loss of the whole brain during anterior regeneration, but not during lateral regeneration or homeostasis in intact worms. Expression of markers of different anterior fated cells are greatly reduced or lost in Smed-prep(RNAi) animals. We find that the ectopic anterior structures induced by abrogation of Wnt signaling also require Smed-prep to form. We use double knockdown experiments with the S. mediterranea ortholog of nou-darake (that when knocked down induces ectopic brain formation) to show that Smed-prep defines an anterior fated compartment within which stem cells are permitted to assume brain fate, but is not required directly for this differentiation process. Smed-prep is the first gene clearly implicated as being necessary for promoting anterior fate and the first homeobox gene implicated in establishing positional identity during regeneration. Together our results suggest that Smed-prep is required in stem cell progeny as they form the anterior regenerative blastema and is required for specifying anterior cell fates and correct patterning.
Introduction
Planaria continue to blossom as a model system for understanding all aspects of regeneration [1]–[3]. A sustained and passionate effort by a number of scientists is pushing planaria to the forefront of the regeneration field, both technically [4], [5] and theoretically [6], and they are finally starting to be directly informative of phenomena in other systems [7]. They provide an opportunity to understand how the replacement of missing tissues from preexisting adult tissue is orchestrated at the molecular level. When amputated along any plane planaria are capable of regenerating all missing tissue and rescaling all structures to the new size of the animal [8].
Recent work has shown that conserved signaling pathways play a role in axial patterning during both regeneration and homeostatic tissue turnover [9]–[13]. In particular Wnt/beta-catenin signaling is necessary for posterior fate during regeneration, with loss of beta-catenin or Wnt signaling leading to all amputations regenerating anterior structures and a gradual loss of posterior identity during homeostasis [9], [10], [12]. Conversely, over activity of Wnt signaling induced by abrogating the expression of negative regulators of the pathways leads to ectopic posterior fate [9]. Further studies have begun to describe the temporal nature of this posterior specification circuit, as well the conserved nature of upstream regulation [14], [15].
Previously elegantly executed manipulative work has uncovered phenomena that suggest that anterior fated tissue can inhibit the regeneration of anterior fate elsewhere [3]. In addition some headway has been made in understanding the potential signaling systems responsible for this [16], [17]. In particular the planarian nou-darake (ndk) gene, an FGF-like receptor, has been shown to be necessary to restrict the formation of anterior-dorsal brain ganglia/cephalic ganglia (CG) to anterior regions [16]. Currently though nothing is known about the instructive signals required to promote anterior fate. We wished to uncover these signals that together must promote anterior fate and correctly pattern the brain as it reforms from stem cell progeny at anterior blastemas.
Given the involvement of conserved pathways already uncovered we hypothesized that other genetic circuits employed to specify positional domains in other animals would be responsible for this process during planarian regeneration. One obvious group of genes for this process would be planarian orthologs of the Hox genes and Hox gene co-factors, These are required for anterior-posterior axis specification in the metazoa [18], [19]. Planarian Hox orthologs have been previously studied, and in some cases are expressed in distinct spatial domains, but have as yet no functions are assigned to them in planaria.
This has led us to consider TALE class homeodomain containing genes, a subset of which act as Hox gene cofactors [18]. Collectively, they are known to modulate the activity of Hox proteins by regulating their localization within the cell and by increasing their binding site specificity, but also have many hox independent roles in development [20]–[23].
Here, Smed-prep, encoding a TALE class homeodomain, is described as the first gene that is necessary to instruct anterior fate and patterning during planarian regeneration.
Results/Discussion
Smed-prep encodes a TALE class homeodomain protein expressed in regeneration blastemas
The Smed-prep transcript was identified in an informatics screen for homeodomain proteins in the Schmidtea mediterranea genome. Searching the S. mediterranea genome identifies other TALE class homeodomain proteins [18], but Smed-prep encodes the only PREP ortholog (Figure 1A). The protein encoded by Smed-prep has high homology to other PREP proteins and contains the conserved features expected of this protein family (Figure S1). In vertebrates, PREP proteins have been implicated in a number of key developmental processes [23], including the correct patterning of anterior structures [21]. The function of Hox and Hox co-factors in planaria remains enigmatic. The fact that these two groups of homeodomains act together to pattern tissues in other systems makes them strong candidates for a role in providing positional information in planarians. For this reason we performed a detailed study of Smed-prep.
Fig. 1. Smed-Prep encodes a TALE Homeobox gene expressed in regenerating blastemas. 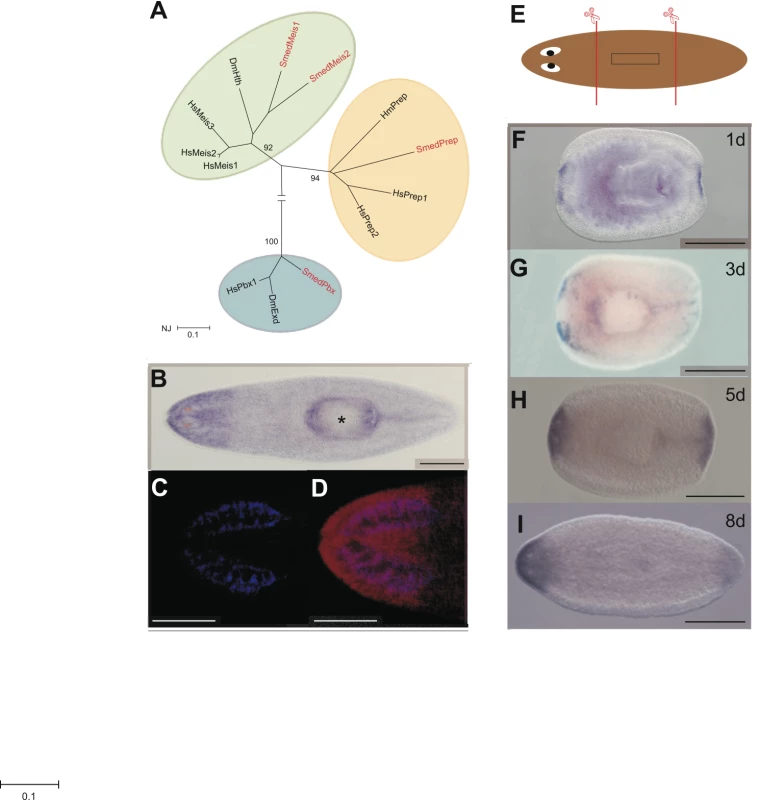
(A) Phylogenetic reconstruction of S. mediterranea TALE Class homeodomain proteins and representative orthologs, with most taxa removed for clarity (Hs: Homo sapien, Dm: Drosophila melanogaster Hm: Hydra magnipapillata), produced using a neighbor joining method and 500 bootstrap replicates. (B) Smed-prep expression in whole worms shows a distinct anterior domain of high expression. (C,D) demonstrate that the posterior margin of high Smed-prep expression coincides with posterior end of the brain. DAPI staining (blue) to highlight the brain (C) combined with false coloring of Smed-prep (red) expression (D). (E) Standard amputation protocol to assess expression during regeneration and regeneration phenotypes of RNAi experiments. Animals are cut pre- and post-pharyngeal to generate regenerating head, trunk and tail fragments. Expression of Smed-prep in regeneration blastemas is present in anterior and posterior blastemas in regenerating trunck pieces at 1 day (F), 3 days (G), 5 days (H), and 8 days (I) after amputation. Expression at 5 days clearly shows an absence of expression in the eye field, posterior expression at 8 days is reduced. All scale bars are 1 mm. Asterix indicates the pharynx. We performed in situ hybridization on whole and regenerating asexual planaria [24], [25]. We find that Smed-prep is expressed at ubiquitously low levels throughout the parenchyma and at higher levels in the head region. The posterior margin of anterior expression coincides with the most posterior position of cephalic ganglia (CG) (Figure 1C and 1D). We also detect low levels of Smed-prep expression in the posterior midline, at higher levels than the broad parenchymal expression, in approximately 50% (39/72) of animals (Figure 1B). Smed-prep expression is not sensitive to irradiation, indicating that Smed-prep is not expressed in, or dependent on, the ‘neoblast’ stem cells (data not shown). During regeneration induced by pre - and post-pharyngeal amputation (Figure 1E) Smed-prep expression is first detected at 24 h and is present in both anterior and posterior blastemas (Figure 1F). New Smed-prep expression is not detected at 6, 12 or 18 hours of regeneration. Expression in the anterior is bilateral up to 3 days but has expanded across the whole blastema at 5 days (Figure 1G and 1H). At 5 days Smed-prep is expressed throughout the anterior compartment with the notable exception of the eye field. We also detect feint expression in the posterior midline of approximately 50% of trunk fragments at 3 (18/41 fragments) and 5 days (23/40 fragments) of regeneration. We observe this in trunk fragments only (Figure 1G and 1H). This expression is absent later and presumably reappears after regeneration is complete and animals reach a homeostatic state (see above). At 8 days of regeneration, posterior blastema expression is reduced while expression in the anterior continues to be high (Figure 1I). This expression pattern led us to hypothesize a role for Smed-prep in patterning regenerating tissue after amputation. In particular expression in whole worms suggested that Smed-prep might have a role in pattering and/or maintaining anterior structures.
Smed-prep(RNAi) results in loss of anterior structures specifically during anterior regeneration
We performed RNAi [26], [27] of Smed-prep to investigate its function during regeneration (see Figure S2 for summary of injection protocols). Smed-prep dsRNA injection before inducing regeneration by amputation (Figure 1E) resulted in all worms having either a cyclops phenotype (Figure 2A) or no eyes at all (Figure 2B, Table 1). All animals had correct early blastema formation, normal levels of neoblast proliferation (data not shown) and no defects in posterior blastema formation (Figure 2A, 2B, 2D, and 2E). A similar cyclops phenotype has been described for a S. mediterranea slit ortholog [28]. Staining with an anti-arrestin VC-1 antibody specific for planarian photoreceptors and associated neurons [29] we observed that the single eye phenotype appeared to represent a fusion of two eyes (Figure S2D, S2E). We detected no other midline defects in regenerating animals that were described for Smed-slit, and Smed-slit expression itself was normal (Figure S2F and S2G). This suggests, in agreement with the Smed-prep expression pattern, that the cyclops phenotype is due to a defect in anterior patterning and fate rather than any midline defects. Control gfp(RNAi) animals had normal eye structure (Figure S2E).
Fig. 2. Smed-prep(RNAi) leads to the loss of anterior fate during regeneration. 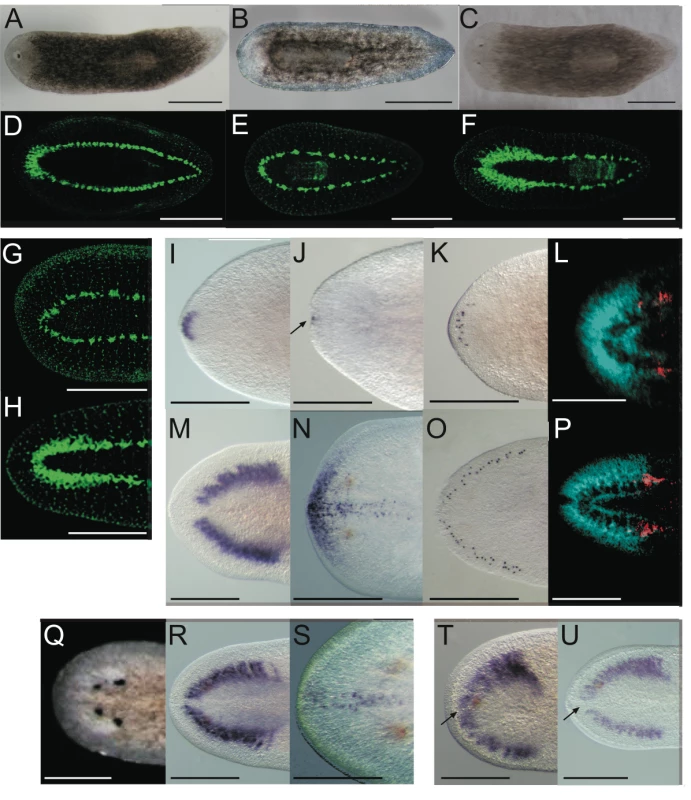
Smed-prep(RNAi) using a standard injecting and cutting protocol (Figure S2A) leads to animals with either one (A) or no eyes (B). Control gfp(RNAi) animals were all normal (C). Staining with the 3C11 monoclonal antibody to synapsin in Smed-prep(RNAi) with one eye (D), animals with no eyes (E), and gfp(RNAi) (F). Smed-prep(RNAi) animals (Figure S2B) (G) and gfp(RNAi) injected during regeneration. Staining with a probe to a glutamate receptor specific to CG/brain, branches, Smed-GluR, confirms reduction of CG structure to the most anterior tip (I). Smed-sFRP-1, a marker of anterior fate, is mostly absent or else confined to the very anterior tip (J). Staining with cintillo (K) shows that the number of these anterior cells is also reduced and restricted to the anterior tips of animals. Staining with the posterior brain marker Smed-WntA (red) shows that in animals where CG/brain is present A/P polarity of the brain (DAPI stained in blue) is maintained (L,P). gfp(RNAi) were normal for all these stains (M–P). Prolonged Smed-prep(RNAi) during homeostasis (Figure S2C) leads to the formation of two new eyes anterior to the original pair (Q) but not to any visible reduction or incorrect patterning of the CG/brain, as shown by Smed-GluR expression (R). The most anterior margin expression of Smed-sFRP-1 is lost in Smed-prep(RNAi) homeostasis worms (S). Smed-prep(RNAi) worms amputated laterally (Figure S2A) are able to regenerate CG, as shown by Smed-GluR expression (T), but the regeneration is not patterned correctly as branches are fused (see arrow in T) compared to gfp(RNAi) animals (U). All panels depict 12 day regenerating trunks except: (G,H) 12 day regenerating tails, (Q,R,S) 28 days homeostasis after first injection, (T,U) 15 days regeneration after lateral regeneration. All scale bars 1 mm. Tab. 1. Summary of phenotypes for Smed-prep(RNAi) experiments. 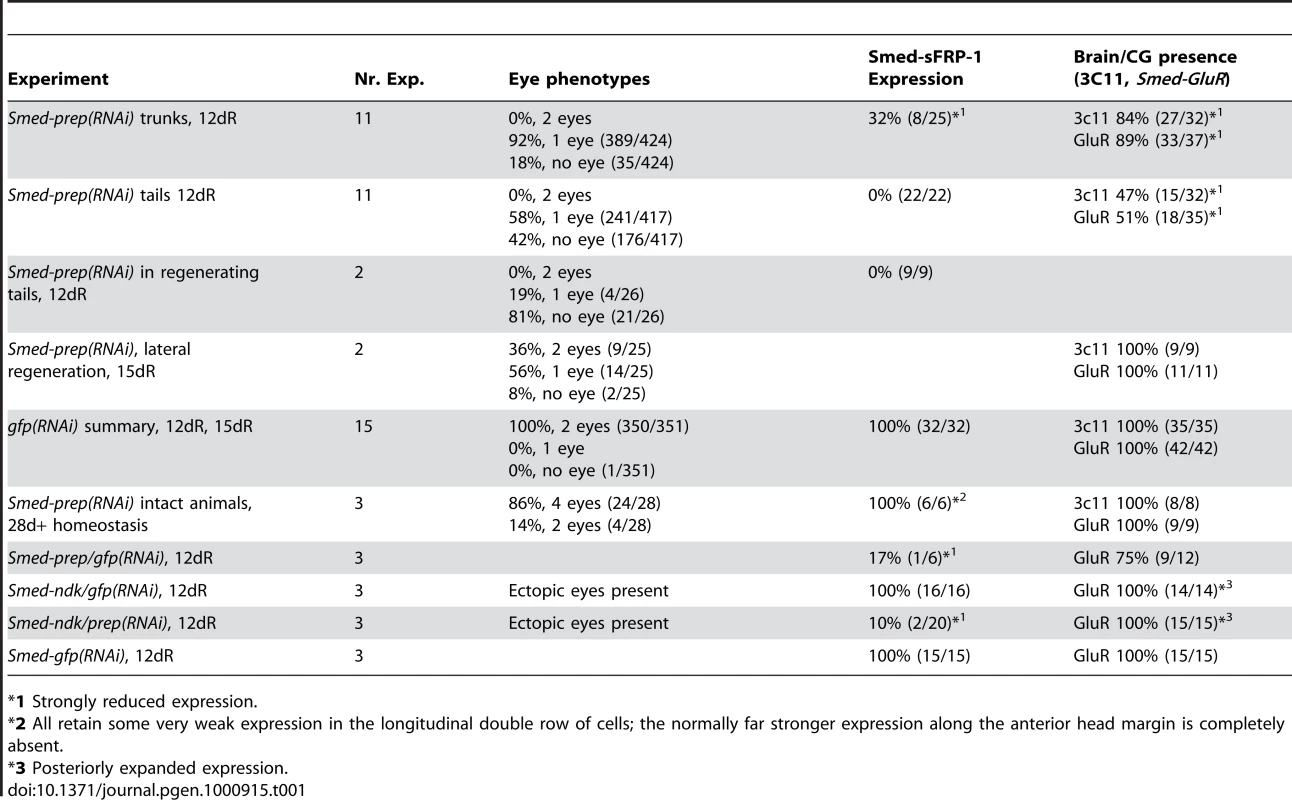
*1 Strongly reduced expression. We investigated the structure of the planarian ventral nerve cords (VNCs) and CG using the anti-SYNORF1 (3C11) cross-reactive monoclonal antibody [30]. We found that in all Smed-prep(RNAi) animals the CG were greatly reduced, with almost no brain at all discernible in the most severely affected RNAi worms (Figure 2D and 2E). In these animals anti-SYNORF1 positive cells do form from differentiating neoblast progeny in the anterior as part of the VNCs. Significantly, anti-SYNORF1 positive cells are present along the whole anteroposterior axis. This suggests, along with correct pharynx and posterior regeneration that Smed-prep(RNAi) does not affect the general ability of stem cells to differentiate. All control gfp(RNAi) animals were normal (Figure 2C and 2F). We confirmed the loss of CG by looking at the expression of Smed-GluR (specific for CG (Figure 2I and 2M). This loss of anterior structures suggests a role for Smed-prep in patterning anterior structures and/or a requirement for Smed-prep in allowing neoblasts to differentiate into CG cells. This phenotype is different from that previously described for the S. mediterranea ortholog of adenomatous polypolis coli (APC), a negative regulator of Wnt signaling. Smed-APC-1(RNAi) results in ectopic posterior fate at anterior blastemas [9].
To build a more exact picture of the requirements for Smed-prep we also investigated its role during regeneration more directly. We injected regenerating animals after amputation and then re-amputated (Figure S2). This approach has previously been used as a proxy to separate regeneration specific effects from homeostatic effects [15]. Control gfp(RNAi) worms regenerated normally but Smed-prep(RNAi) worms failed to make eyes and CG almost entirely (Figure 2G and 2H, Table 1). All animals did regenerate normal VNCs within regenerated anterior tissue. This confirms that new Smed-prep expression during regeneration is required to properly replace anterior structures.
To investigate whether Smed-prep was required specifically for stem cell progeny to differentiate to CG or instead primarily for global anterior fates we investigated the expression of cintillo [31] and Smed-sFRP-1 [9], [12]. These genes represent two different anterior markers that are not expressed in CG cells. We find that both cintillo and Smed-sFRP-1 expression are greatly reduced or absent in Smed-prep(RNAi) animals at 12 days of regeneration (Figure 2J and 2K). In the case of Smed-sFRP-1 expression we observed a correlation between the strength of the Smed-prep(RNAi) phenotype and whether any Smed-sFRP-1 expression was detectable. Those animals that maintained a single eye (and therefore some CG) also had some remaining Smed-sFRP-1 expression. Animals with stronger phenotypes (no eyes) had no detectable anterior Smed-sFRP-1 expression. All gfp(RNAi) animals had normal expression for both these markers (Figure 2N and 2O). Together these data suggest that Smed-prep is required for correct anterior blastema fate patterning during regeneration, rather than solely for CG formation by differentiating neoblasts.
This loss of anterior markers led us to consider whether Smed-prep(RNAi) leads to a homeotic like posteriorisation of the planarian body plan. We found no evidence for this by looking at the relative position of the regenerating or fully formed pharynx, the expression of a medial marker Smed-Tcen49 [32], or by looking at the expression of posterior markers such as Smed-HoxD [10]. Thus we infer that Smed-prep(RNAi) leads to a reduction in the formation of anterior structures, but neither a change to posterior fate at anterior blastemas nor an expansion in posterior or medial fates in existing tissues (Figure S2J, S2K, S2L, and S2M). We also found that early Smed-sFRP-1 expression at anterior blastemas at 24 hours of regeneration is absent in Smed-prep(RNAi) animals. This suggests Smed-prep acts to provide anterior fate and pattern the anterior blastema, after polarity is set (Figure S2H and S2I).
The planarian brain and the planarian head have distinct A/P polarity, as is the case in other animals [17]. Smed-prep expression is higher in the anterior and lateral margins of the planarian head (Figure 1B). We wished to know whether this was a reflection of Smed-prep having a role in defining different A/P fates within the anterior blastema itself. In this case any remaining brain fated tissues observed in Smed-prep(RNAi) animals (Table 1) would be expected to have posterior brain fate. By investigating the expression of Smed-WntA, a marker of the posterior brain [17] we found that Smed-prep(RNAi) animals that regenerated one eye and some CG also maintained antero-posterior identity within their much reduced anterior structures (Figure 2L and 2P). In these animals Smed-WntA still labels a posterior domain of the remaining CG. This suggests that Smed-prep is required to specify an anterior field of cells in which further A/P patterning occurs.
Smed-prep is required for anterior patterning but not for brain maintenance or regeneration during homeostasis or lateral regeneration
We performed long term Smed-prep(RNAi) in whole worms, to assess its role during normal homeostasis and tissue turnover. Long-term knockdown did not result in loss or proportional reduction of anterior structures or CG/Brain (Figure 2R, Table 1). However, Smed-prep(RNAi) worms developed a new pair of photoreceptors anterior to the original pair (Figure 2Q). This result suggests that Smed-prep expression in the anterior of whole worms is required for correct positioning of the photoreceptors during homeostasis but not for CG maintenance. Smed-sFRP-1 expression was also affected in these animals, with loss of anterior margin and lateral expression, but maintenance of weaker ventral antero-medial expression (Figure 2S). This provides more evidence to suggest that Smed-sFRP-1 expression is dependent on Smed-prep expression. These data show that Smed-prep has different roles in establishing anterior structures and their subsequent maintenance.
The finding that the CG were not reduced in homeostasis led us to consider whether Smed-prep(RNAi) would affect the lateral regeneration of anterior structures. We reasoned that if Smed-prep was not required for CG maintenance during homeostasis, then alternative anterior maintenance mechanisms must be active during homeostasis. These alternate mechanisms could also be sufficient to orchestrate lateral regeneration, a scenario where existing anterior structures are left partially intact. We cut Smed-prep(RNAi) worms longitudinally (Figure S2A) and observed regeneration. We found that Smed-prep(RNAi) worms were able to laterally regenerate all structures, with correct scaling, and subsequent normal behavior. While some worms did not regenerate a second eye correctly, all animals regenerated lateral CG. However, on looking at the pattern of the CG structure in more detail we noticed that the bilateral CG fused at the anterior tip (Figure 2T and 2U). In this regenerative scenario Smed-prep(RNAi) animals can regenerate antero-laterally but CG structures are not patterned correctly. This indicates that while Smed-prep is specifically required for the replacement of missing anterior structures when they are absent, it is not required to generate missing anterior fated structures during antero-lateral regeneration, i.e. when one side of the brain is still present. Instead, it is only required for the formation of correct pattern during this regenerative scenario. It seems likely that the remaining anterior tissue contains cues, generated downstream of Smed-prep during normal anterior regeneration, that are sufficient to direct neoblast progeny to CG fate.
Double Smed-prep/nou-darake(RNAi) shows that Smed-prep is required for anterior patterning but not for brain differentiation
Our experiments thus far suggest that Smed-prep is required for anterior patterning and fate. To formally rule out the possibility that Smed-prep is also directly required during anterior regeneration for stem cell differentiation into CG we utilized the previously described nou-darake (ndk) RNAi phenotype [16]. RNAi of this FGF-like receptor gene leads to ectopic posterior expansion of CG during homeostasis and regeneration. We predicted that if Smed-prep was required for anterior patterning but not for neoblast differentiation then double Smed-prep/ndk(RNAi) worms would display expanded CG differentiation, but with aberrant anterior patterning and loss of anterior marker expression. Smed-prep/gfp(RNAi) and Smed-ndk/gfp(RNAi) animals regenerated with reduced and expanded CG respectively compared to gfp(RNAi) worms (Figure 3B and 3C). Smed-prep/ndk(RNAi) animals had expanded CG but this expansion was patterned incorrectly (Figure 3D). The CG of Smed-prep/Smed-ndk(RNAi) animals are fused at the anterior tip, similar to Smed-prep(RNAi) laterally regenerated animals (Figure 3D). Both gfp(RNAi) and smed-ndk/gfp(RNAi) animals have normally patterned bilateral CG (Figure 3A and 3C). To test if this mispatterning was concomitant with the loss of anterior fate we also looked at Smed-sFRP-1 expression. Whereas Smed-sFRP-1 expression was normal in Smed-ndk(RNAi) animals after regeneration it was absent or greatly reduced in Smed-prep/ndk(RNAi) animals (Figure 3E–3G). This suggests that Smed-prep specifies an anterior domain during regeneration and that stem cell progeny normally differentiate to form CG only within this domain. This restriction requires activity of Smed-ndk, which is also expressed in an anterior domain. In double Smed-prep/ndk(RNAi) animals the loss of Smed-ndk removes this restriction on neoblast progeny, allowing them to adopt CG fate without the presence of Smed-prep expression, but does not rescue the defects in anterior patterning.
Fig. 3. Double Smed-prep/Smed-ndk(RNAi) and double Smed-prep/Smed-beta-catenin-1(RNAi) phenotypes further define the role of Smed-prep. 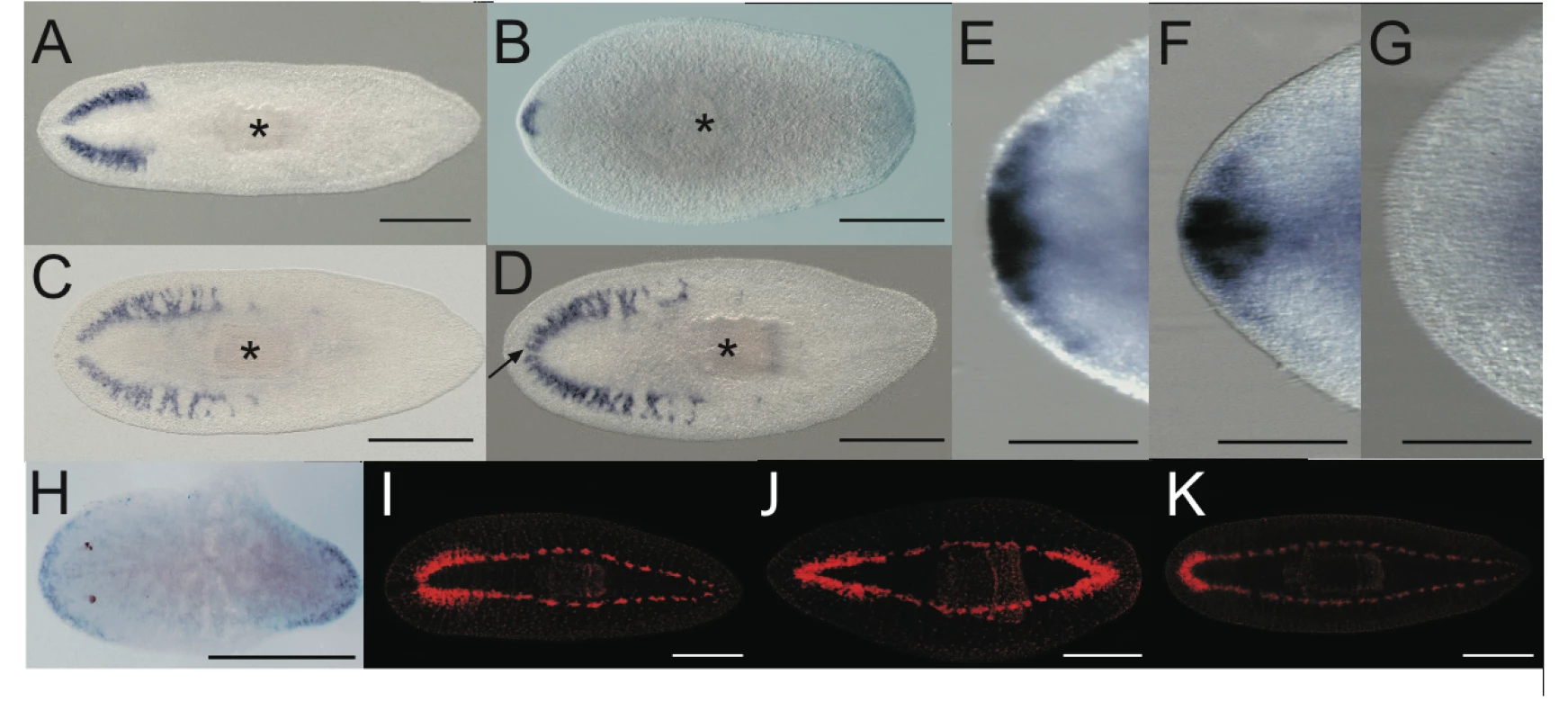
Smed-GluR expression in gfp(RNAi) (A), Smed-prep/gfp(RNAi) (B), Smed-ndk/gfp(RNAi) (C), and Smed-prep/ndk(RNAi) (D) animals. Smed-prep/ndk(RNAi) (D) animals have ectopic CG cells and have fused bilateral CG branches (arrow). Smed-prep/ndk(RNAi) (G) animals also fail to correctly express the anterior marker Smed-sFRP-1, which is expressed in gfp(RNAi) (E) and Smed-ndk/gfp(RNAi) (F) animals. Smed-beta-catenin-1(RNAi) animals (H) ectopically express Smed-prep at the “new” anterior end and Smed-beta-catenin-1/gfp(RNAi) animals regenerate heads at both blastemas of regenerating fragments (J). The regeneration of anterior structures is greatly reduced or entirely absent in posterior blastemas in Smed- prep/beta-catenin-1(RNAi) (K) and regneration is normal control (I) gfp(RNAi) animals, whereas the regenerated head in Smed-prep/beta-catenin-1(RNAi) shows the expected head reduction of Smed- prep(RNAi). All panels are trunk pieces accept (H) which is a head. All pieces are 15 day regenerants. All scales bars are 1 mm except (E–G) which are 500 µm. Smed-prep is required for formation of ectopic anterior structures in Smed-beta-catenin-1(RNAi) animals
Wnt signaling is central in patterning the antero-posterior axis of planarians by promoting posterior fate [9], [10], [12], [15]. Given the finding that Smed-prep is not required for CG maintenance or formation during homeostasis and lateral regeneration respectively, it remained unclear whether Smed-prep would be required for the ectopic anterior structures observed when Wnt signaling is attenuated. We found that when Smed-beta-catenin-1(RNAi) results in head regeneration at both anterior and posterior blastemas [3]–[5], ectopic and prolonged expression of Smed-prep in these new heads is observed (Figure 3H). In addition Smed-prep/beta-catenin-1(RNAi) reduced anterior structures at both ends (Figure 3K). As Smed-prep expression is initially present at both posterior and anterior blastemas our data suggest that active Wnt signaling in the posterior blastema suppresses Smed-prep action at posterior blastemas post-transcriptionally.
Smed-prep is the first gene clearly implicated as being necessary for promoting anterior fate during regeneration in S. mediterranea. We propose that after initial polarity determination, involving Wnt signals and other as yet unknown mechanisms, Smed-prep expression in neoblast progeny determines an anterior field of cells in which anterior structures differentiate and are patterned. At posterior blastemas Smed-prep activity is inhibited post-transcriptionally by Wnt activity. This now provides the opportunity to discover downstream genes that are required for further fine patterning during anterior regeneration, as some of these are likely transcriptional targets of Smed-prep activity.
In other animals the function of PREP TALE class homeodomains remains rather poorly defined compared to those of other TALE class family genes. In the both major invertebrate genetics models, C. elegans and D. melanogaster, a direct ortholog of PREP TALE class homeodomains is absent [18]. Interestingly both worms and flies contain MEIS orthologs (unc-62 and homothorax respectively) that have broad roles in specifying fate during development [33], [34] and other members of the nematode and arthropod phyla do have PREP orthologs [18]. The finding that PREP is involved in zebra fish brain development may suggest that PREP has an evolutionary conserved role in anterior fates. Broader phylogenetic study of its function is required to test this [21]. Here, we show that Smed-prep expression and function delineates the whole anterior domain, including all regions of the brain. Previous studies of Hox and Hox co-factor function have not implicated these two groups of genes in defining the most anterior structures of other vertebrates [35] or arthropods [36].
Significantly, the requirement for Smed-prep is observably different during homeostasis and different regenerative scenarios. This illustrates that the genetic networks available to solve different regenerative scenarios may be diverse and are likely to depend on the informational/signaling capacity of the differentiated portion of starting tissue. In addition it is the first time that homeobox transcription factors have been directly implicated in A/P patterning in planaria. We suspect that other conserved homeodomain proteins will also play core roles in specifying positional information during regeneration.
Materials and Methods
Animals
All experiments were performed with a clonal line originally generated from a single animal of the asexual strain of the planarian S. mediterranea collected in Montjuïc (provided by Professor Emili Saló i Boix) maintained at 20°C in tap water treated with activated charcoal and buffered with 0.5 ml/L 1 M NaHCO3. Planarians were fed veal liver and starved for at least one week prior to experiments.
Isolation of Smed-prep
To identify planarian homologues of TALE transcription factors we searched a local database of Version 3.1 of the S. mediterranea Genome Project for orthologs of mammalian TALE genes (http://genome.wustl.edu/genomes). The contigs 018898 and 020093 containing Smed-prep were analyzed using Vector NTI (Invitrogen) and sequence data supplemented by using RACE (Ambion RLM Race Kit). The primers Sm-Prep-Forward with sequence ATTGCTACTAGAGCAATGTGAACAAGC and Sm-Prep-Reverse with sequence ATTCTGCGTCGGGCATTGAT amplify a 810 bp fragment which was used for whole mount ISH hybridization and RNAi knockdown. PREP and TALE proteins sequences were taken from Mukherjee at al [18] and alignments checked with the CLUSTAL [37]. Phylogenetic reconstruction was conducted using MEGA version 4 using the bootstrapped neighbour-joining method [38]. The Smed-prep sequence has been submitted to GenBank with accession number GU290186.
RNAi
DsRNAs were synthesized as described previously [39]. Control animals were injected with dsRNA of GFP that has no homology in the planarian genome. DsRNA microinjection was performed as described elsewhere [27]. For injection schedules please refer to Figure S2. For double RNAi experiments concentrations for each gene were maintained at 1 µg/µl after mixing and for GFP controls 2 µg/µl was injected.
Whole-mount ISH hybridization, immuno-staining, and imaging
Whole mount ISH hybridization was carried out as described previously [25] with modifications described in [40] and [24]. The paraformaldehyde solution for the fixation step was prepared fresh and adjusted to pH 9.5 using NaOH.
For immuno-staining animals were killed in 2% HCl for 5 min on ice and then fixed in Carnoy's solution for 2 h at 4°C. After fixation, samples were processed as described elsewhere [41], [42]. The following primary antibodies were used: anti-SYNORF1, a mouse monoclonal antibody specific for synapsin (Developmental Studies HybridomaBank, dilution of 1∶25) and anti-arrestin VC-1, a mouse monoclonal antibody specific for planarian photosensitive cells (kindly provided by Hidefumi Orii, used at a dilution of 1∶15,000). Goat anti-mouse secondary antibody conjugated to Alexa 488 or Alexa 546 (Molecular Probes) was used at a 1∶400 dilution.
Brightfield pictures were taken on a Zeiss Discovery V8 from CarlZeiss using an AxioCam MRC from CarlZeiss. Fluorescent pictures were taken on a Leica MZ16F fluorescence stereomicroscope using a Leica DFC 300Fx camera (Leica Lasertechnik, Heidelberg). Confocal laser scanning microscopy was performed with a LeicaSP2 confocal laser scanning microscope (CLSM) (Leica Lasertechnik, Heidelberg).
Supporting Information
Zdroje
1. AgataK
UmesonoY
2008 Brain regeneration from pluripotent stem cells in planarian. Philos Trans R Soc Lond B Biol Sci 363 2071 2078
2. ReddienPW
Sanchez AlvaradoA
2004 Fundamentals of planarian regeneration. Annu Rev Cell Dev Biol 20 725 757
3. SaloE
2006 The power of regeneration and the stem-cell kingdom: freshwater planarians (Platyhelminthes). Bioessays 28 546 559
4. ReddienPW
BermangeAL
MurfittKJ
JenningsJR
Sanchez AlvaradoA
2005 Identification of genes needed for regeneration, stem cell function, and tissue homeostasis by systematic gene perturbation in planaria. Dev Cell 8 635 649
5. ReddienPW
OviedoNJ
JenningsJR
JenkinJC
Sanchez AlvaradoA
2005 SMEDWI-2 is a PIWI-like protein that regulates planarian stem cells. Science 310 1327 1330
6. UmesonoY
AgataK
2009 Evolution and regeneration of the planarian central nervous system. Dev Growth Differ 51 185 195
7. GlazerA
WilkinsonA
BackerCB
LapanS
GutzmanJH
2009 The Zn Finger protein Iguana impacts Hedgehog signaling by promoting ciliogenesis. Dev Biol
8. MorganTH
1900 Regeneration in Planarians. Archiv Fur Entwicklungsmechanik Der Organismen (copy in filing cabinet) 10 58 119
9. GurleyKA
RinkJC
Sanchez AlvaradoA
2008 Beta-catenin defines head versus tail identity during planarian regeneration and homeostasis. Science 319 323 327
10. IglesiasM
Gomez-SkarmetaJL
SaloE
AdellT
2008 Silencing of Smed-{beta}catenin1 generates radial-like hypercephalized planarians. Development 135 1215 1221
11. MolinaMD
SaloE
CebriaF
2007 The BMP pathway is essential for re-specification and maintenance of the dorsoventral axis in regenerating and intact planarians. Dev Biol 311 79 94
12. PetersenCP
ReddienPW
2008 Smed-betacatenin-1 is required for anteroposterior blastema polarity in planarian regeneration. Science 319 327 330
13. ReddienPW
BermangeAL
KiczaAM
Sanchez AlvaradoA
2007 BMP signaling regulates the dorsal planarian midline and is needed for asymmetric regeneration. Development 134 4043 4051
14. AdellT
SaloE
BoutrosM
BartschererK
2009 Smed-Evi/Wntless is required for beta-catenin-dependent and -independent processes during planarian regeneration. Development 136 905 910
15. PetersenCP
ReddienPW
2009 A wound-induced Wnt expression program controls planarian regeneration polarity. Proc Natl Acad Sci U S A 106 17061 17066
16. CebriaF
KobayashiC
UmesonoY
NakazawaM
MinetaK
2002 FGFR-related gene nou-darake restricts brain tissues to the head region of planarians. Nature 419 620 624
17. KobayashiC
SaitoY
OgawaK
AgataK
2007 Wnt signaling is required for antero-posterior patterning of the planarian brain. Dev Biol 306 714 724
18. MukherjeeK
BurglinTR
2007 Comprehensive analysis of animal TALE homeobox genes: new conserved motifs and cases of accelerated evolution. J Mol Evol 65 137 153
19. RyanJF
MazzaME
PangK
MatusDQ
BaxevanisAD
2007 Pre-bilaterian origins of the Hox cluster and the Hox code: evidence from the sea anemone, Nematostella vectensis. PLoS ONE 2 e153 doi:10.1371/journal.pone.0000153
20. BerthelsenJ
ZappavignaV
FerrettiE
MavilioF
BlasiF
1998 The novel homeoprotein Prep1 modulates Pbx-Hox protein cooperativity. EMBO J 17 1434 1445
21. DeflorianG
TisoN
FerrettiE
MeyerD
BlasiF
2004 Prep1.1 has essential genetic functions in hindbrain development and cranial neural crest cell differentiation. Development 131 613 627
22. LaurentA
BihanR
OmilliF
DeschampsS
PellerinI
2008 PBX proteins: much more than Hox cofactors. Int J Dev Biol 52 9 20
23. MoensCB
SelleriL
2006 Hox cofactors in vertebrate development. Dev Biol 291 193 206
24. Gonzalez-EstevezC
ArseniV
ThambyrajahRS
FelixDA
AboobakerAA
2009 Diverse miRNA spatial expression patterns suggest important roles in homeostasis and regeneration in planarians. Int J Dev Biol 53 493 505
25. UmesonoY
WatanabeK
AgataK
1997 A planarian orthopedia homolog is specifically expressed in the branch region of both the mature and regenerating brain. Dev Growth Differ 39 723 727
26. FireA
XuS
MontgomeryMK
KostasSA
DriverSE
1998 Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391 806 811
27. Sanchez AlvaradoA
NewmarkPA
1999 Double-stranded RNA specifically disrupts gene expression during planarian regeneration. Proc Natl Acad Sci U S A 96 5049 5054
28. CebriaF
GuoT
JopekJ
NewmarkPA
2007 Regeneration and maintenance of the planarian midline is regulated by a slit orthologue. Dev Biol 307 394 406
29. SakaiF
AgataK
OriiH
WatanabeK
2000 Organization and regeneration ability of spontaneous supernumerary eyes in planarians -eye regeneration field and pathway selection by optic nerves. Zoolog Sci 17 375 381
30. CebriaF
2008 Organization of the nervous system in the model planarian Schmidtea mediterranea: an immunocytochemical study. Neurosci Res 61 375 384
31. OviedoNJ
NewmarkPA
Sanchez AlvaradoA
2003 Allometric scaling and proportion regulation in the freshwater planarian Schmidtea mediterranea. Dev Dyn 226 326 333
32. BuenoD
VispoM
SanchoV
RomeroR
2001 Maintenance of A/P body regions in planarians by tcen49, a putativ cystine-knot neurotrophin. Belg J Zool 131S1 89 95
33. Van AukenK
WeaverD
RobertsonB
SundaramM
SaldiT
2002 Roles of the Homothorax/Meis/Prep homolog UNC-62 and the Exd/Pbx homologs CEH-20 and CEH-40 in C. elegans embryogenesis. Development 129 5255 5268
34. AldazS
MorataG
AzpiazuN
2005 Patterning function of homothorax/extradenticle in the thorax of Drosophila. Development 132 439 446
35. WilsonSW
HouartC
2004 Early steps in the development of the forebrain. Dev Cell 6 167 181
36. LynchJA
BrentAE
LeafDS
PultzMA
DesplanC
2006 Localized maternal orthodenticle patterns anterior and posterior in the long germ wasp Nasonia. Nature 439 728 732
37. HigginsDG
ThompsonJD
GibsonTJ
1996 Using CLUSTAL for multiple sequence alignments. Methods Enzymol 266 383 402
38. TamuraK
DudleyJ
NeiM
KumarS
2007 MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24 1596 1599
39. BoutrosM
KigerAA
ArmknechtS
KerrK
HildM
2004 Genome-wide RNAi analysis of growth and viability in Drosophila cells. Science 303 832 835
40. NogiT
LevinM
2005 Characterization of innexin gene expression and functional roles of gap-junctional communication in planarian regeneration. Dev Biol 287 314 335
41. BasyukE
BertrandE
JournotL
2000 Alkaline fixation drastically improves the signal of in situ hybridization. Nucleic Acids Res 28 E46
42. CebriaF
NewmarkPA
2005 Planarian homologs of netrin and netrin receptor are required for proper regeneration of the central nervous system and the maintenance of nervous system architecture. Development 132 3691 3703
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2010 Číslo 4
-
Všechny články tohoto čísla
- Whole-Genome SNP Association in the Horse: Identification of a Deletion in Myosin Va Responsible for Lavender Foal Syndrome
- Human Telomeres Are Hypersensitive to UV-Induced DNA Damage and Refractory to Repair
- Fragilities Caused by Dosage Imbalance in Regulation of the Budding Yeast Cell Cycle
- Admixture Mapping Scans Identify a Locus Affecting Retinal Vascular Caliber in Hypertensive African Americans: the Atherosclerosis Risk in Communities (ARIC) Study
- Activation of Estrogen-Responsive Genes Does Not Require Their Nuclear Co-Localization
- Genetic Tests for Ecological and Allopatric Speciation in Anoles on an Island Archipelago
- A Pax3/Dmrt2/Myf5 Regulatory Cascade Functions at the Onset of Myogenesis
- Two New Loci for Body-Weight Regulation Identified in a Joint Analysis of Genome-Wide Association Studies for Early-Onset Extreme Obesity in French and German Study Groups
- Hypomethylation of a LINE-1 Promoter Activates an Alternate Transcript of the MET Oncogene in Bladders with Cancer
- Candidate Causal Regulatory Effects by Integration of Expression QTLs with Complex Trait Genetic Associations
- Combined Inactivation of pRB and Hippo Pathways Induces Dedifferentiation in the Retina
- Allele-Specific Virulence Attenuation of the HopZ1a Type III Effector via the ZAR1 Resistance Protein
- Down-Regulation of Honey Bee Gene Biases Behavior toward Food Rich in Protein
- A Microarray-Based Genetic Screen for Yeast Chronological Aging Factors
- Actin-Related Protein Arp6 Influences H2A.Z-Dependent and -Independent Gene Expression and Links Ribosomal Protein Genes to Nuclear Pores
- Phosphorylation of the Conserved Transcription Factor ATF-7 by PMK-1 p38 MAPK Regulates Innate Immunity in
- A -Regulatory Signature for Chordate Anterior Neuroectodermal Genes
- Genetic Analysis of Fin Development in Zebrafish Identifies Furin and Hemicentin1 as Potential Novel Fraser Syndrome Disease Genes
- Assembly of a 40 Mb Eukaryotic Genome from Short Sequence Reads: , a Model Organism for Fungal Morphogenesis
- Trait-Associated SNPs Are More Likely to Be eQTLs: Annotation to Enhance Discovery from GWAS
- Absence of Evidence for MHC–Dependent Mate Selection within HapMap Populations
- The TALE Class Homeobox Gene Defines the Anterior Compartment for Head Regeneration
- Cyclic Expression of Lhx2 Regulates Hair Formation
- Genetic Evidence for Hybrid Trait Speciation in Butterflies
- Epigenetic Regulation of a Murine Retrotransposon by a Dual Histone Modification Mark
- Chromosome 9p21 SNPs Associated with Multiple Disease Phenotypes Correlate with Expression
- S Phase Progression in Human Cells Is Dictated by the Genetic Continuity of DNA Foci
- The Next Generation Becomes the Now Generation
- Acts as a Tumor Suppressor in a Murine Retinoblastoma Model by Facilitating Tumor Cell Death
- Genome-Wide Association Study of Lp-PLA Activity and Mass in the Framingham Heart Study
- The Five Zinc Transporters Undergo Different Evolutionary Fates towards Adaptive Evolution to Zinc Tolerance in
- MicroRNA–Directed siRNA Biogenesis in
- Deletion of the WD40 Domain of LRRK2 in Zebrafish Causes Parkinsonism-Like Loss of Neurons and Locomotive Defect
- Incipient Balancing Selection through Adaptive Loss of Aquaporins in Natural Populations
- GTPase Activity Plays a Key Role in the Pathobiology of LRRK2
- Natural Single-Nucleosome Epi-Polymorphisms in Yeast
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Whole-Genome SNP Association in the Horse: Identification of a Deletion in Myosin Va Responsible for Lavender Foal Syndrome
- Admixture Mapping Scans Identify a Locus Affecting Retinal Vascular Caliber in Hypertensive African Americans: the Atherosclerosis Risk in Communities (ARIC) Study
- Genetic Tests for Ecological and Allopatric Speciation in Anoles on an Island Archipelago
- Human Telomeres Are Hypersensitive to UV-Induced DNA Damage and Refractory to Repair
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



