-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
Reklama: Dissecting the Molecular Interface between Pathogen and Plant
article has not abstract
Published in the journal: . PLoS Pathog 8(11): e32767. doi:10.1371/journal.ppat.1002955
Category: Pearls
doi: https://doi.org/10.1371/journal.ppat.1002955Summary
article has not abstract
Fungal diseases of plants represent one of the most eminent threats to agriculture. Given the food needs of a growing world population and that more and more crops are devoted to fuel production, the necessity to develop crops with better resistance to disease is increasing. To accomplish this, the mechanisms that plant pathogenic fungi use to colonize plants need to be elucidated. As of now, there are only few examples/models in which this can be done on a functional, genome-wide level, taking into account both the pathogen and its host plant [1]. The fungus Ustilago maydis (U. maydis) is one of these examples. It is a member of the smut fungi: a large group of parasites infecting mostly grasses, including several important crop plants such as maize (Figure 1B), wheat, barley, and sugar cane. Smut fungi are biotrophs, i.e., parasites that need the living host plant to complete their sexual life cycle [2], [3]. They do not establish prominent feeding structures like the related, haustoria-forming rust fungi. During penetration, the host plasma membrane invaginates and completely encases the intracellular hyphae (Figure 1A), establishing an extended interaction zone [4] mediating the exchange of molecules between fungus and host. In contrast to most smut fungi that cause a systemic infection, remaining symptomless until the plant flowers, U. maydis can infect all above-ground parts of the maize plant but fails to spread systemically. U. maydis induces local tumors in which spores develop (Figure 1B) – a unique feature that allows detection of symptoms in corn seedlings less than a week after syringe infection with high levels of inoculum. This, together with the toolbox developed for reverse genetics, cell biology, and functional studies, has contributed to its status as a model for biotrophic basidiomycete fungi [5]. Here the current level of our understanding of the elaborate molecular crosstalk between U. maydis and its host plant will be discussed.
Fig. 1. Disease symptoms and schematic presentation of effector cocktail use in different maize organs and tissues infected by U. maydis. 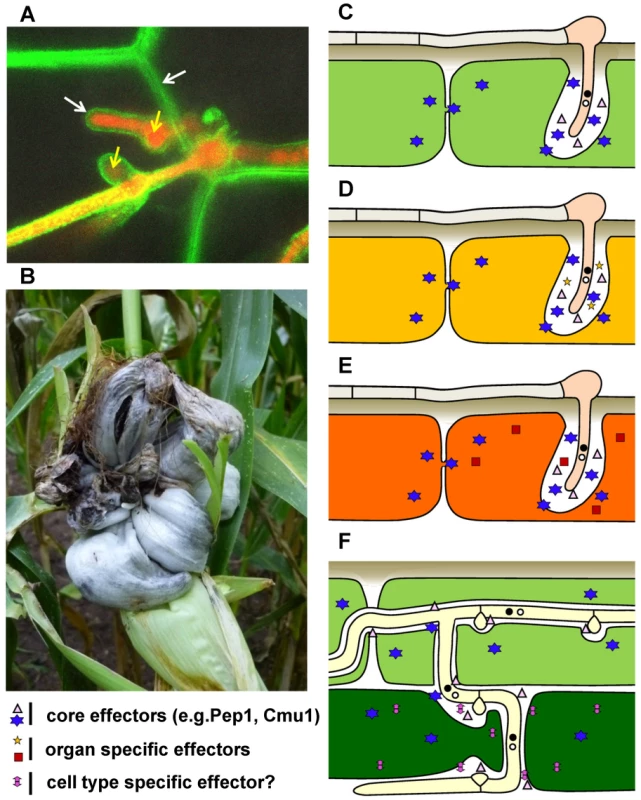
A) Confocal microscopy of a U. maydis strain expressing cytosolic mRFP (yellow arrows) during intracellular growth in epidermal maize cells expressing PIN-YFP as a plasma membrane marker (white arrows). B) U. maydis tumor on field-grown maize plant (picture kindly provided by S. Krombach). C–F depict schematically the different tissues infected by U. maydis (the width of the interaction zone between hyphae and host plasma membrane is not drawn to scale): C) epidermal cell of an infected maize seedling (light green); D) epidermal cell of an infected mature leaf (yellow); E) epidermal cell of infected tassel (orange; F) epidermal cell (light green) and mesophyll cells (dark green) of infected seedling. Core effectors, organ-specific effectors, and cell type–specific effectors with either apoplastic or cytoplasmic function inside plant cells are indicated with different symbols. Secreted Effectors of U. maydis
Sequencing of the U. maydis genome and transcriptional profiling of different infection stages paved the path for the discovery of effector genes that govern interaction with the host plant. Effectors are broadly defined as microbe-derived secreted molecules that shape interaction with the host. In lower eukaryotes, effector identification is largely restricted to conventionally secreted proteins, as these can be recognized through the presence of a signal peptide. The U. maydis genome codes for about 550 predicted secreted proteins of which more than 50% are novel, lacking known Interpro domains [6], [7]. Many of these novel genes reside in gene clusters, are upregulated during host colonization, and encode effectors with a virulence function [6], [8]. Comparative approaches have shown that many of these novel effectors are also found in related smut fungi [8], [9]. They are either highly conserved or poorly conserved, leading to speculation that they may target equally conserved or variable host proteins, respectively [8], [9]. A recent study revealed that effector genes are differentially expressed in different infected maize organs and that some effector mutants affect tumor formation only in specific organs [10]. This mirrors the finding that different maize organs express distinct sets of proteins and suggests that the ability of U. maydis to induce tumors in different maize organs relies on its ability to reprogram different plant developmental states to suit its own needs, i.e., for fungal proliferation in tumor tissue [10]. Skibbe et al. [10] suggest a two-step process in which a first set of “core” effectors (Figure 1C–E) is used to suppress plant defense responses during penetration, and a second set of specific effectors responds to maize organ–specific properties (organ-specific effectors, Figure 1D, E). This indicates that U. maydis is able to sense the different developmental conditions of its host and reacts by secreting a cocktail of specifically tuned effectors for reprogramming these tissues. Linking effectors with virulence may thus require infecting different maize organs during different developmental stages rather than performing seedling infections with mutant strains, as is common practice nowadays. It is even conceivable that effectors could act in a cell type–specific manner, i.e., there could be effectors that are needed in epidermal tissue, and a different effector set might be required in mesophyll cells (Figure 1F). To identify such cell type–specific effectors, one could assume that the respective genes are transcriptionally upregulated in specific cell types and apply laser-dissection methods together with next-generation sequencing methods.
Effectors Acting in the Biotrophic Interface between U. maydis Hyphae and Host
During the early phase of maize infection, U. maydis is recognized by the plant via conserved molecular patterns (PAMPs). This leads to salicylate-dependent defense responses, a typical response of plants to biotrophs [11], [12]. These plant defense responses are considered to be overcome/suppressed with the help of the set of core effector molecules that are upregulated during penetration. One of these core effectors is Pep1 (Figure 1C–F). pep1 mutants are able to form normal appressoria but are arrested during penetration and induce strong plant defense responses that include the massive transcriptional upregulation of the maize-secreted peroxidase POX12 and an accumulation of H2O2 at attempted sites of penetration [13].
A biologically active Pep1-mCherry fusion protein localizes to the biotrophic interaction zone, suggesting that Pep1 is an apoplastic effector [13]. In a recent study, Pep1 is shown to interact directly with POX12 and to inhibit its activity [14]. Since the main sources for reactive oxygen species (ROS) in the apoplast are NADPH oxidases and peroxidases, Hemetsberger et al. [14] go on to show that biotrophic development of the pep1 mutant can be partially restored by scavenging ROS as well as by silencing peroxidase POX12. The inhibition of plant peroxidases by Pep1 appears to be rather unspecific, as horse radish peroxidase is also inhibited [14]. Thus, the Pep1 effector targets an apoplastic peroxidase, a critical component of the PAMP-triggered defense program.
The recent analysis of a cluster of four genes transcriptionally upregulated during biotrophic development revealed that two of those genes, pit1 and pit2, are important for tumor induction [15]. pit1 as well as pit2 mutants are able to colonize maize plants but are unable switch to strong proliferation and tumor induction. While pit2 codes for a small secreted effector that accumulates in the biotrophic interphase, Pit1 is a transmembrane protein that localizes to the hyphal tips, moving early endosomes as well as vacuolar structures [15]. Intriguingly, pit1 and pit2 mutants induce transcriptional programs in infected maize plants that are indistinguishable, suggesting a yet unknown functional link between the apoplastic effector Pit2 and the transmembrane protein Pit1 [15].
U. maydis Effectors Acting inside Plant Cells
In all plant-pathogen systems where respective analyses have been done, the repertoire of effectors consists of those functioning in the apoplast and those taken up by plant cells [16]. In U. maydis, a secreted chorismate mutase, Cmu1, serves as the first example of an effector that is translocated into the host cell. The cmu1 gene of U. maydis is among the most strongly induced genes during plant colonization, and the Cmu1 protein is the most abundant fungal protein detected in the apoplast [17]. By complementation in yeast as well in vitro enzymatic assays, Cmu1 was shown to be a chorismate mutase. Chorismate is the branching metabolite of the shikimate pathway. Chorismate mutase catalyzes the conversion of chorismate to prephenate, which is further converted to an array of different phenylpropanoid compounds. Chorismate is also the precursor for aromatic amino acids and the plant defense hormone salicylic acid. In plants, the shikimate pathway resides in plastids [18]. Nevertheless, cytosolic plant chorismate mutases exist, and by two hybrid data, Cmu1 was shown to form heterodimers with the plastidic as well as the cytosolic isoforms [17], [19]. Immunolocalization studies indicate a cytosolic localization for Cmu1 after translocation into the host plant cell. In accordance with metabolic profiles of infected maize, a rechanneling of the chorismate flow is suggested by the cooperative action of the cytosolic maize chorismate mutase together with Cmu1, leading to a reduction of available chorismate for salicylic acid biosynthesis [17]. The ability of Cmu1 to spread locally to neighboring yet uninfected host cells (most likely via plasmodesmata) has been interpreted as metabolic priming, leading to lower salicylic acid levels, that prepares cells for the upcoming colonization by U. maydis [17]. Based on the presence of cmu1-related genes in other smut fungi [17], Cmu1 is also considered to be a core effector (Figure 1C–F).
The Interface between U. maydis Hyphae and Host as the Site for Obtaining Nutrients
For the acquisition of nutrients, biotrophic fungi have to divert the metabolism of the host to provide them with nutrients via the extracellular, biotrophic interphase. This is mechanistically not trivial, as the extracellular release of monosaccharides has been shown to trigger plant defense responses [20]. U. maydis is shown to express a novel, plasma membrane-localized saccharose transporter (Srt1) during its biotrophic phase whose deletion strongly affects virulence. Srt1 is an H+-symporter specific for sucrose and displays an unusually high substrate affinity. These features are perfectly in-line with the needs of a biotroph: Srt1 guarantees efficient carbon supply and transports the disaccharide saccharose without producing apoplastic signals that trigger plant defenses [21]. Its functional role as a saccharose transporter during biotrophic development was corroborated by demonstrating that the srt1 deletion phenotype can be complemented by the saccharose transporter AtSuc9 from Arabidopsis thaliana. Due to its low Km at a pH of 5.5 that is relevant for the apoplast, Srt1 can compete efficiently with plant saccharose transporters [22].
U. maydis infections lead to massive changes in the metabolome of infected plant tissue [11]. These metabolic changes could all result from repressing/modulating defense signaling pathways on different levels. Alternatively, the strongly elevated metabolites observed during infection could be of nutritional value for U. maydis, which would make this pathogen a “molecular farmer.” Yet another possibility would be that some of the metabolites induced are used by U. maydis to defend the habitat against other microbes. Compounds like DIMBOA (2,4-dihydroxy-7-methoxy-1,4-benzoxazin-3-one), sesquiterpenes, and defensins could conceivably have such a role. Genes involved in the synthesis of these compounds like Bx1 (benzoxazinless 1), catalyzing the initial DIMBOA biosynthesis step, the sequiterpene cyclase umi2, and the defensin-related gene umi11 are upregulated in tumor tissue [23].
With more genome sequences of related smut fungi with highly syntenic genomes becoming available [6], [8], [9], the identification of conventionally secreted effectors has become an easier task as many of these genes are only poorly conserved. Comparative genomics also allow for identification of effector sets that might have a role in determining host specificity. In addition, comparative genomics is a valuable tool for defining conserved effector domains for functional assays. However, to fully comprehend and appreciate the manipulative toolbox of a biotrophic pathogen like U. maydis, it will be necessary to decipher the functions of the >250 unknown effectors and to link these functions with the observed transcriptomic and metabolomic changes in the host. Given that many secreted effectors are species, genus, or family specific [9], [24]–[26], the huge challenge that needs to be met in the future is to find out whether eukaryotic plant pathogens target the same or different pathways in their respective hosts.
Zdroje
1. DeanR, van KunJA, PretoriusZA, Hammond-KosackKE, Di PietroA, et al. (2012) The top 10 fungal pathogens in molecular plant pathology. Mol Plant Pathol 13 : 414–430.
2. BanuettF (1995) Genetics of Ustilago maydis, a fungal pathogen that induces tumors in maize. Annu Rev Genet 29 : 179–208.
3. BrefortT, DoehlemannG, Mendoza-MendozaA, ReissmannS, DjameiA, et al. (2009) Ustilago maydis as a pathogen. Annu Rev Phytopathol 47 : 423–445.
4. BauerR, OberwinklerF, VánkyK (1997) Ultrastructural markers and systematics in smut fungi and allied taxa. Can J Bot 75 : 1273–1314.
5. VollmeisterE, SchipperK, BaumannS, HaagC, PohlmannT, et al. (2012) Fungal development of the plant pathogen Ustilago maydis. FEMS Microbiol Rev 36 : 59–77.
6. KamperJ, KahmannR, BolkerM, MaLJ, BrefortT, et al. (2006) Insights from the genome of the biotrophic fungal plant pathogen Ustilago maydis. Nature 444 : 97–101.
7. MuellerO, KahmannR, AguilarG, Trejo-AguilarB, WuA, et al. (2008) The secretome of the maize pathogen Ustilago maydis. Fungal Genet Biol 45 Suppl 1: S63–70.
8. SchirawskiJ, MannhauptG, MunchK, BrefortT, SchipperK, et al. (2010) Pathogenicity determinants in smut fungi revealed by genome comparison. Science 330 : 1546–1548.
9. LaurieJD, AliS, LinningR, MannhauptG, WongP, et al. (2012) Genome comparison of barley and maize smut fungi reveals targeted loss of RNA silencing components and species-specific presence of transposable elements. Plant Cell 24 : 1733–45.
10. SkibbeDS, DoehlemannG, FernandesJ, WalbotV (2010) Maize tumors caused by Ustilago maydis require organ-specific genes in host and pathogen. Science 328 : 89–92.
11. DoehlemannG, WahlR, HorstRJ, VollLM, UsadelB, et al. (2008) Reprogramming a maize plant: transcriptional and metabolic changes induced by the fungal biotroph Ustilago maydis. Plant J 56 : 181–195.
12. GlazebrookJ (2005) Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu Rev Phytopathol 43 : 205–227.
13. DoehlemannG, van der LindeK, AßmannD, SchwammbachD, HofA, et al. (2009) Pep1, a secreted effector protein of Ustilago maydis, is required for successful invasion of plant cells. PLoS Pathog 5: e1000290 doi:10.1371/journal.ppat.10000290
14. HemetsbergerC, HerrbergerC, ZechmannB, HillmerM, DoehlemannG (2012) The Ustilago maydis effector Pep1 suppresses plant immunity by inhibition of host peroxidase activity. PLoS Pathog 8: e1002684 doi:10.1371/journal.ppat.1002684
15. DoehlemannG, ReissmannS, AssmannD, FleckensteinM, KahmannR (2011) Two linked genes encoding a secreted effector and a membrane protein are essential for Ustilago maydis-induced tumour formation. Mol Microbiol 81 : 751–766.
16. StergiopoulosI, de WitPJ (2009) Fungal effector proteins. Annu Rev Phytopathol 47 : 233–263.
17. DjameiA, SchipperK, RabeF, GhoshA, VinconV, et al. (2011) Metabolic priming by a secreted fungal effector. Nature 478 : 395–398.
18. TzinV, GaliliG (2010) New insights into the shikimate and aromatic amino acids biosynthesis pathways in plants. Mol Plant 3 : 956–972.
19. EberhardJ, EhrlerTT, EppleP, FelixG, RaeseckeHR, et al. (1996) Cytosolic and plastidic chorismate mutase isozymes from Arabidopsis thaliana: molecular characterization and enzymatic properties. Plant J 10 : 815–821.
20. HerbersK, MeuwlyP, MetrauxJP, SonnewaldU (1996) Salicylic acid-independent induction of pathogenesis-related protein transcripts by sugars is dependent on leaf developmental stage. FEBS Lett 397 : 239–244.
21. WahlR, WippelK, GoosS, KämperJ, SauerN (2010) A novel high-affinity sucrose transporter is required for virulence of the plant pathogen Ustilago maydis. PLoS Biol 8: e1000303 doi:10.1371/journal.pbio.1000303
22. WippelK, SauerN (2012) Arabidopsis SUC1 loads the phloem in suc2 mutants when expressed from the SUC2 promoter. J Exp Bot 63 : 669–679.
23. BasseCW (2005) Dissecting defense-related and developmental transcriptional responses of maize during Ustilago maydis infection and subsequent tumor formation. Plant Physiol 138 : 1774–1784.
24. LevesqueCA, BrouwerH, CanoL, HamiltonJP, HoltC, et al. (2010) Genome sequence of the necrotrophic plant pathogen Pythium ultimum reveals original pathogenicity mechanisms and effector repertoire. Genome Biol 11: R73.
25. KemenE, GardinerA, Schultz-LarsenT, KemenAC, BalmuthAL, et al. (2011) Gene gain and loss during evolution of obligate parasitism in the white rust pathogen of Arabidopsis thaliana. PLoS Biol 9: e1001094 doi:10.1371/journal.pbio.1001094
26. LinksMG, HolubE, JiangRH, SharpeAG, HegedusD, et al. (2011) De novo sequence assembly of Albugo candida reveals a small genome relative to other biotrophic oomycetes. BMC Genomics 12 : 503.
Štítky
Hygiena a epidemiologie Infekční lékařství Laboratoř
Článek vyšel v časopisePLOS Pathogens
Nejčtenější tento týden
2012 Číslo 11- Jak souvisí postcovidový syndrom s poškozením mozku?
- Měli bychom postcovidový syndrom léčit antidepresivy?
- Farmakovigilanční studie perorálních antivirotik indikovaných v léčbě COVID-19
- 10 bodů k očkování proti COVID-19: stanovisko České společnosti alergologie a klinické imunologie ČLS JEP
-
Všechny články tohoto čísla
- Egyptian H5N1 Influenza Viruses—Cause for Concern?
- Epigenetics of Host–Pathogen Interactions: The Road Ahead and the Road Behind
- Environmental Regulation of Prions in Yeast
- : Dissecting the Molecular Interface between Pathogen and Plant
- A Wolf in Sheep's Clothing: SV40 Co-opts Host Genome Maintenance Proteins to Replicate Viral DNA
- Exploring New Biological Functions of Amyloids: Bacteria Cell Agglutination Mediated by Host Protein Aggregation
- A Trade-off between the Fitness Cost of Functional Integrases and Long-term Stability of Integrons
- Intracellular Vesicle Acidification Promotes Maturation of Infectious Poliovirus Particles
- Distinct Effects on Diversifying Selection by Two Mechanisms of Immunity against
- Whole Genome Sequencing Reveals Local Transmission Patterns of in Sympatric Cattle and Badger Populations
- Early Mechanisms of Pathobiology Are Revealed by Transcriptional Temporal Dynamics in Hippocampal CA1 Neurons of Prion Infected Mice
- Revised Phylogeny and Novel Horizontally Acquired Virulence Determinants of the Model Soft Rot Phytopathogen SCC3193
- : Where Does It Live?
- The Role of Auxin-Cytokinin Antagonism in Plant-Pathogen Interactions
- PLOS Pathogens
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- A Wolf in Sheep's Clothing: SV40 Co-opts Host Genome Maintenance Proteins to Replicate Viral DNA
- Intracellular Vesicle Acidification Promotes Maturation of Infectious Poliovirus Particles
- Revised Phylogeny and Novel Horizontally Acquired Virulence Determinants of the Model Soft Rot Phytopathogen SCC3193
- The Role of Auxin-Cytokinin Antagonism in Plant-Pathogen Interactions
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



