-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaTranscription coactivator and lncRNA duet evoke Hox genes
article has not abstract
Published in the journal: . PLoS Genet 13(6): e32767. doi:10.1371/journal.pgen.1006797
Category: Perspective
doi: https://doi.org/10.1371/journal.pgen.1006797Summary
article has not abstract
Mammalian genomes are pervasively transcribed [1, 2] to produce thousands of long noncoding RNAs (lncRNAs) [3–5], transcripts that are more than 200 nucleotides in length that do not code for proteins. Although only a handful of functional lncRNAs have been well characterized to date, recent work suggests that some lncRNAs have crucial roles in the control of gene expression during both normal development and disease, through multiple mechanisms [6, 7]. As new lncRNAs are being discovered at a rapid pace, their molecular mechanisms are continuing to be enriched and diversified. For example, a few lncRNAs have been shown to affect expression of nearby genes through recruitment of protein regulatory complexes [8–10], while it is suggested that others function akin to enhancers and local regulators in cis [11, 12]. In fact, such local gene-regulatory mechanisms have been invoked to explain the observation that lncRNA expression is often correlated with the expression of nearby genes—the so-called “guilt by association” [13].
The cis-acting mechanisms of lncRNAs are largely unknown. Whether loci encoding lncRNAs function via their lncRNA transcripts or DNA elements is often unclear. An important question in the field centers around distinguishing between at least 2 possible mechanisms—either the function of many of the lncRNAs that appear to regulate genes in cis depends on the RNAs themselves or their effect is mediated by enhancer-like activity of underlying DNA elements in the lncRNA locus, the act of transcription, and/or splicing of lncRNAs [14, 15]. Furthermore, the molecular components regulating the expression of lncRNAs remain mostly unexplored. An understanding of the possible commonalities of the disparate underlying mechanisms may facilitate instructive and predictive models of lncRNA function.
Pradeepa et al. [16] set out to elucidate 1 mechanism of lncRNA regulation and function using the HoxA locus as their model. The Bickmore group had previously shown that positive cofactor 4 (PC4) and splicing factor 2 (SF2) interacting protein (Psip1), also known as lens epithelium-derived growth factor (LEDGF), played an important role in the regulation of Hox genes [17] and had more recently demonstrated the role of the p75 isoform of Psip1 (Psip1/p75) in recruiting the Trithorax/mixed lineage leukemia (MLL) complex to expressed Hox genes [18]. Interestingly, loss of Psip1 led not only to reduced binding of the MLL complex and loss of histone 3 lysine 4 trimethylation (H3K4me3) at the distal HoxA genes, it also resulted in complete loss of expression of the lncRNA HoxA transcript at the distal tip (Hottip), transcribed in an antisense direction away from the distal end of the 5′ HoxA cluster [10]. This suggested that Psip1 might function as a transcriptional regulator of Hottip expression. Following up on this possible connection, the authors showed that knockdown of Psip1/p52, the p52 isoform of Psip1, or Hottip, led to down-regulation of multiple 5′ HoxA genes; knockdown of p52 also strongly down-regulated Hottip expression. Consistent with these observations, MLL occupancy was significantly reduced across posterior HoxA genes upon knockdown of p52 or Hottip compared to controls.
The authors next took advantage of CRISPR-Cas9 technology to create gene-body deletions of Hottip. Loss of Hottip expression led to decreased expression of posterior (HoxA13, HoxA11, and HoxA10) and increased expression of anterior (HoxA2, HoxA6, and HoxA7) HoxA genes. To find direct genomic targets of Hottip, chromatin isolation by RNA purification [19] (ChIRP) was performed, which demonstrated specific occupancy of Hottip RNA over the promoters of HoxA13 and HoxA11. Importantly, ectopic activation of full-length Hottip via dCas9-mediated transcriptional activation showed specific induction of posterior (HoxA13, HoxA11, and HoxA10) but not anterior HoxA or HoxD genes. Additionally, premature termination of Hottip RNA by insertion of a synthetic polyadenylation cassette downstream of the Hottip transcription start site significantly reduced HoxA13 and HoxA11 mRNA levels in vitro and in vivo, firmly establishing a role for the intact Hottip lncRNA molecule and distinguishing its requirement from the act of transcription at the Hottip locus in the regulation of gene expression in cis. Taken together, the study provides a solid mechanism for the control of posterior HoxA gene transcription through activation of Hottip lncRNA by Psip1/p52 (Fig 1).
Fig. 1. Model of positive cofactor 4 (PC4) and splicing factor 2 (SF2) interacting protein (Psip1) action. 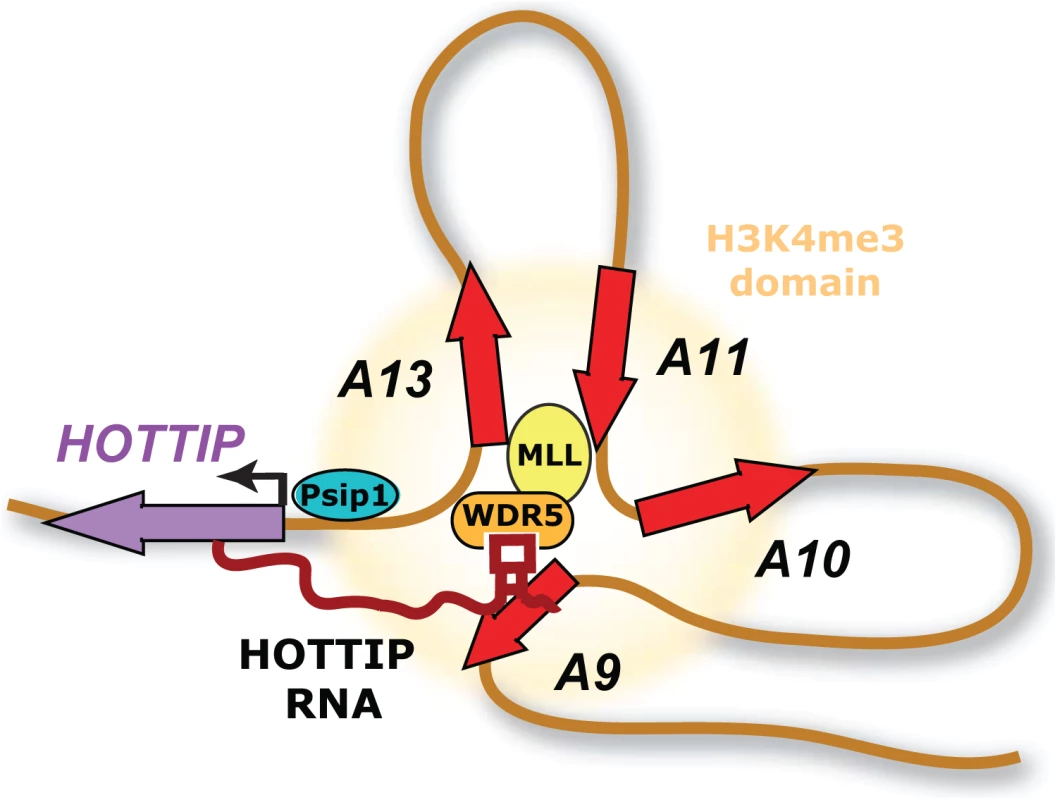
Psip1 activates the transcription of Hottip long noncoding RNA (lncRNA) in distal limb cells. Hottip lncRNA in turn stimulates the transcription of 5′ HoxA genes by enforcing H3K4me3 chromatin modification. Hottip’s selective effect on 5′ HoxA genes is due to the chromosome looping that brings 5′ HoxA genes into the vicinity of Hottip RNA emanating from the Hottip DNA locus. This mechanism transmits the spatial information in DNA looping into chemical information in chromatin modification and gene activation. Abbreviations: WDR5, WD repeat-containing protein 5; MLL, mixed lineage leukemia. Several interesting and important questions remain. Psip1 binds to 5′ HoxA genes but somehow still requires Hottip RNA to activate these genes. Thus, the concept of transcription coactivator using an enhancer-like RNA to amplify or expand its reach may apply to other pairs as well. These findings also provide substantial insights into the regulation of Hottip, the expression of which is dysregulated in a number of human cancers. Hottip appears to act locally near its site of transcription, but the mechanism through which Hottip RNA localizes specifically to the distal HoxA genes in cis is unknown. It is possible that Hottip takes advantage of the existing 3-dimensional chromosomal structure at the distal HoxA loci [10] to affect the local transcription landscape. This would be an attractive mechanism given the now-appreciated highly conserved hierarchical organization of the genome [20], in which fundamental structural units serve to guide regulatory elements or, in this case, specific RNA transcripts to their cognate promoters. These interactions could provide insight into the way in which the 3-dimensional organization of the genome reflects alterations in lineage and stage-specific transcriptional programs that govern cell fate, with lncRNAs such as Hottip acting as well-timed molecular switches. This may represent a fundamental property of mammalian gene regulatory networks, whose mechanisms for specificity and the general applicability represent key areas for future investigation.
Zdroje
1. Mattick JS. RNA regulation: a new genetics? Nat Rev Genet. Nature Publishing Group; 2004;5 : 316–323. doi: 10.1038/nrg1321 15131654
2. Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, et al. RNA Maps Reveal New RNA Classes and a Possible Function for Pervasive Transcription. Science. 2007;316 : 1484–1488. doi: 10.1126/science.1138341 17510325
3. Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10 : 155–159. doi: 10.1038/nrg2521 19188922
4. Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009;136 : 629–641. doi: 10.1016/j.cell.2009.02.006 19239885
5. Wilusz JE, Sunwoo H, Spector DL. Long noncoding RNAs: functional surprises from the RNA world. Genes & Development. 2009;23 : 1494–1504. doi: 10.1101/gad.1800909 19571179
6. Wang KCK, Chang HYH. Molecular mechanisms of long noncoding RNAs. Molecular Cell. 2011;43 : 904–914. doi: 10.1016/j.molcel.2011.08.018 21925379
7. Fatica A, Bozzoni I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet. 2014;15 : 7–21. doi: 10.1038/nrg3606 24296535
8. Lee JT. Lessons from X-chromosome inactivation: long ncRNA as guides and tethers to the epigenome. Genes & Development. 2009;23 : 1831–1842. doi: 10.1101/gad.1811209 19684108
9. Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R, et al. The Air Noncoding RNA Epigenetically Silences Transcription by Targeting G9a to Chromatin. Science. 2008;322 : 1717–1720. doi: 10.1126/science.1163802 18988810
10. Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011;472 : 120–124. doi: 10.1038/nature09819 21423168
11. Ørom UA, Derrien T, Beringer M, Gumireddy K, Gardini A, Bussotti G, et al. Long noncoding RNAs with enhancer-like function in human cells. Cell. 2010;143 : 46–58. doi: 10.1016/j.cell.2010.09.001 20887892
12. Guil S, Esteller M. Cis-acting noncoding RNAs: friends and foes. Nat Struct Mol Biol. 2012;19 : 1068–1075. doi: 10.1038/nsmb.2428 23132386
13. Rinn JL, Chang HY. Genome Regulation by Long Noncoding RNAs. Annu Rev Biochem. 2012;81 : 145–166. doi: 10.1146/annurev-biochem-051410-092902 22663078
14. Bassett AR, Akhtar A, Barlow DP, Bird AP, Brockdorff N. Considerations when investigating lncRNA function in vivo. Elife. 2014.
15. Engreitz JM, Haines JE, Perez EM, Munson G, Chen J, Kane M, et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature Publishing Group. 2016;539 : 452–455. doi: 10.1038/nature20149 27783602
16. Pradeepa MM, McKenna F, Taylor GCA, Bengani H, Grimes GR, Wood AJ, et al. (2017) Psip1/p52 regulates posterior Hoxa genes through activation of lncRNA Hottip. PLoS Genet 13(4): e1006677. doi: 10.1371/journal.pgen.1006677 28384324
17. Sutherland HG, Newton K, Brownstein DG, Holmes MC, Kress C, Semple CA, et al. Disruption of Ledgf/Psip1 results in perinatal mortality and homeotic skeletal transformations. Mol Cell Biol. American Society for Microbiology; 2006;26 : 7201–7210. doi: 10.1128/MCB.00459-06 16980622
18. Pradeepa MM, Grimes GR, Taylor GCA, Sutherland HG, Bickmore WA. Psip1/Ledgf p75 restrains Hox gene expression by recruiting both trithorax and polycomb group proteins. Nucleic Acids Res. 2014;42 : 9021–9032. doi: 10.1093/nar/gku647 25056311
19. Chu CC, Qu KK, Zhong FLF, Artandi SES, Chang HYH. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Molecular Cell. 2011;44 : 667–678. doi: 10.1016/j.molcel.2011.08.027 21963238
20. Lupiáñez DG, Spielmann M, Mundlos S. Breaking TADs: How Alterations of Chromatin Domains Result in Disease. Trends Genet. 2016;32 : 225–237. doi: 10.1016/j.tig.2016.01.003 26862051
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2017 Číslo 6
Nejčtenější v tomto čísle- Lokiarchaea are close relatives of Euryarchaeota, not bridging the gap between prokaryotes and eukaryotes
- Transcription coactivator and lncRNA duet evoke Hox genes
- Optimal sequencing strategies for identifying disease-associated singletons
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Současné možnosti léčby obezity
nový kurzAutoři: MUDr. Martin Hrubý
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



